- The paper demonstrates that phenotype discovery time scales inversely with phenotype frequency, leading to the 'arrival of the frequent' in evolving populations.
- It derives a mean-field model and validates it via simulations on random and RNA GP maps, confirming theoretical predictions across diverse evolutionary regimes.
- The results highlight that biased genotype-phenotype maps constrain adaptation, favoring frequently accessed yet suboptimal phenotypes and challenging classical evolutionary models.
Bias in Genotype-Phenotype Maps and Its Role in Evolutionary Dynamics
Overview
This paper investigates the profound evolutionary consequences arising from the highly non-uniform mapping of genotypes to phenotypes in biological systems. By rigorously analyzing both abstract random GP (genotype-phenotype) maps and more realistic RNA sequence–structure mappings, the authors demonstrate that the inherent bias in these maps can cause certain phenotypes—those mapping from disproportionately many genotypes—to be repeatedly explored and fixed in populations, sometimes at the expense of even fitter but rarer phenotypes. This "arrival of the frequent" effect results from extreme non-ergodicity in the evolutionary process, challenging traditional population genetic frameworks that implicitly assume accessible fitness landscapes and sufficiently rapid exploration of phenotype space.
Mean-Field Theory and Scaling of Discovery Time
The core of the paper is the derivation and empirical validation of a mean-field model for the discovery time Tp of new phenotypes. The model assumes that, for most biologically relevant genotype-phenotype maps, three properties generally hold:
- Genotypic Redundancy: NG≫NP, resulting in vast neutral networks.
- Neutral Exploration: Movement within a neutral space enables populations to sample a broader set of phenotypic neighbors.
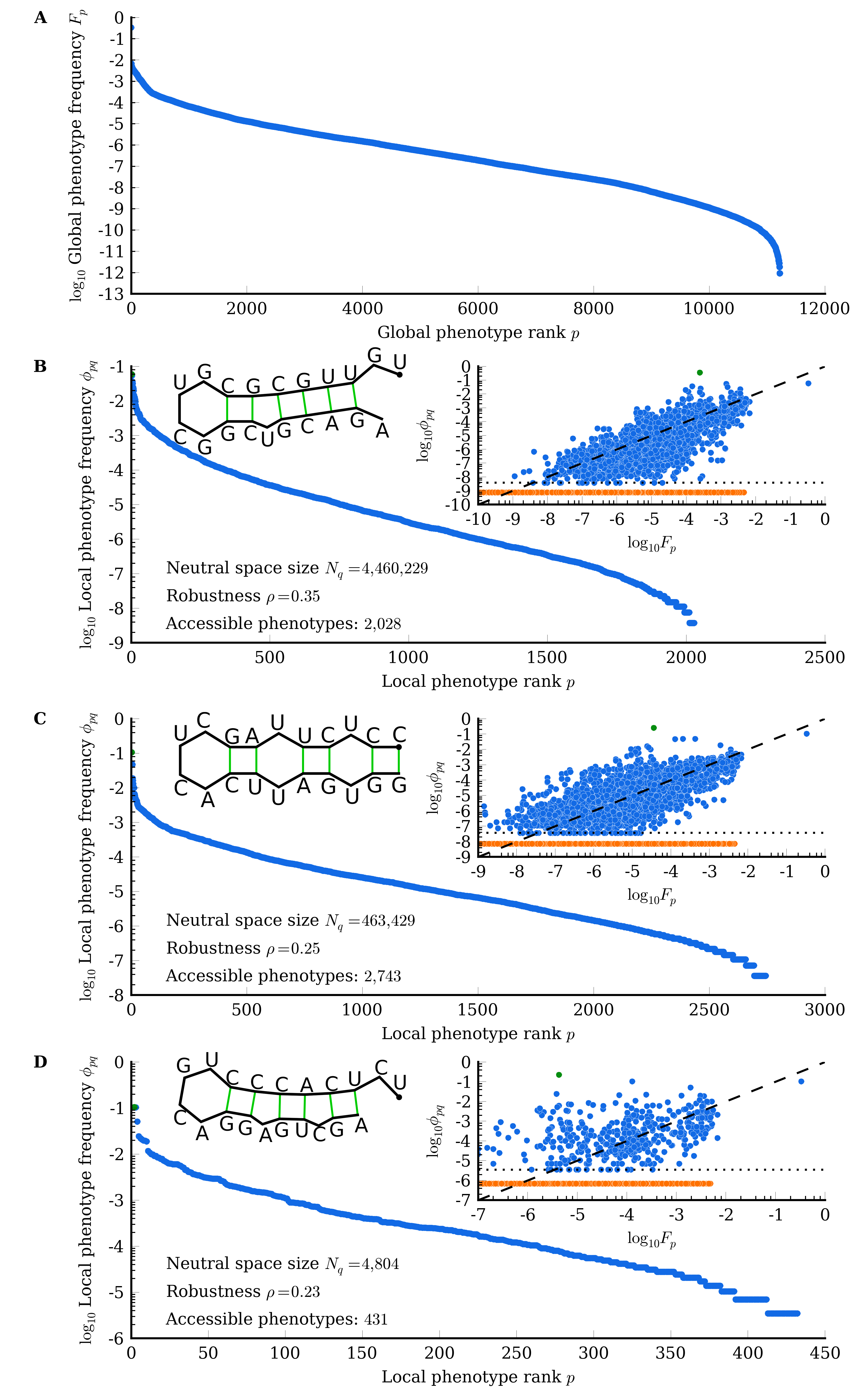
- Phenotypic Bias: The distribution of phenotype frequencies Fp is heavily skewed, such that most genotypes map to a small subset of phenotypes.
By formalizing these principles, the authors show analytically that the probability of mutating from a genotype in neutral space Nq to a phenotype p, ϕpq, scales to leading order with Fp (global phenotype frequency). The mean time to first discover phenotype p is therefore inversely proportional to its frequency, Tp∼1/Fp, across a wide range of parameters involving population size N, mutation rate μ, and genome length L.
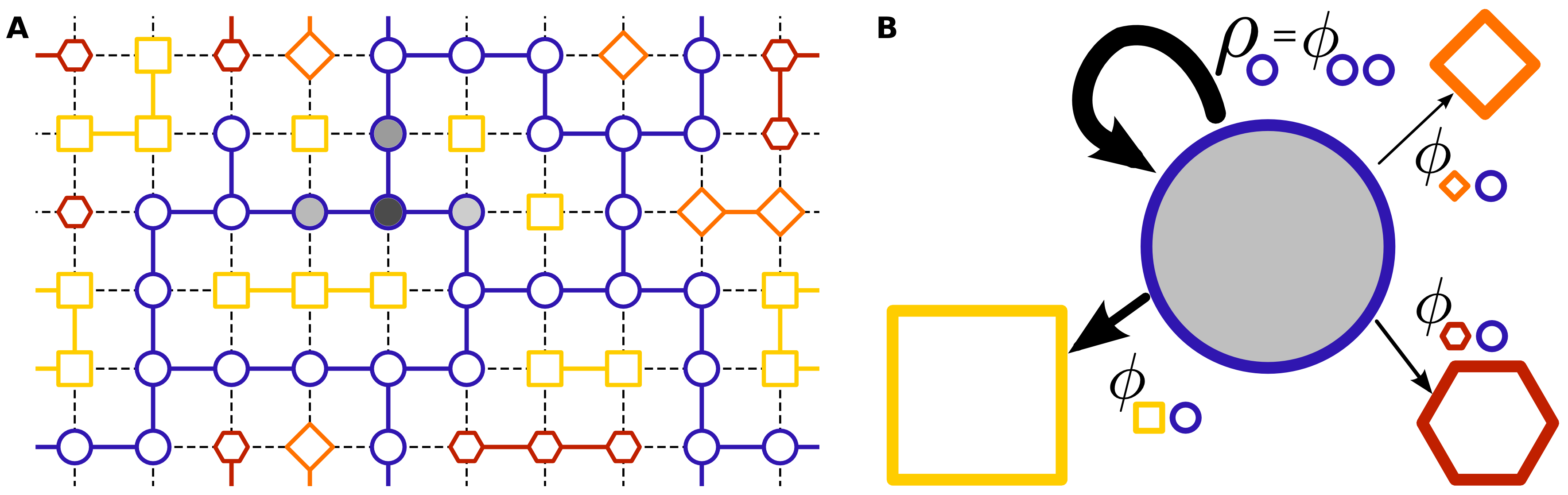
Figure 1: Schematic of the mean-field neutral space model demonstrating how phenotype discovery is governed by the structure of the genotype-phenotype map.
Validation on Random and RNA GP Maps
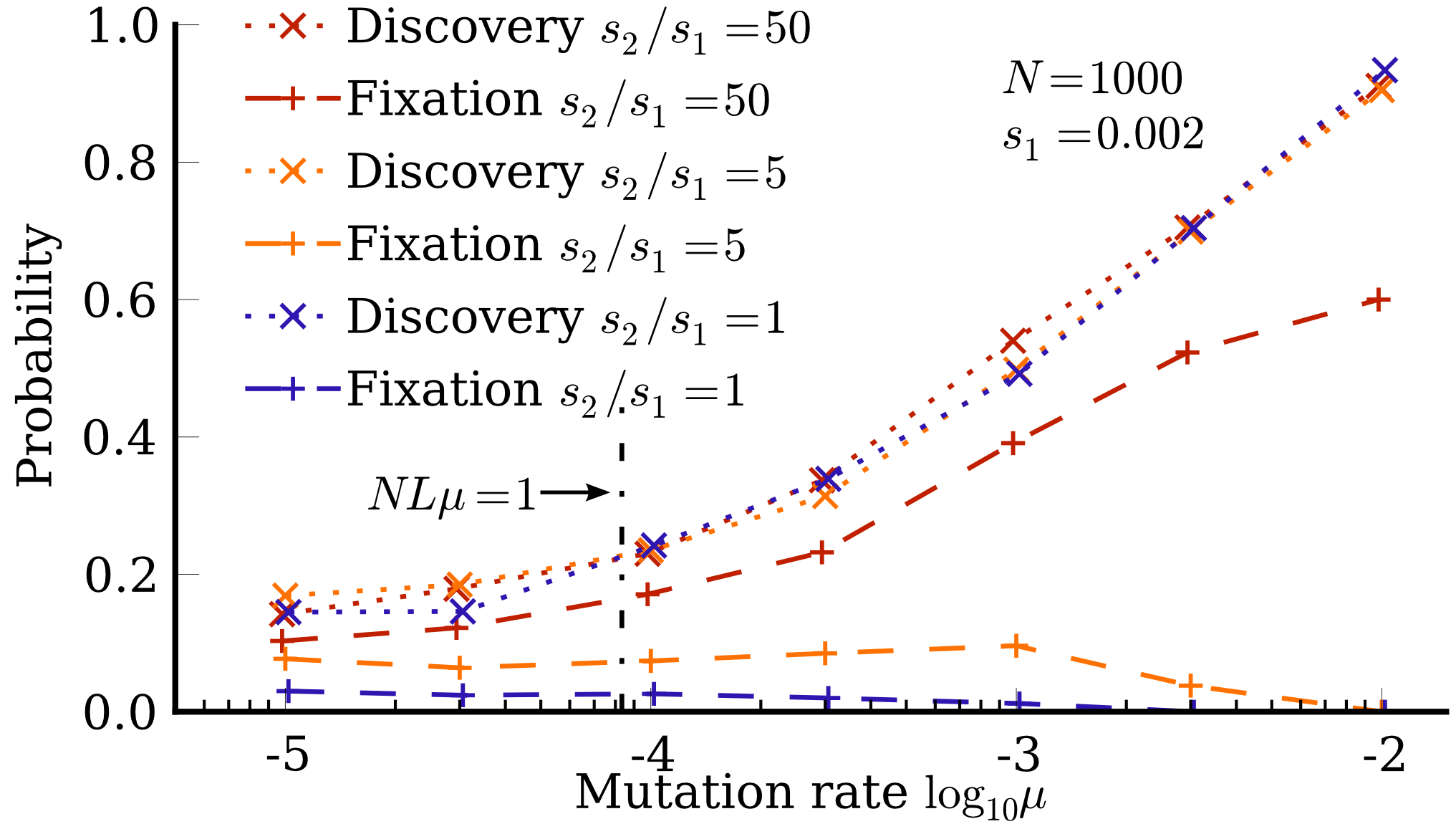
Simulations were performed for both stylized random GP maps and realistic RNA secondary structure mappings (L=12 and L=20):
Phenotypic Bias and Adaptive Trajectories
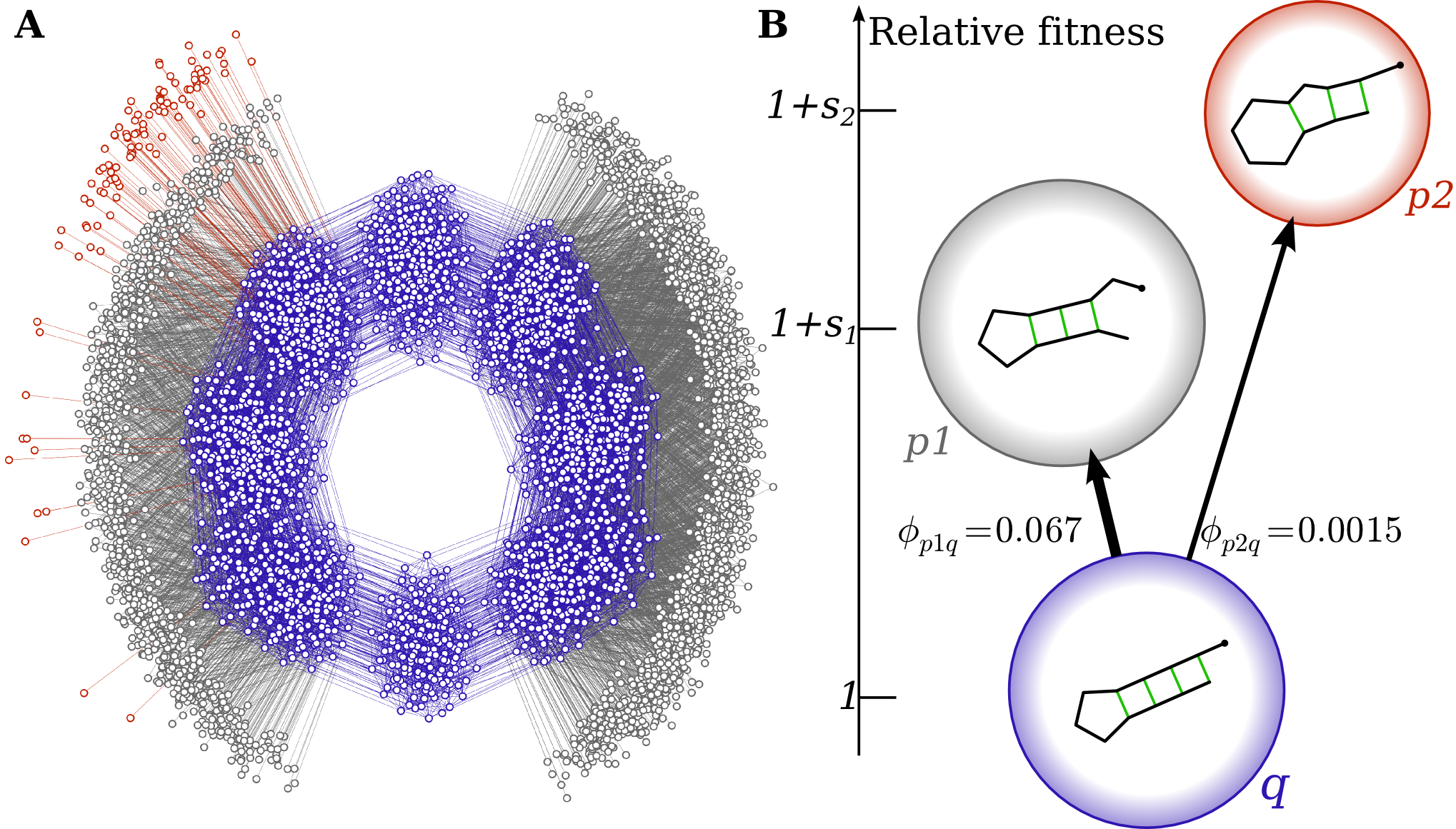
The practical implication is that the likelihood of a phenotype's appearance—and thus its possible fixation and adaptive role—is determined less by its absolute fitness and more by its mutational accessibility (encoded by Fp or ϕpq). The "arrival of the frequent" becomes critical in adaptive scenarios: When several mutant phenotypes are more fit than the incumbent, the one with higher local entry flux is likely to fix, even if an alternate phenotype is far fitter:
These results contradict the expectations of standard Wright-Fisher and origin-fixation models, which postulate that the fittest accessible phenotype will generally fix. Here, the bias in the GP map fundamentally distorts the adaptive walk by skewing the supply of variation. This effect is further accentuated in natural systems due to the high dimensionality and size of genotype space.
Broader Implications and Theoretical Insights
These findings suggest several key theoretical and practical implications for evolutionary biology and related computational fields:
Future Directions
Future research may focus on the interplay between the structure of GP maps and the dynamics of adaptation in even larger and more complex genotype spaces—with an emphasis on higher-order mutational connectivity, recombination, and horizontal transfer. Incorporating these results into evolutionary algorithms may also dramatically influence the design of optimization landscapes and inform the development of more robust, evolvable systems in synthetic biology and AI, where the mapping between "genotype" and "function" is explicitly constructed or learned.
Conclusion
The paper rigorously demonstrates that the inherent bias in GP maps constitutes a critical constraint on evolutionary outcomes. The mutational arrival rates of novel phenotypes, multi-orders-of-magnitude apart due to this bias, robustly steer populations toward frequent phenotypes even under strong selection for rarer, higher-fitness options. This "arrival of the frequent" mechanism disrupts standard evolutionary predictions, highlighting a previously underappreciated source of evolutionary contingency and restriction. The results evoke a re-examination of evolutionary assumptions in both biological and artificial evolving systems, urging an integrated modeling of the genotype-phenotype map in evolutionary theory.