- The paper introduces a deep multi-instance learning (MIL) framework that eliminates the need for detailed manual annotations in whole mammogram classification.
- It presents three approaches—max pooling, label assignment, and sparse MIL—that exploit patch-level features from convolutional neural networks to enhance malignancy detection.
- Results on the INbreast dataset demonstrate significant performance improvements and better localization of malignant regions over traditional methods.
Overview
The paper "Deep Multi-instance Networks with Sparse Label Assignment for Whole Mammogram Classification" proposes a novel approach for mammogram classification leveraging deep multi-instance learning (MIL) techniques. The authors focus on the challenges associated with conventional mammogram classification methods, which typically require extensive manual annotation and specialized models for detecting such annotations during testing. By eliminating the need for costly annotations, the paper introduces three end-to-end trained deep multi-instance networks—max pooling-based, label assignment-based, and sparse deep multi-instance learning networks—to enhance classification performance on whole mammograms.
Methodology
Deep Multi-instance Network Framework
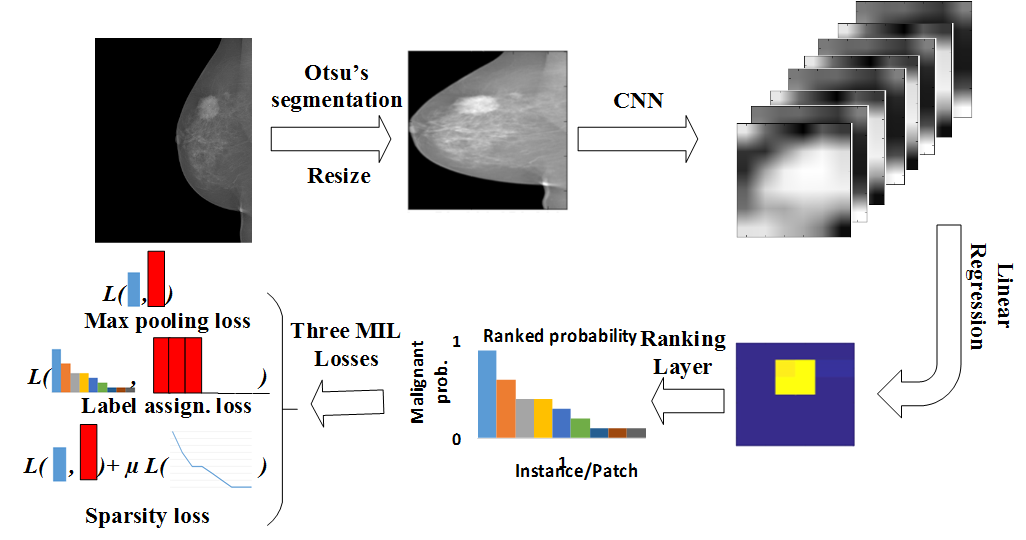
Figure 1: The proposed deep multi-instance network framework. First, we use Otsu's segmentation to remove the background and resize the mammogram to 224\times224. Second, the deep multi-instance network accepts the resized mammogram as input to the convolutional layers. Third, linear regression with weight sharing is employed for the malignant probability of each position from the convolutional neural network (CNN) feature maps of high channel dimensions. Then the responses of the instances/patches are ranked. Lastly, the learning loss is calculated using max pooling loss, label assignment, or sparsity loss for the three different schemes.
The authors propose a deep multi-instance network framework, consisting of convolutional layers followed by a ranking layer and a multi-instance loss layer. The design harnesses convolutional neural networks (CNNs) to efficiently derive features across multiple patches simultaneously. The patches are then evaluated for malignancy probability using a logistic regression layer with shared weights. By treating each patch of a mammogram as an instance, the entire mammogram is conceptualized as a bag of instances, transforming the classification problem into an MIL problem.
Multi-instance Learning Variants
Max Pooling-based Learning
The max pooling assumption posits that if any instance within the bag is positive, the entire bag is positive. Thus, the malignant probability of a mammogram is determined by the highest probability among its patches. This scheme, however, limits the exploitation of data by focusing solely on the patch with the highest probability, excluding informative patches with lower probabilities.
Label Assignment-based Learning
The label assignment-based approach extends the MIL to consider multiple top patches when assigning labels, rather than relying on a single maximum probability. The method assumes that the highest probability patches correlate with the mammogram's label. By assigning positive labels to the k highest-ranking patches, it exploits more data and offers better training dynamics, albeit with challenges in determining the optimal k.
Sparse Multi-instance Learning
The sparse MIL technique incorporates prior knowledge about mass sparsity, enforcing a constraint where only a fraction—approximately 2\%—of the patches belong to a malignant region. It thus balances the assumptions between max pooling and label assignment, utilizing an adaptive approach for training.
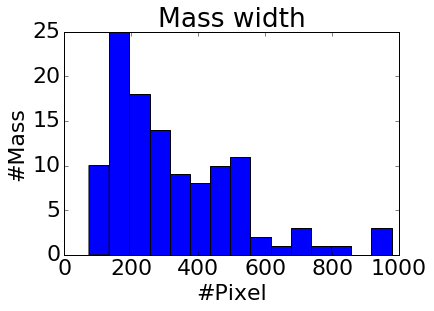
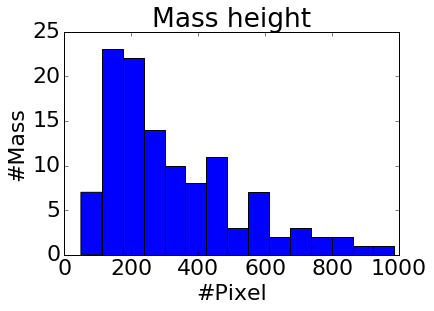
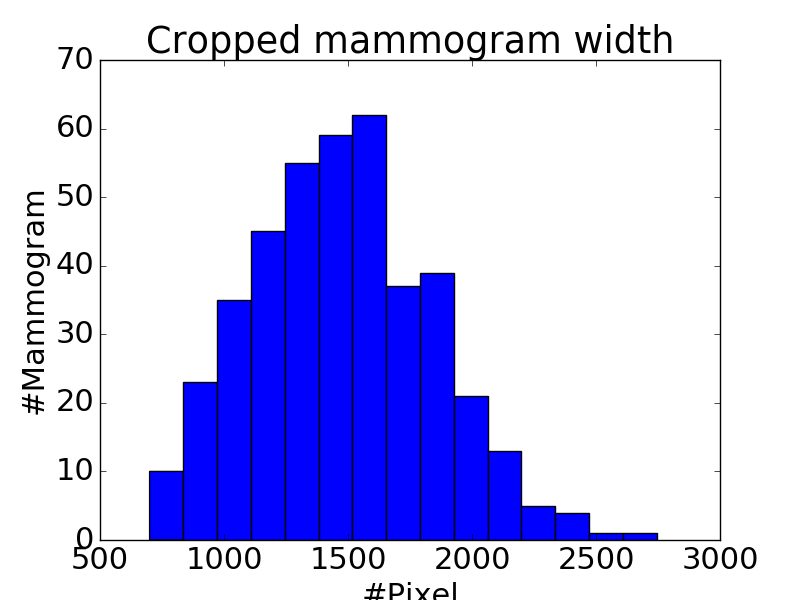
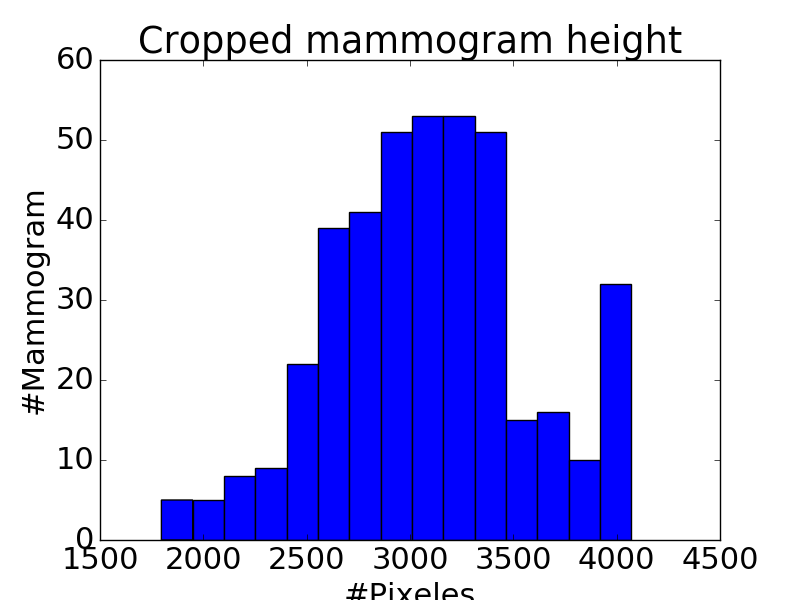
Figure 2: Histograms of mass width (a) and height (b), mammogram width (c) and height (d). Compared to the size of whole mammogram (1,474 \times 3,086 on average after cropping), the mass of average size (329 \times 325) is tiny, and takes about 2\% of a whole mammogram.
Experimental Evaluation
Results on the INbreast Dataset
The proposed networks were evaluated on the INbreast dataset, demonstrating significantly improved performance over traditional techniques reliant on extensive annotations. The deep MIL approaches consistently outperform previous segmentation and detection-based methods, showcasing robust classification capabilities without manual label dependencies.
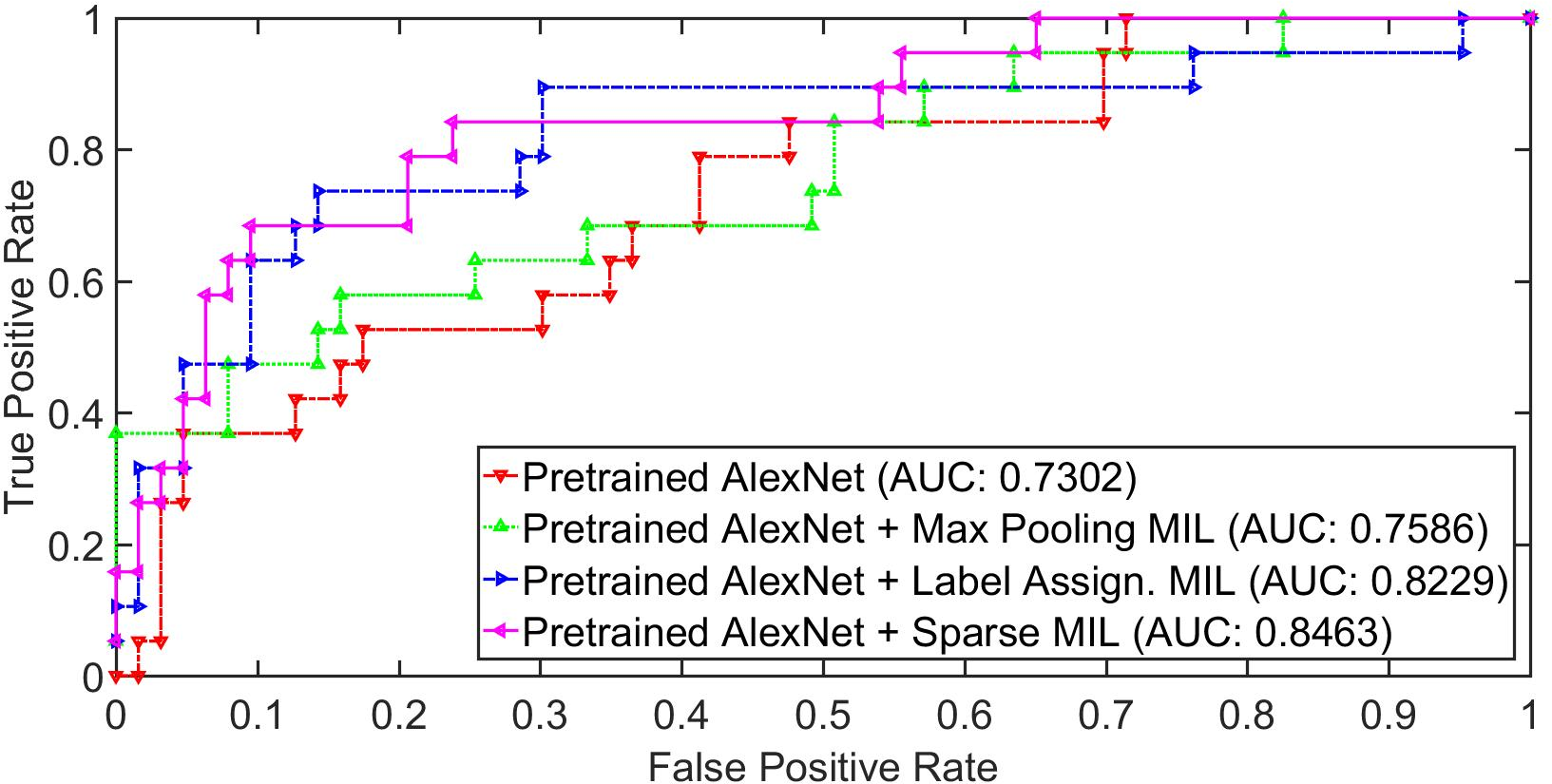
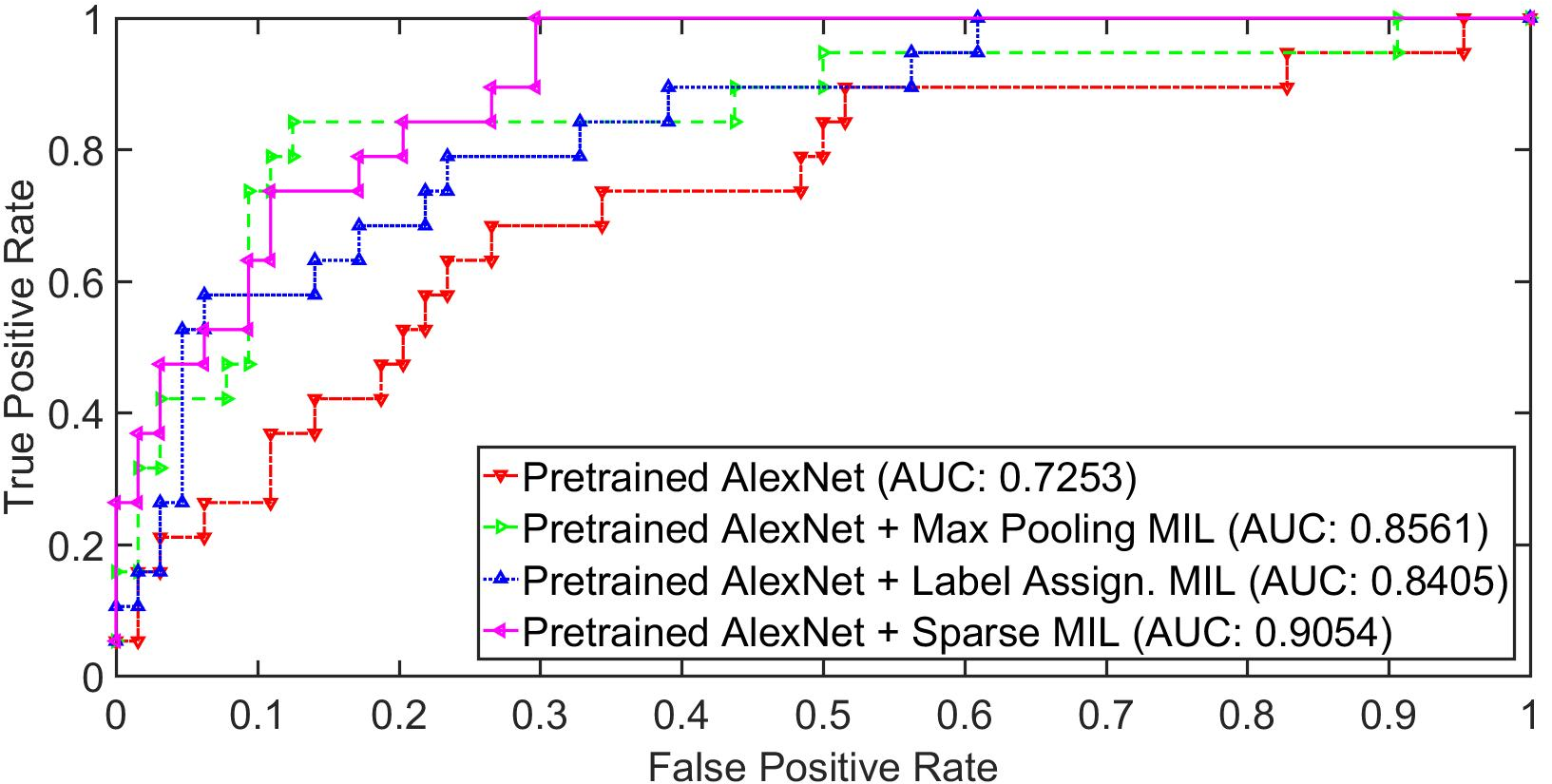
Figure 3: The ROC curve on fold 2 (a) and fold 4 (b) using pretrained AlexNet, pretrained AlexNet with max pooling multi-instance learning, pretrained AlexNet with label assigned multi-instance learning, pretrained AlexNet with sparse multi-instance learning. The proposed deep multi-instance networks improve greatly over the baseline pretrained AlexNet model.
Network Visualization and Insights
Visualization of the predicted malignant probabilities for patches in different mammograms reveals the efficacy of the MIL approaches in identifying mass regions accurately. Notably, while max pooling-based networks may overlook certain malignant patches, label assignment-based and sparse MIL approaches offer improved localization and label consistency.
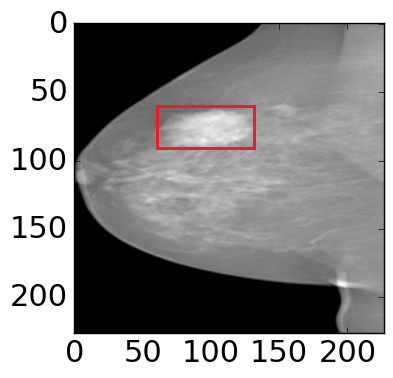
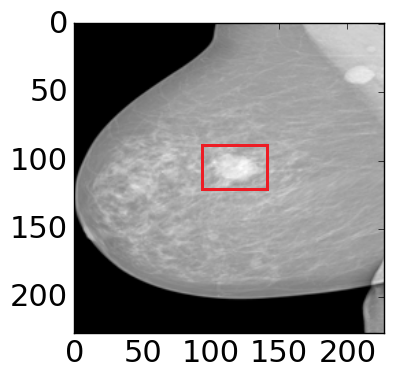
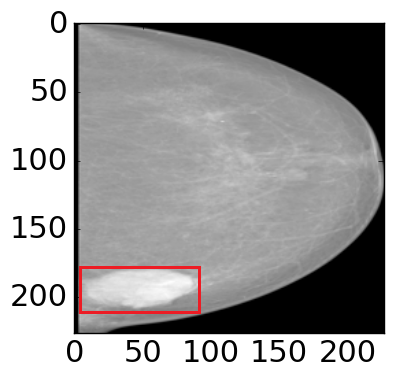
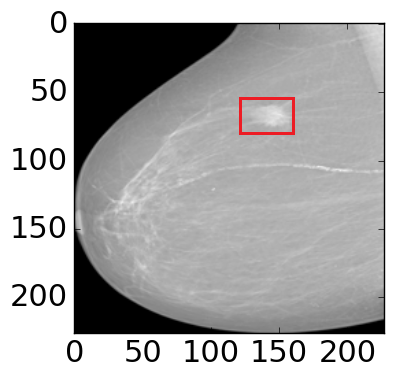
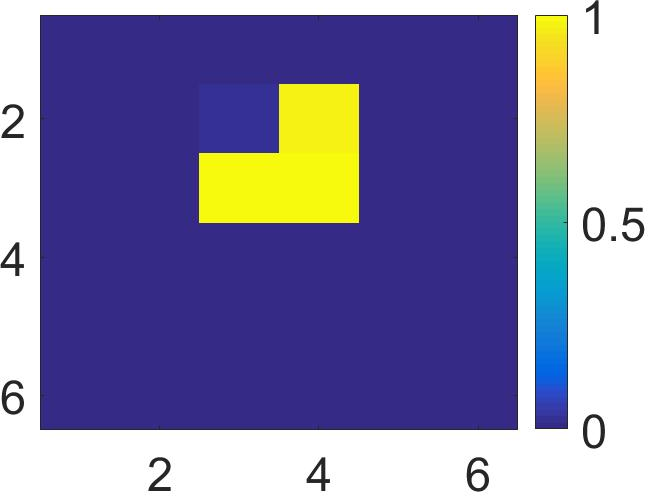
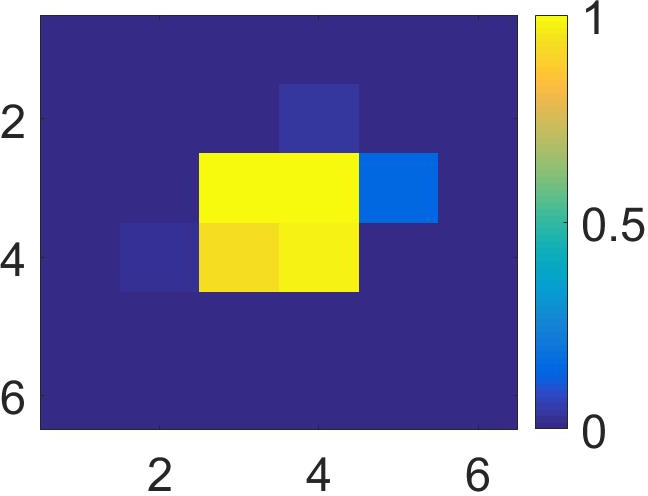
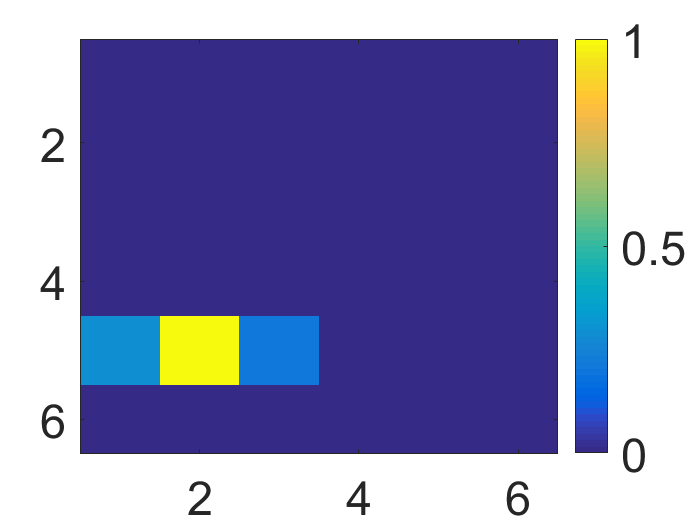
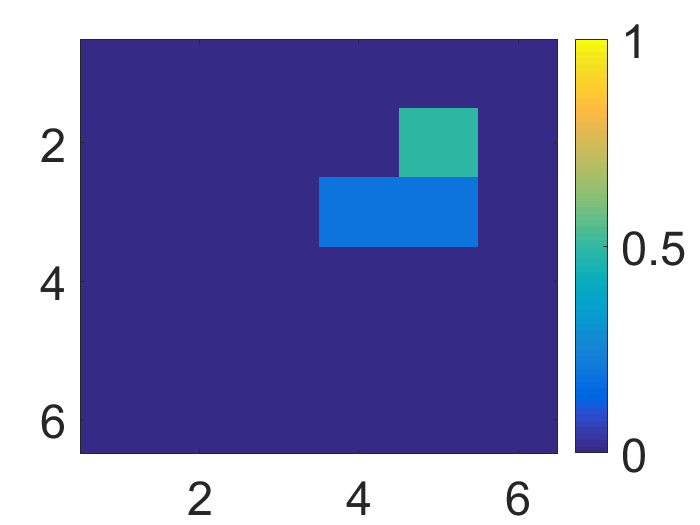
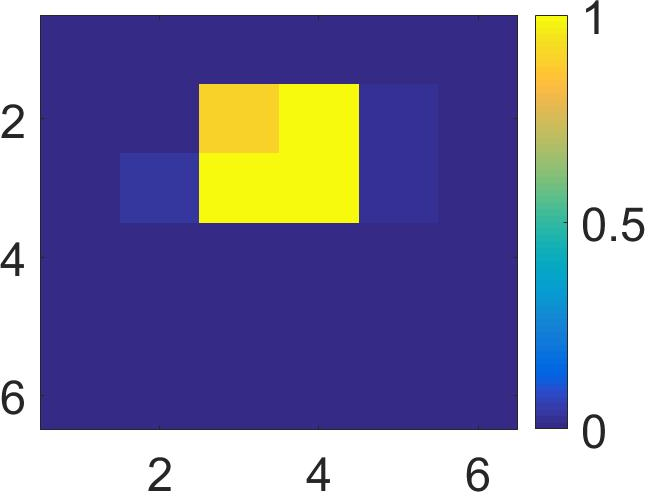
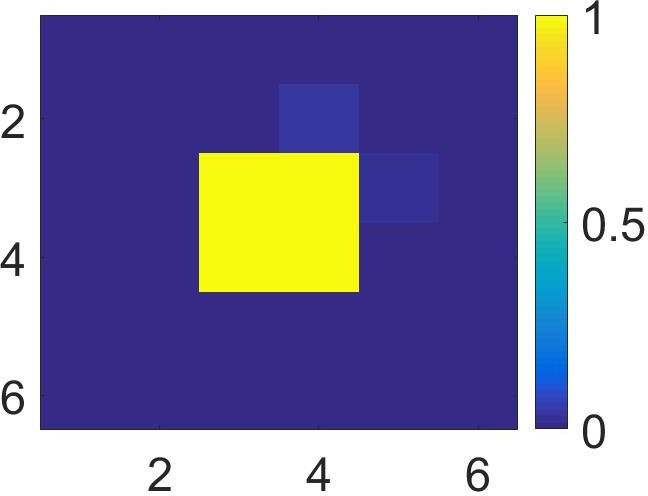
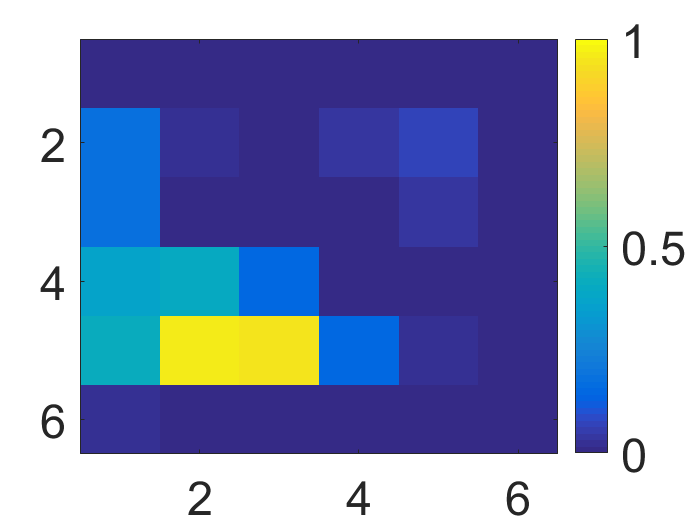
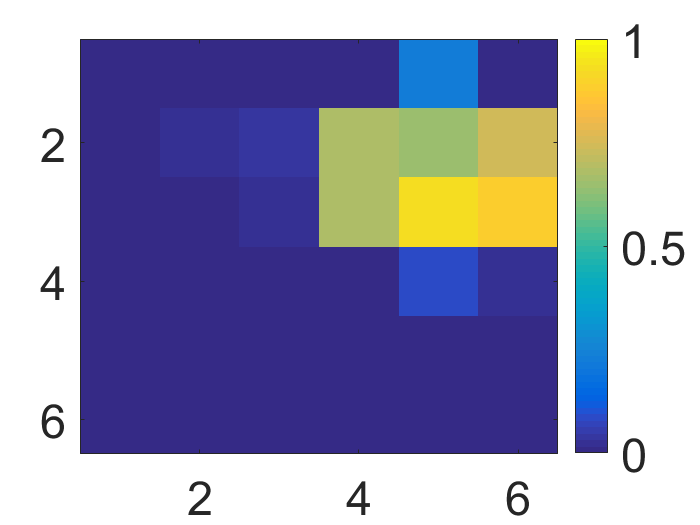
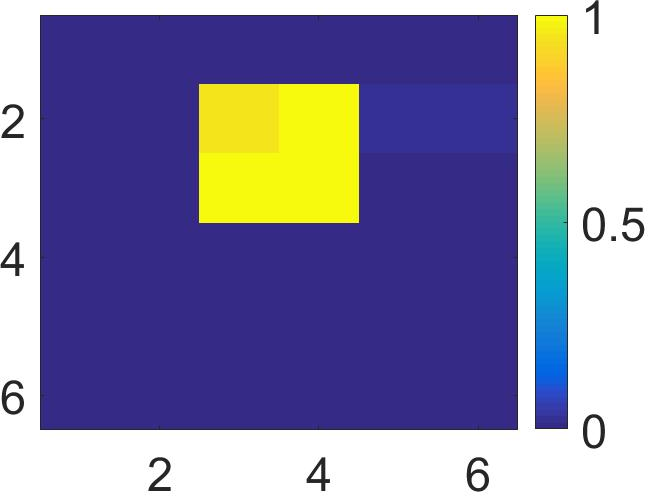
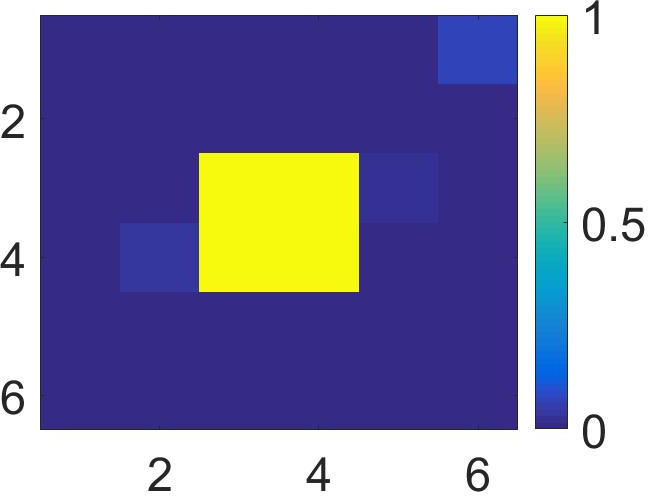
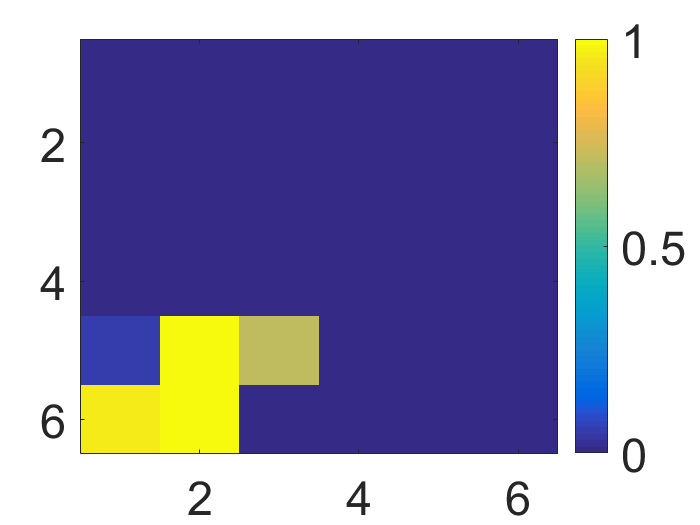
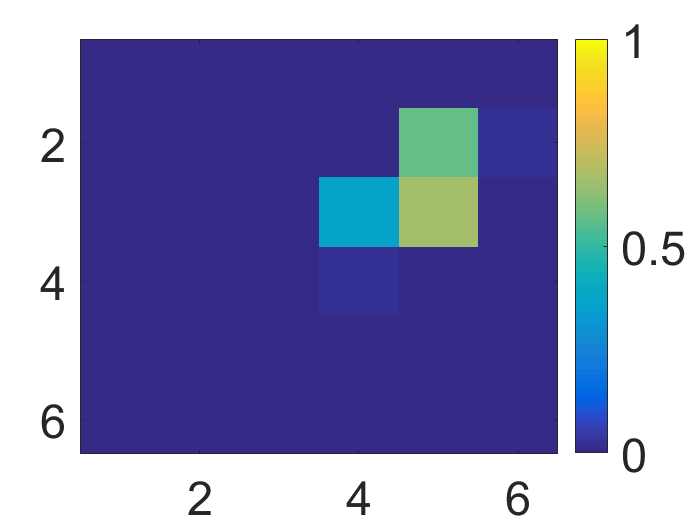
Figure 4: The visualization of predicted malignant probabilities for instances/patches in four different resized mammograms. The first row is the resized mammogram. The red rectangle boxes are mass regions from the annotations on the dataset. The color images from the second row to the last row are the predicted malignant probability from linear regression layer for (a) to (d) respectively, which are the malignant probabilities of patches/instances. Max pooling-based, label assignment-based, sparse deep multi-instance networks are in the second row, third row, fourth row respectively. Max pooling-based deep multi-instance network misses some malignant patch for mammogram (a), (c) and (d). Label assignment-based deep multi-instance network mis-classifies patches into malignant in (d).
Implications and Future Directions
This study introduces a framework that not only improves mammogram classification accuracy and reduces dependency on annotated data but also provides insights for extending MIL to other domains, such as pathology or radiology, where high-quality annotation is scarce. Future research could focus on multi-scale modeling enhancements, adaptive label assignment strategies, and wider applications to other medical imaging problems.
Conclusion
The research contributes a sophisticated MIL framework that significantly elevates mammogram classification performance. It provides a paradigm shift away from annotation-heavy approaches, demonstrating the utility of deep learning in medical image analysis under reduced annotation conditions. This has broad potential applications, especially in fields requiring automated diagnosis without comprehensive pre-labeled data sets.






















