3D Convolutional Neural Networks for Brain Tumor Segmentation: A Comparison of Multi-resolution Architectures
Abstract: This paper analyzes the use of 3D Convolutional Neural Networks for brain tumor segmentation in MR images. We address the problem using three different architectures that combine fine and coarse features to obtain the final segmentation. We compare three different networks that use multi-resolution features in terms of both design and performance and we show that they improve their single-resolution counterparts.
Summary
- The paper demonstrates that multi-resolution 3D CNN architectures significantly improve segmentation accuracy, achieving Dice score gains of over 40 points compared to single-resolution models.
- It compares three architectures—3DNet_1, 3DNet_2, and 3DNet_3—with the two-pathway model (3DNet_3) best integrating local and contextual features despite higher computational costs.
- Key methodological insights include the use of non-uniform patch sampling and strategic fusion of fine and global features to address class imbalance and heterogeneity in brain tumor segmentation.
3D Convolutional Neural Networks for Brain Tumor Segmentation: Multi-resolution Architectures and Comparative Analysis
Introduction and Motivation
Automatic brain tumor segmentation in MRI images is a critical step for diagnosis, treatment planning, and longitudinal assessment in neuro-oncology. The segmentation problem is compounded by substantial class imbalance, fuzziness in tumor borders, and high inter-patient heterogeneity. This study proposes and critically evaluates three distinct 3D fully convolutional neural network (CNN) architectures that incorporate multi-resolution pathways, leveraging both local and contextual features to address the challenge of accurately segmenting gliomas and associated structures from MRI volumes (1705.08236).
Dataset, Sampling, and Training Paradigms
The models are trained and validated on the BRATS dataset, which aggregates multi-modal MR images (T1, T1-contrast, T2, FLAIR) with expert annotations demarcating necrotic core, edema, enhancing core, and non-enhancing tumor regions. Given the severe class imbalance (tumor classes occupy a minor fraction of the volume), the training regimen employs a non-uniform sampling strategy: training patches are sampled such that the central voxel is equally likely to be tumor or background, striking a balance that mitigates prevalent biases inherent in both naive and strictly equiprobable sampling strategies.
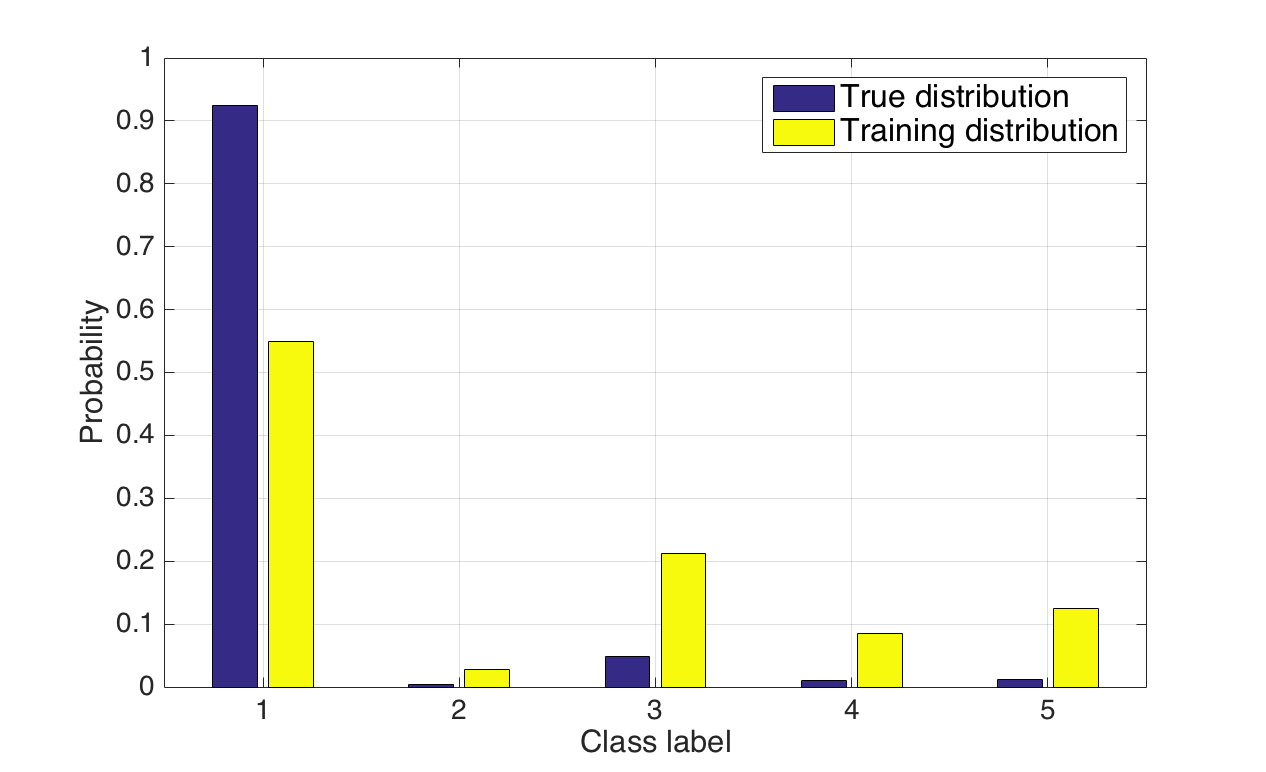
Figure 1: True dataset class distributions (blue) versus the more balanced training patch distribution (yellow) employed to address the foreground-background imbalance problem.
Hardware constraints dictated the use of 643 voxel patches and small batch sizes, with downstream use of hybrid dense-inference to maximize efficiency within GPU memory limitations. All models utilize small convolutional kernels (33) and integrate batch normalization to facilitate stable training in the deep 3D setting.
Multi-resolution 3D CNN Architectures
Three architectures are formulated to systematically probe the effects of fusing contextual and local feature representations:
3DNet_1: VGG-based FCN with Multi-scale Skip Connections
3DNet_1 extends the VGG-FCN paradigm with 3D convolutions. Skip connections are realized via 1×1×1 convolutions at multiple depths, producing class scores at different scales. These are summed and then upsampled to the input’s spatial resolution, ensuring the prediction integrates receptive fields of 403, 923, and 2123 voxels (i.e., hierarchical aggregation of context).

Figure 2: Schematic of the 3DNet_1 architecture demonstrating hierarchical skip connections and concatenated upsampling.
3DNet_2: 3D U-Net with Symmetric Encoder-Decoder Paths
3DNet_2 implements a 3D extension of the canonical U-net. Feature maps from multiple encoder resolutions are concatenated into the decoder at corresponding spatial scales, facilitating fusion of hierarchical features and enabling precise voxel-level segmentation with a maximum receptive field of 1403 voxels.

Figure 3: Schematic of the 3DNet_2 U-net architecture, showing contracting and expanding paths with multi-scale feature concatenations.
3DNet_3: Two-pathway Modified DeepMedic
Inspired by DeepMedic, 3DNet_3 comprises parallel convolutional paths with distinct receptive field sizes: a "local" path without pooling captures fine details (173 receptive field), while a second, deeper path aggregates global context (1363 receptive field). Both outputs are concatenated for the final prediction, allowing for joint exploitation of detailed and contextual cues.

Figure 4: Schematic of the 3DNet_3 two-path architecture, where separate paths capture local and global context and are merged by concatenation.
Experimental Results: Single- vs. Multi-resolution Networks
Ablation studies contrast each multi-resolution architecture with its single-resolution counterpart (i.e., with skip connections or parallel paths disabled). Multi-resolution variants consistently yield superior Dice coefficients, notably for the whole tumor, core, and active regions.
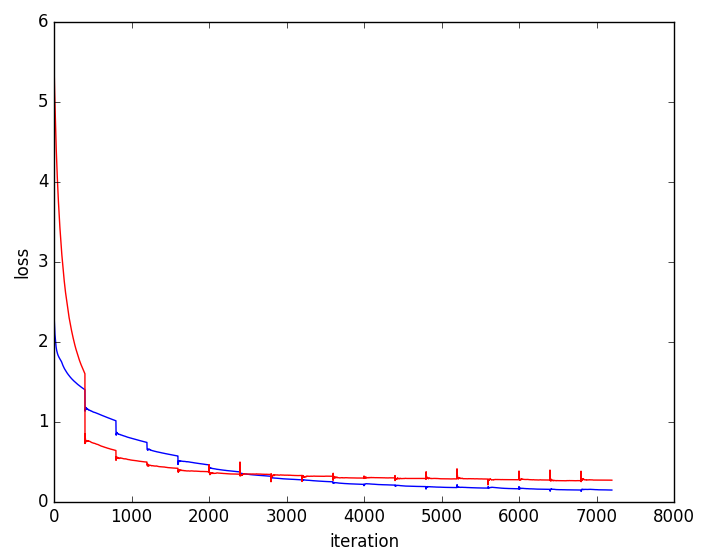
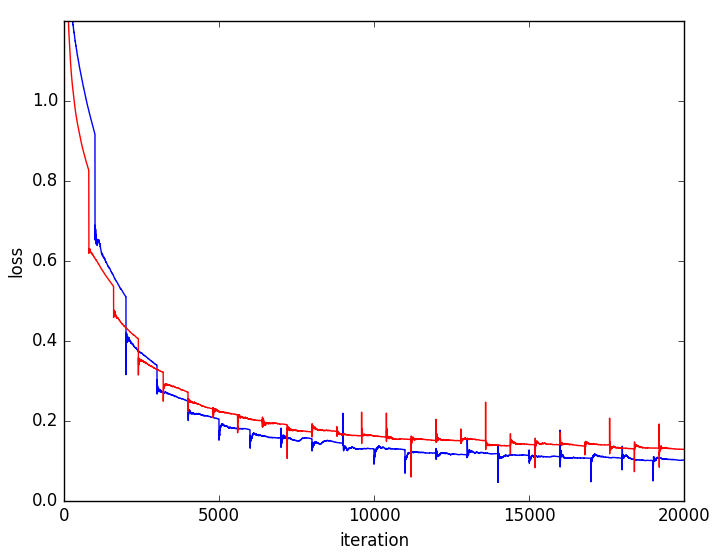
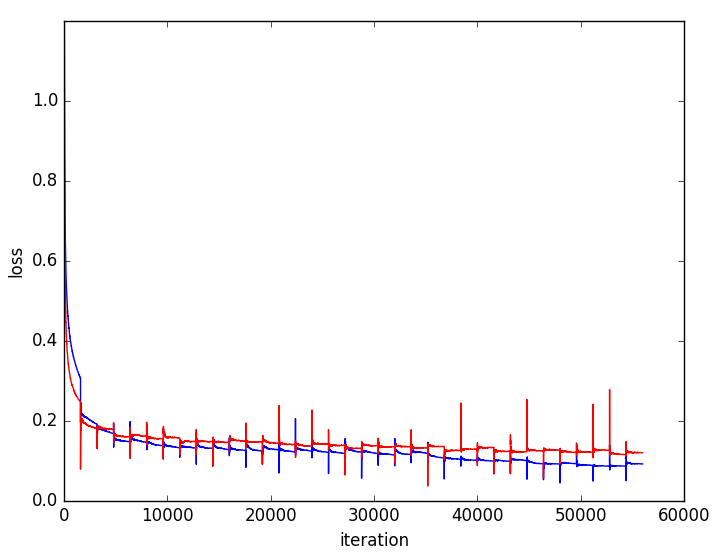
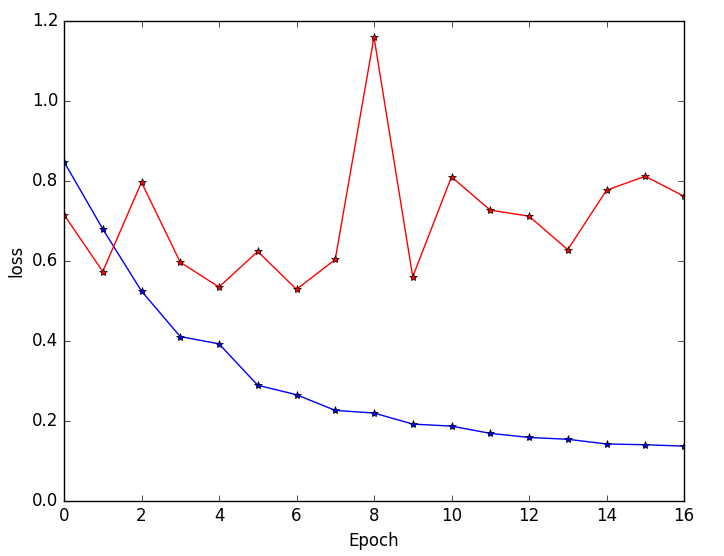
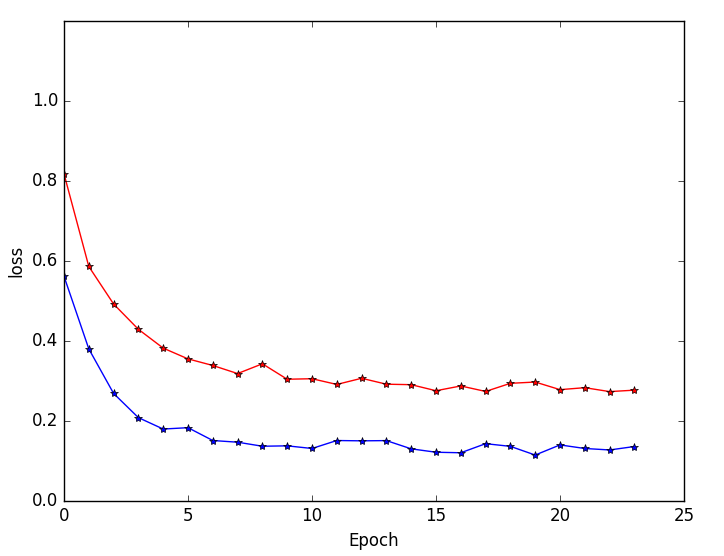
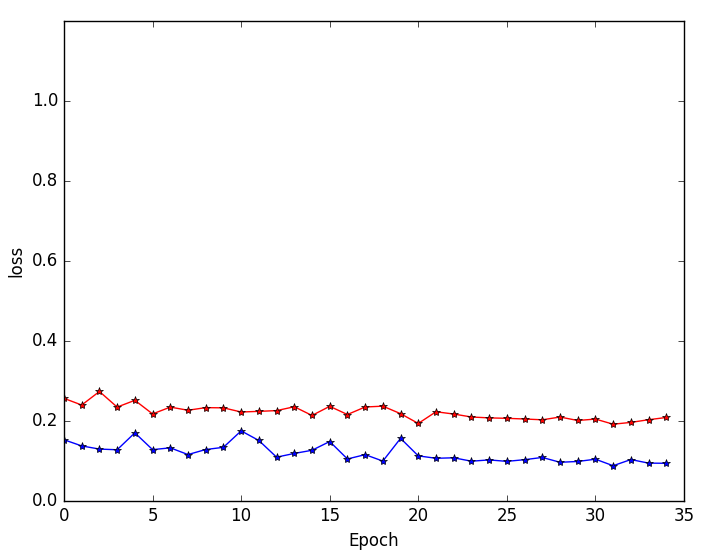
Figure 5: Training and validation loss curves highlighting faster convergence and superior generalization for multi-resolution (blue) versus single-resolution (red) networks.
Single-resolution networks suffer either from inability to localize fine tumor boundaries (when using only coarse/contextual features) or reduced detection recall (with purely local/limited context). For instance, the single-path 3DNet_1 fails to segment tumors even in large regions, while the single-path 3DNet_3 (small receptive field) exhibits finer segmentation but at greater susceptibility to false positives.
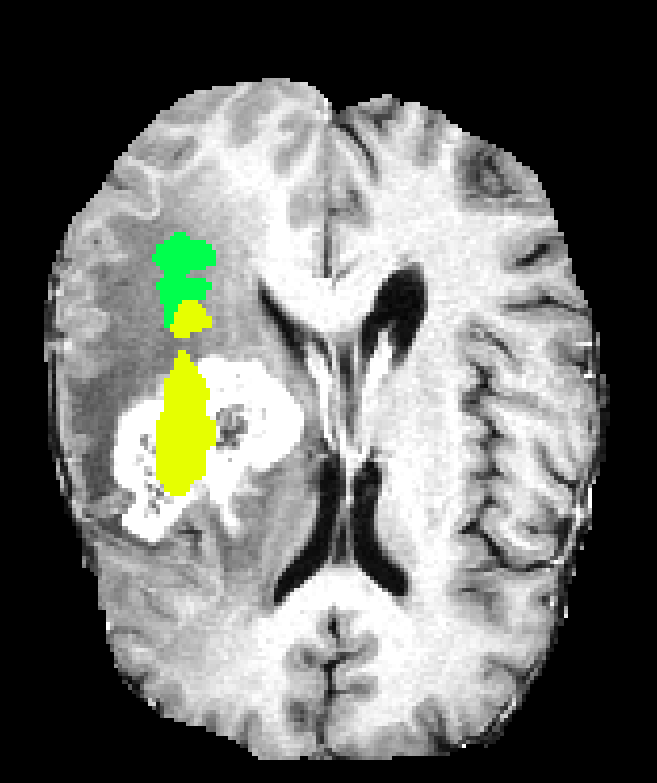
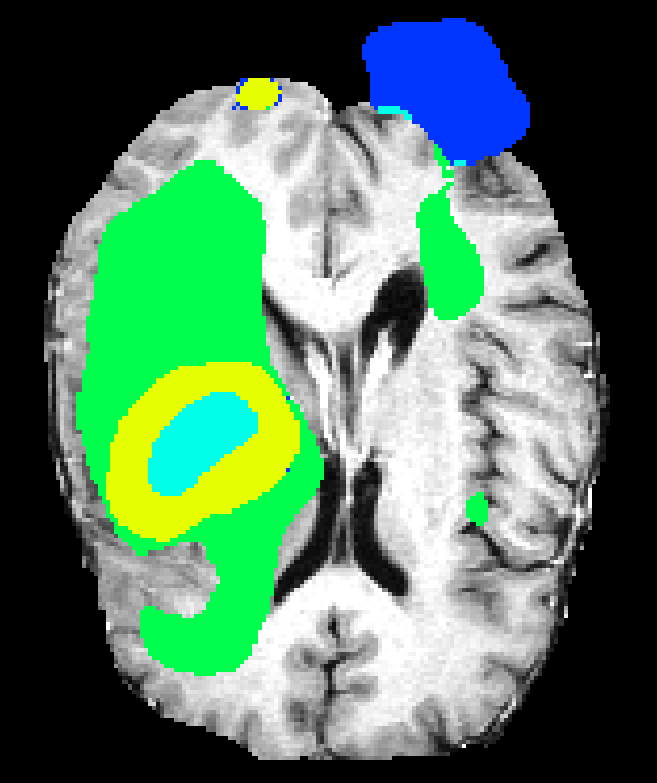
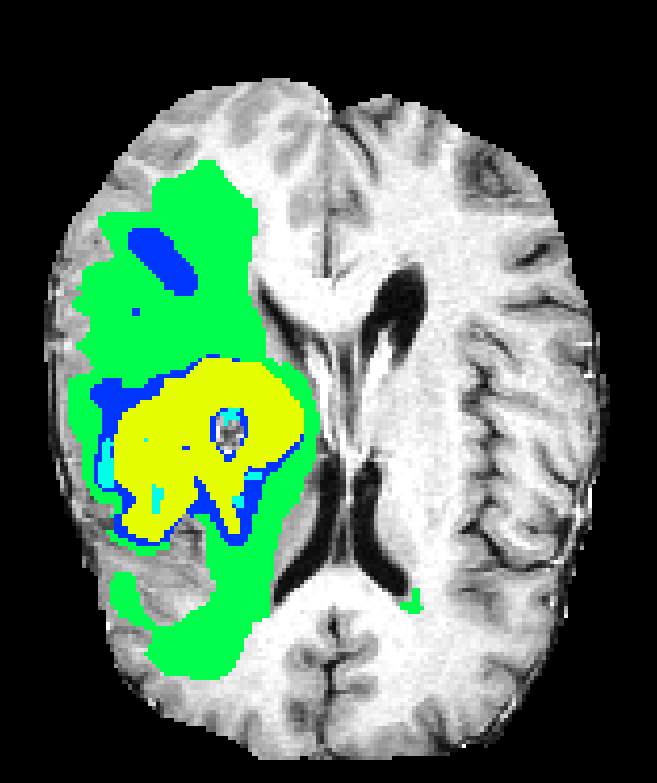
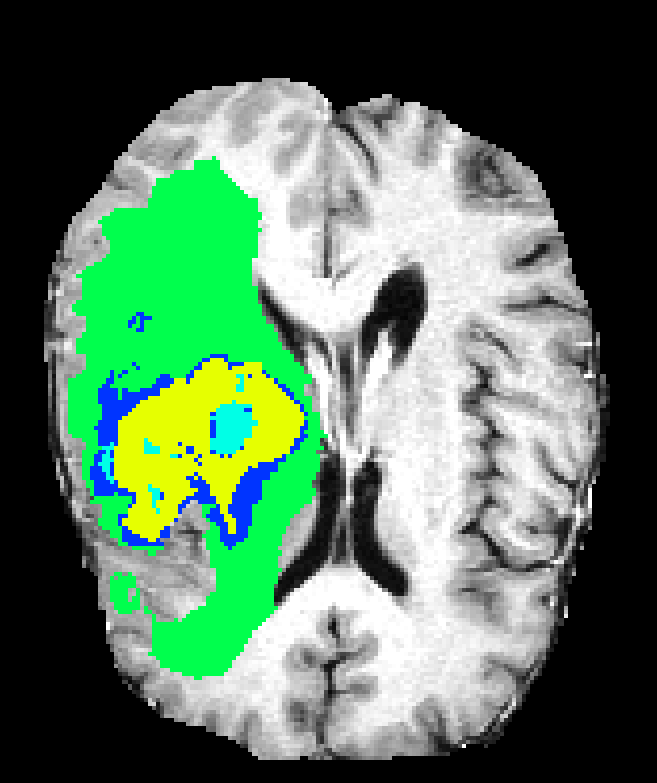
Figure 6: Qualitative segmentations showing intra-tumoral regions across three single-resolution architectures, each affected by different characteristic failure modes.
On validation data, multi-resolution 3DNet_3 achieves Dice scores of 91.7 (whole), 83.6 (core), and 76.8 (enhancing core), while the single-resolution baseline degrades significantly, especially for core and enhancing substructures.
Comparative Performance of Multi-resolution Architectures
When directly compared, all multi-resolution networks significantly outperform their single-resolution precursors. 3DNet_3 attains the highest Dice coefficients, especially for difficult subregions, but at noticeably higher computational cost and GPU memory due to its local path eschewing pooling layers.
While all architectures demonstrate strong discrimination of edema and large tumor masses, challenges remain for highly heterogeneous or small intra-tumoral subregions. 3DNet_2 and 3DNet_1, although computationally more efficient and permitting usage of more filters, lose fine boundary specificity in some cases due to excessive pooling.
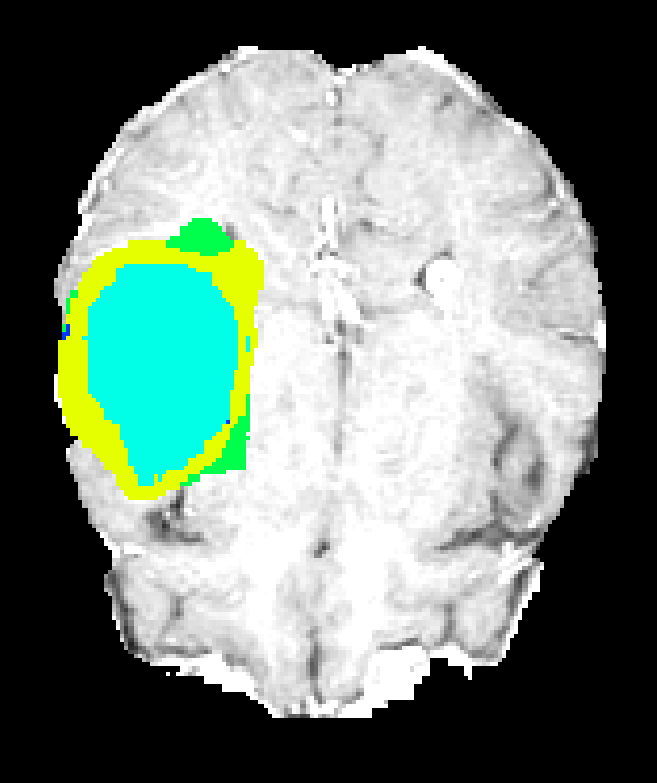
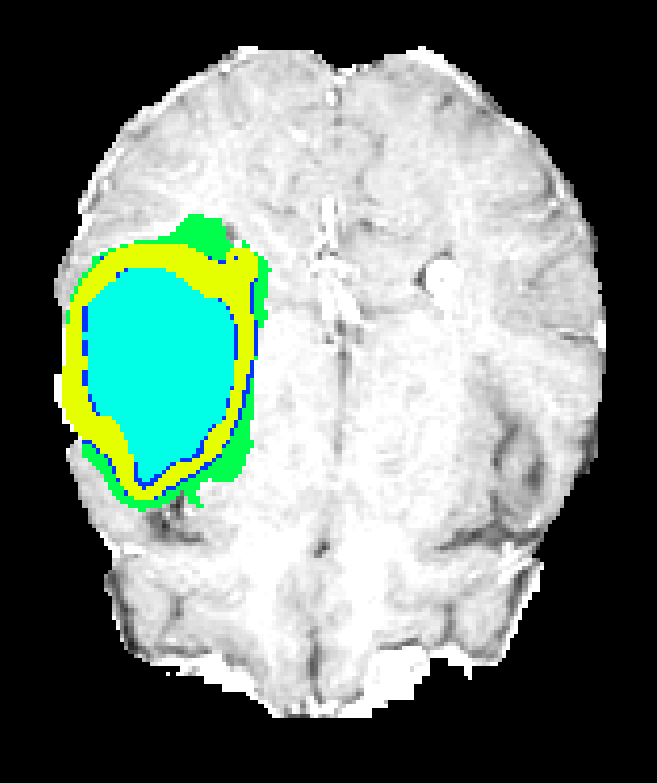
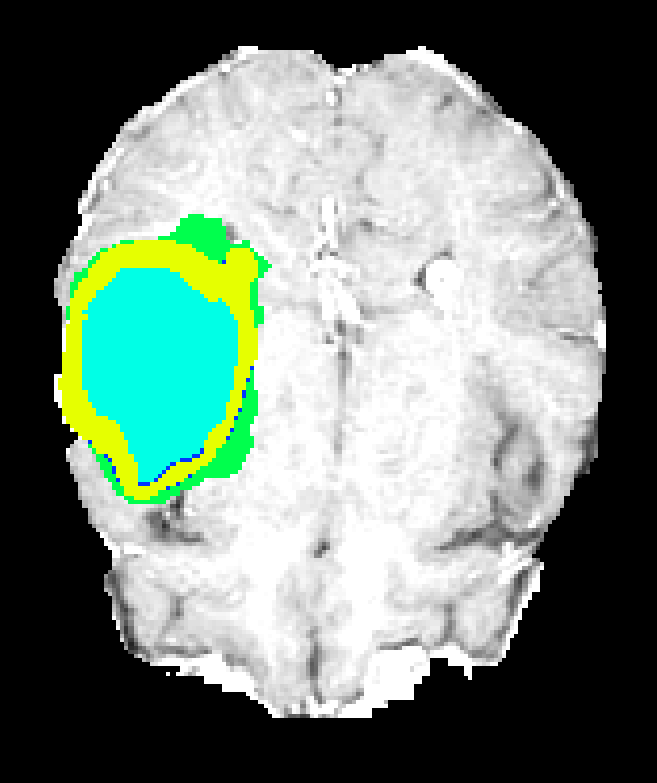
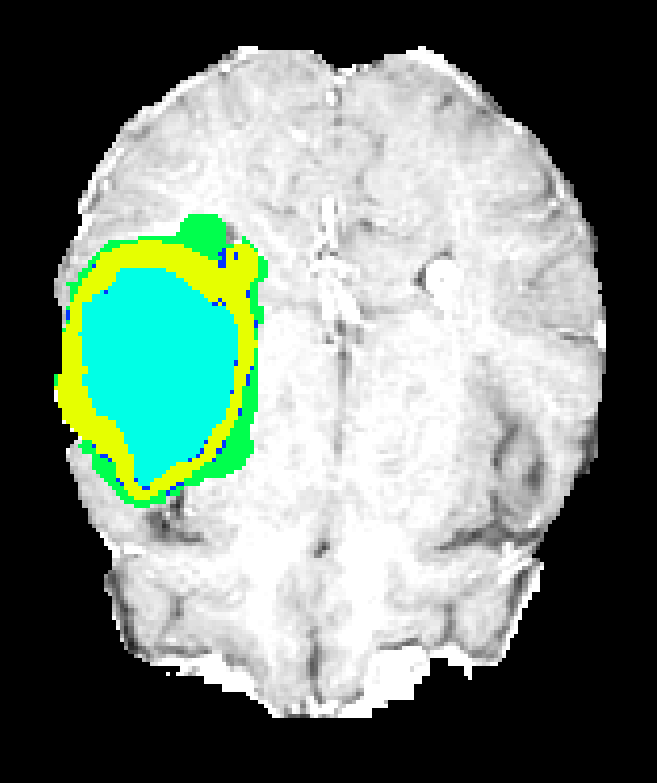
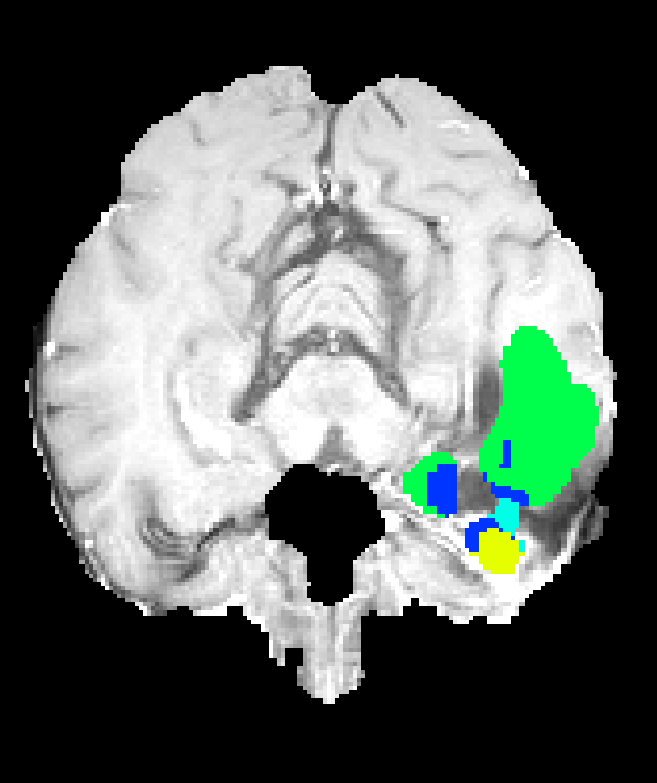
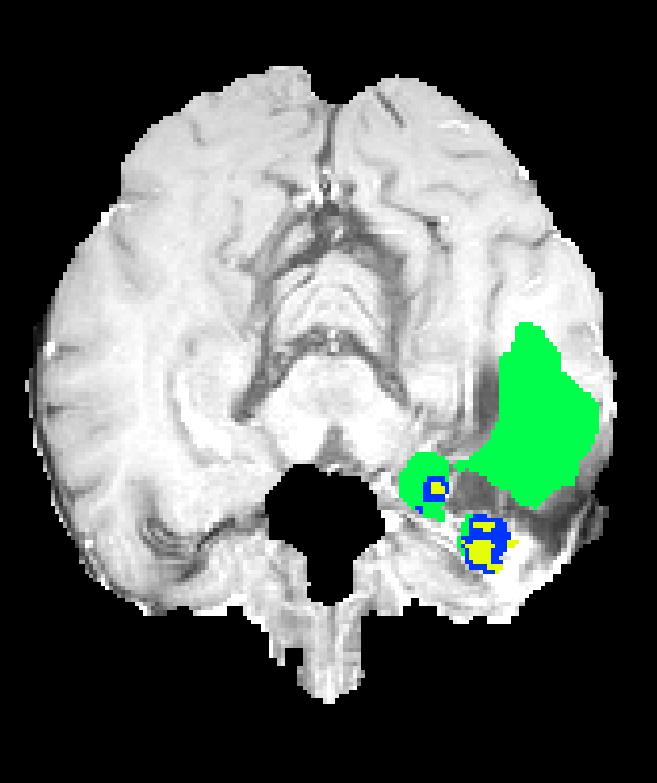
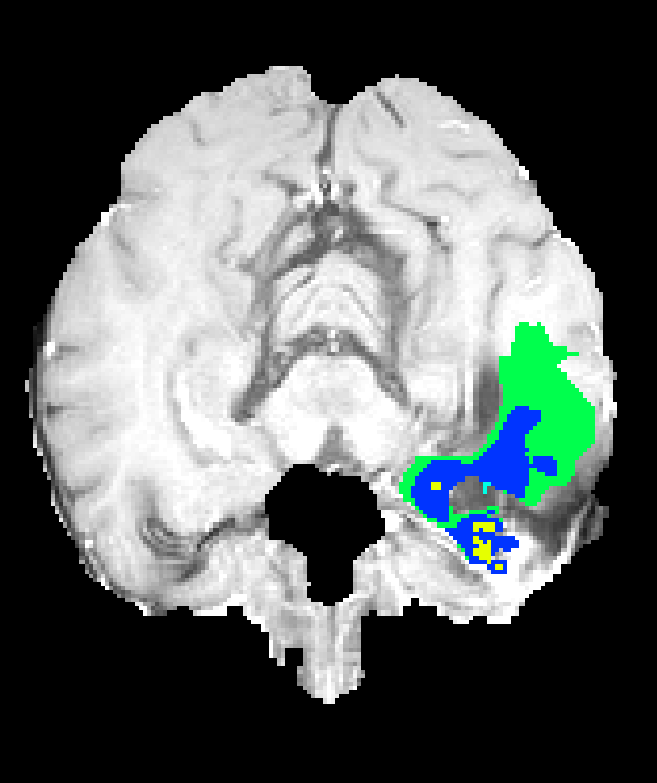
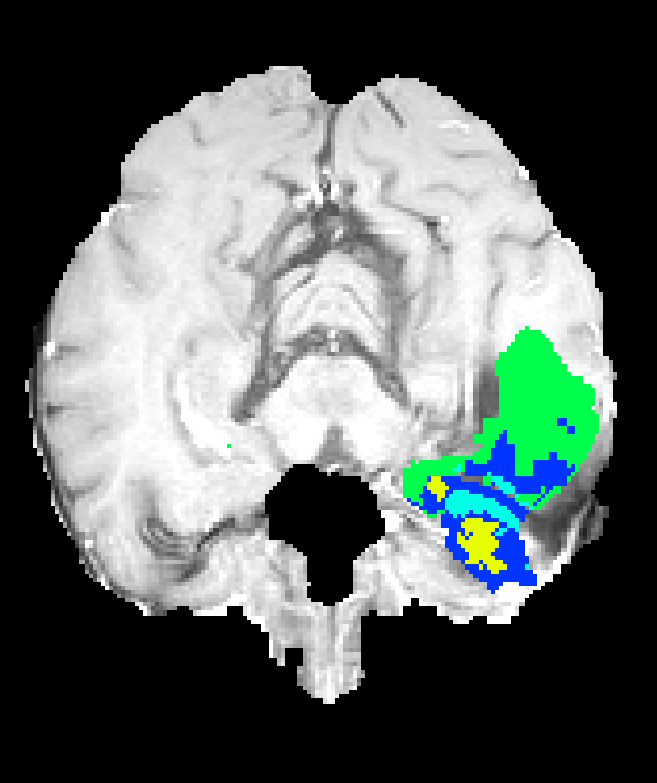
Figure 7: Axial plane qualitative results across architectures. All models localize gross tumor; only models with explicit fine-scale pathways capture high variability within heterogenous regions.
Discussion and Implications
Strong claims in the paper are:
- All multi-resolution 3D CNNs tested greatly outperform their single-resolution counterparts across all clinical targets (Dice improvement on whole tumor: >40 points).
- 3DNet_3, despite having the fewest parameters due to hardware constraints, yields the best Dice score on challenging subregions, indicating architectural inductive bias (local path without pooling) can outweigh raw depth or parameter count.
- Memory and computational demands form the chief bottleneck for 3D models, necessitating carefully engineered input, batch size, and parameterization trade-offs.
Practical implications: Deploying multi-resolution 3D CNNs yields robust segmentations suitable for integration into clinical workflows. However, inference speed and memory footprint are limiting factors for real-time application or high-throughput analysis unless additional model pruning, quantization, or distributed inference strategies are adopted.
Theoretical implications: The analysis supports the critical requirement for explicit multi-scale feature fusion, rather than reliance on receptive field growth alone, in volumetric segmentation. Furthermore, the results suggest that complementing deep contextual features with unpooled, high-resolution analysis is advantageous, especially in the context of highly variable pathologies.
Future developments: Anticipated directions include end-to-end differentiable post-processing (e.g., CRFs as RNNs), hybrid explicit attention mechanisms to further adaptively weigh context versus local structure, and architecture search approaches for maximizing accuracy within strict hardware constraints. Semi-supervised and transfer learning approaches could further reduce annotation requirements for rare tumor subtypes.
Conclusion
The study provides a comprehensive evaluation of multi-resolution 3D CNN architectures for brain tumor segmentation in MRI, empirically demonstrating that integrating both local and contextual features via parallel paths or skip connections is essential for state-of-the-art performance. Among the tested designs, those incorporating explicit high-resolution pathways without pooling (3DNet_3) exhibit the best efficacy on the BRATS dataset, particularly for subregion segmentation. The findings have tangible implications for computational neuro-oncology and offer clear guidance for architectural choices in future volumetric medical image analysis systems.
(1705.08236)
Paper to Video (Beta)
No one has generated a video about this paper yet.
Whiteboard
No one has generated a whiteboard explanation for this paper yet.
Paper Prompts
Sign up for free to create and run prompts on this paper using GPT-5.
Top Community Prompts
Open Problems
We haven't generated a list of open problems mentioned in this paper yet.
Continue Learning
- How does multi-resolution feature integration contribute to improved segmentation performance in 3D CNNs?
- What role does non-uniform sampling play in mitigating class imbalance in this study?
- How do the receptive field sizes of the different architectures affect their segmentation outcomes?
- What are the computational challenges and trade-offs observed with the 3DNet_3 two-pathway model?
- Find recent papers about multi-resolution medical image segmentation.
Related Papers
- Identifying the Best Machine Learning Algorithms for Brain Tumor Segmentation, Progression Assessment, and Overall Survival Prediction in the BRATS Challenge (2018)
- Cascaded V-Net using ROI masks for brain tumor segmentation (2018)
- Deep Learning Enables Automatic Detection and Segmentation of Brain Metastases on Multi-Sequence MRI (2019)
- High-Resolution Breast Cancer Screening with Multi-View Deep Convolutional Neural Networks (2017)
- An interpretable classifier for high-resolution breast cancer screening images utilizing weakly supervised localization (2020)
- Predicting Alzheimer's disease: a neuroimaging study with 3D convolutional neural networks (2015)
- Multi-resolution Super Learner for Voxel-wise Classification of Prostate Cancer Using Multi-parametric MRI (2020)
- A review: Deep learning for medical image segmentation using multi-modality fusion (2020)
- Fast and Accurate 3D Medical Image Segmentation with Data-swapping Method (2018)
- Deep Neural Networks for Anatomical Brain Segmentation (2015)
Authors (4)
Collections
Sign up for free to add this paper to one or more collections.