- The paper introduces a novel deep normative modeling framework that leverages Neural Processes for mixed-effect modeling, addressing the scalability and kernel limitations of traditional Gaussian Process methods.
- It integrates 3D-CNN layers within a Tensor Gaussian Predictive Process to capture spatial correlations and estimate predictive uncertainties in MRI data.
- The approach significantly improves diagnostic accuracy for ADHD and reduces computational complexity from cubic to linear with increasing sample sizes.
Neural Processes Mixed-Effect Models for Deep Normative Modeling of Clinical Neuroimaging Data
Introduction
The paper introduces a novel framework for deep normative modeling based on Neural Processes (NPs) to address limitations of Gaussian Process Regression (GPR) in analyzing neuroimaging data. Normative modeling, which maps biological and clinical covariates, can reveal biomarkers of psychiatric disorders; however, GPR's scalability issues and fixed kernel requirements hinder its application on large datasets with spatially correlated neuroimaging data. Neural processes offer a solution by implementing mixed-effect models that capture feature representations and covariance structures in neuroimaging through stochastic processes.
Methods
Mixed-Effect Modeling of MRIs
The framework applies NPs within the Tensor Gaussian Predictive Process (TGPP), defining a mixed-effect model for neuroimaging. It includes fixed-effects modeled linearly and random-effects accounting for spatial variations and noise. The model estimates predictive uncertainty by sampling global latent variables, enhancing the flexibility and scalability of normative modeling.
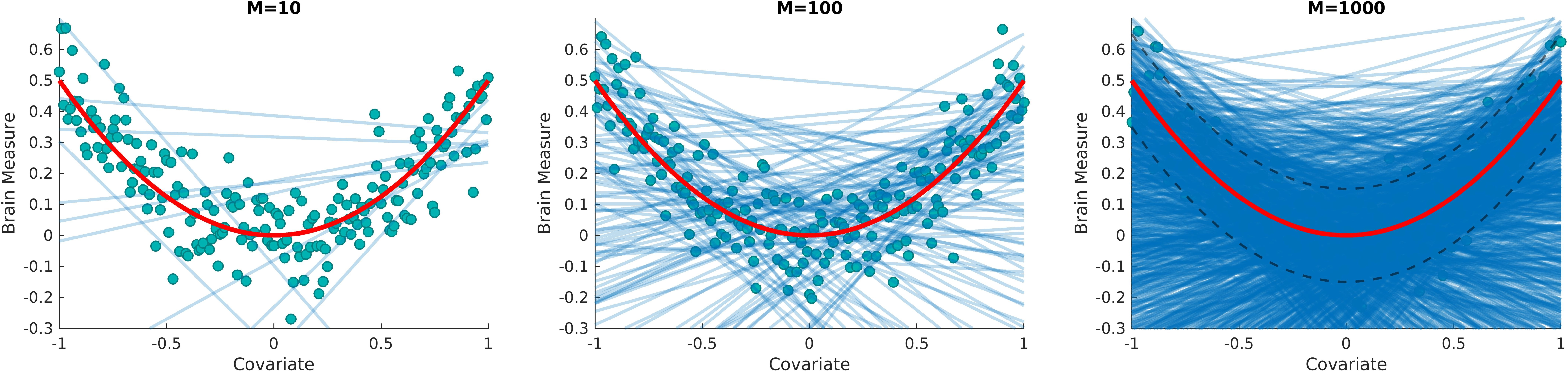
Figure 1: A schematic illustration on approximating the distribution of a non-linear target function (red curve) from the distribution of linear context functions (blue lines).
Neural Processes for MRI Analysis
Neural Processes are employed to learn distributions over functions rather than individual data points, benefiting modeling of multivariate normative data. The approach utilizes 3D-CNN layers to preserve spatial structures in MRI data. Encoder and decoder networks facilitate extracting and reconstructing neuroimaging features, with latent variables encoding covariance properties crucial for statistical map derivations.
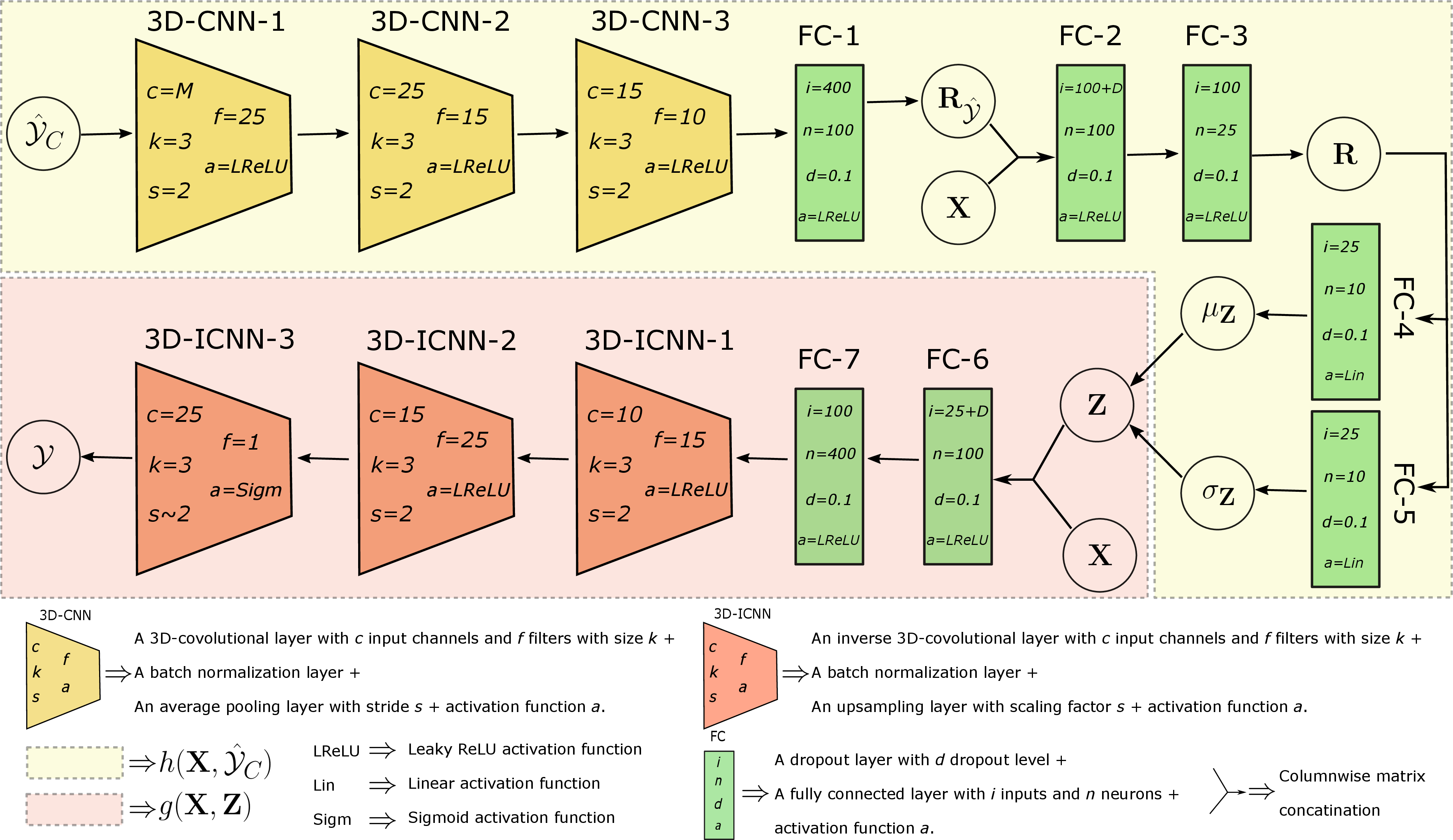
Figure 2: An example NP architecture for mixed-effect modeling of MRIs.
Implementation Strategy
The methodology involves encoding MRI data and covariates using convolutional neural architectures to estimate normative models, which can then be used to predict psychiatric disorders in an unsupervised setting. The generative model calculates aleatoric and epistemic uncertainties as part of normative probability maps, enabling further delineation of disease-specific deviations.
Results and Analysis
Neural processes demonstrated significant improvements over multi-task Gaussian processes in identifying ADHD patients from a clinical fMRI dataset, particularly enhancing detection accuracy in unsupervised diagnostic settings. The precision in mapping deviations and associations with impulsivity across different anatomical brain areas underlines the potential of the NP approach in practical applications.
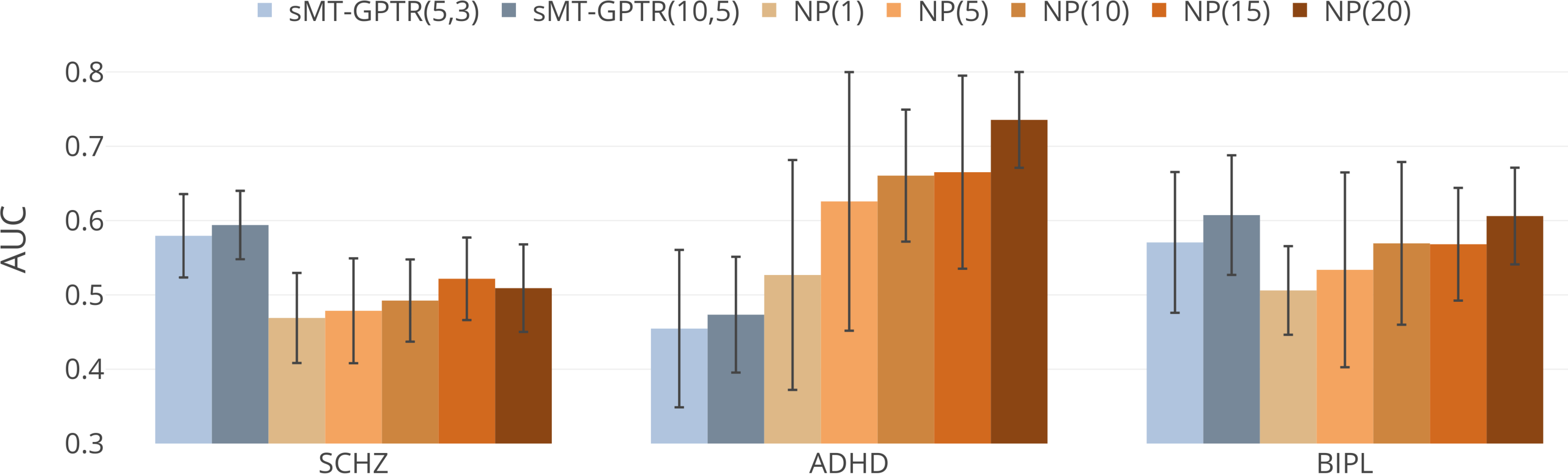
Figure 3: Comparison between novelty detection performances of normative models derived by sMT-GPTR and NPs.
Discussion
Computational Efficiency
Neural Processes offer computational advantages through scalable modeling, reducing complexity from cubic to linear concerning sample sizes. This aligns well with the needs for large cohort analyses, especially in high-dimensional neuroimaging contexts where traditional methods falter.
Implications and Future Directions
The extension of context/target points to functions has opened new paths for mixed-effect modeling, allowing considerations of structured variations in spatial data. The paper advocates for the potential expansion of NP applications beyond neuroimaging, affecting varied domains like genomics and epidemiology.
Conclusion
The application of Neural Processes to mixed-effect models in neuroimaging establishes a new pathway for deep normative modeling, effectively improving diagnostic predictions while maintaining computational feasibility for large datasets. The approach serves as a bridge to wider applicability across health-related and neuroscientific research, suggesting further explorations into larger and more complex clinical datasets.


