- The paper presents a deep learning model that integrates genotype and environmental data to improve crop yield prediction accuracy.
- It employs a dual-DNN architecture with 21 hidden layers, using dropout, batch normalization, and residual shortcuts to capture non-linear interactions.
- The approach outperforms traditional methods, though challenges remain with the 'black box' nature of deep neural networks for biological interpretation.
Crop Yield Prediction Using Deep Neural Networks
The paper "Crop Yield Prediction Using Deep Neural Networks" presents a sophisticated approach to predicting crop yields using deep learning algorithms. This research addresses the challenge of accurately predicting crop yields by leveraging extensive datasets and state-of-the-art modeling techniques. The model's superior accuracy compared to traditional methods is highlighted by its performance in the 2018 Syngenta Crop Challenge.
Introduction
Crop yield prediction involves understanding the complex interactions between genotype (G), environment (E), and their interactions (G×E). Traditional approaches often simplify these interactions by considering only additive effects, which may overlook critical dynamics in genotype-environment interactions. Machine learning, notably deep neural networks (DNNs), is well-suited for capturing complex, non-linear relationships, as it models the yield as an implicit function of genotype and environmental factors.
Methodology
Data Preprocessing
The research utilized data from the 2018 Syngenta Crop Challenge, including genotype data, environmental data comprising soil and weather variables, and yield performance records. Effective preprocessing of genotype data involved reductions and imputations that maintained predictive accuracy.
Weather Prediction
Weather plays a vital role in yield prediction. The authors implemented neural networks to predict weather variables due to the presence of inherent non-linearities in weather data. The ability of neural networks to learn these non-linear dynamics without predefined models was crucial.
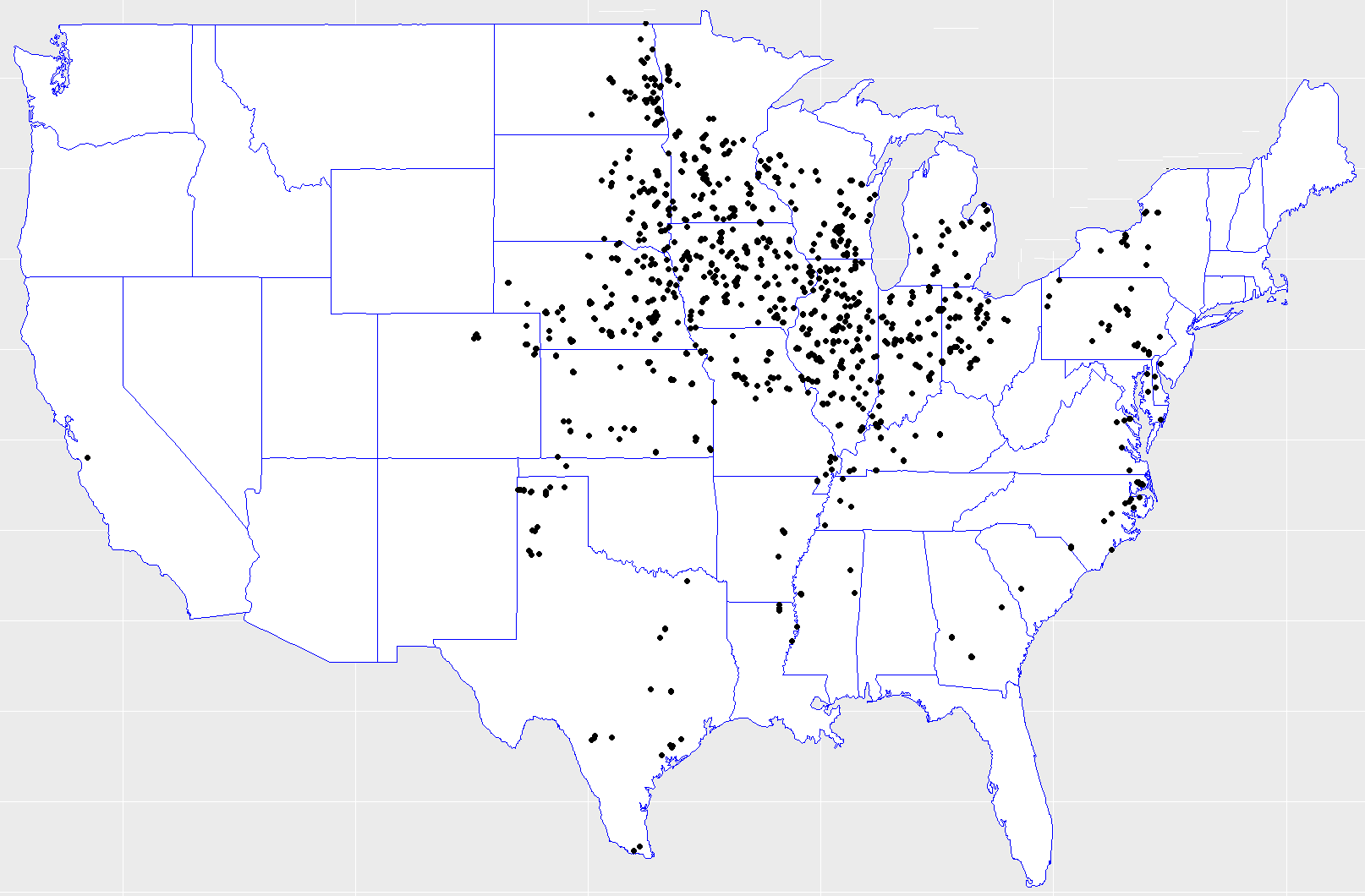
Figure 1: Hybrids locations across the United States. Data collected from the 2018 Syngenta Crop Challenge.
Deep Neural Networks for Yield Prediction
The core of the prediction model involved training two separate DNNs for yield and check yield prediction, respectively, and utilizing the yield difference as a derived measure. The architecture included 21 hidden layers with 50 neurons each, utilizing advanced techniques like batch normalization, dropout, and residual shortcuts to mitigate issues like vanishing gradients.
(Figure 2)
Figure 2: Deep neural network structure for yield or check yield prediction, illustrating the data flow and connectivity.
Results and Analysis
The DNN model demonstrated enhanced performance over traditional models such as Lasso, shallow neural networks, and regression trees, particularly excelling in predicting yield and check yield with the RMSE being approximately 11% of their mean values. The model, however, faced challenges with accurately predicting yield differences due to higher variance in this metric.
Additionally, the study conducted feature selection using guided backpropagation to identify important variables within genotype and environmental datasets. This method successfully reduced input space dimensionality without significant prediction loss.
(Figure 3)
Figure 3: Bar plot of estimated effects of soil conditions, highlighting the relative importance of different soil factors.
Conclusion
The paper demonstrates the efficacy of deep learning methods for crop yield prediction, emphasizing the model's ability to handle complex interactions between genotype and environmental factors. The influence of accurate weather predictions on model performance underscores the necessity of integrating improved weather forecasting into crop yield models.
Challenges remain, primarily the "black box" nature of DNNs, which limits direct biological inference. Future research should aim to develop more interpretable models that retain high predictive accuracy while offering deeper insights into genotype-environment interactions.
