- The paper demonstrates that supervised radiomic feature selection enhances lung nodule classification, achieving top SVM accuracy with 8 features.
- It compares supervised and unsupervised ranking methods applied to shape and texture features extracted from CT images.
- The study shows that optimal feature subsets can outperform larger sets, improving classification efficiency and accuracy.
Radiomic Feature Selection for Lung Cancer Classifiers
Introduction
The paper "Radiomic feature selection for lung cancer classifiers" (2003.07098) investigates the role of radiomic features in the classification of lung nodules using machine learning algorithms. This study addresses the gap in literature regarding effective feature ranking and the optimal number of features required for nodule classification. The authors explore the impact of supervised and unsupervised feature selection techniques on classifiers, emphasizing Support Vector Machines (SVM) and Naive Bayes algorithms.
Methodology
The workflow of the experimental setup involves several critical steps beginning with the segmentation of lung nodules from CT imaging data, followed by radiomic feature extraction. Supervised and unsupervised feature ranking algorithms are applied to select the most discriminative features, which are then utilized to train machine learning classifiers for the purpose of nodule classification.

Figure 1: Work flow of the proposed experimental setup.
Radiomic features, including those related to shape and texture, are computed using the PyRadiomics module. Features are classified into various categories such as GLDM, GLCM, and GLSZM. A pre-processing step involves one-way ANOVA tests to identify discriminative features which are ranked using both supervised and unsupervised methods.
Classification Models
The study utilizes SVM and Naive Bayes classifiers to categorize nodules as benign or malignant. SVM operates through constructing a hyper-plane that best separates the classes, while Naive Bayes applies probabilistic reasoning grounded in Bayes' theorem. Both classifiers are evaluated based on accuracy, specificity, and sensitivity using cross-validation techniques.
Figure 2: Segmentation of a malignant nodule from Lung1 database in (a) 2-D axial slice (b) 3-D axial slice.
Feature Selection Techniques
Feature selection is performed using supervised methods such as Fisher Score, ReliefF, and fNCA, which consider class labels for ranking. Unsupervised methods like minimum correlation, Laplacian Score, and MCFS focus on intrinsic feature characteristics without reference to class labels. The paper provides a detailed comparison of these approaches and emphasizes that supervised feature selection generally yields superior classification outcomes.
Figure 3: Feature extraction and Feature reduction with respect to feature class.
Results
The findings of the study include exceptional classification performance with the highest accuracy achieved using SVM trained with 8 features from the supervised ranking approach. The paper notes that increasing the number of features from 2 to 20 does not always result in improved performance, highlighting that fewer features can sometimes suffice for accurate classification.
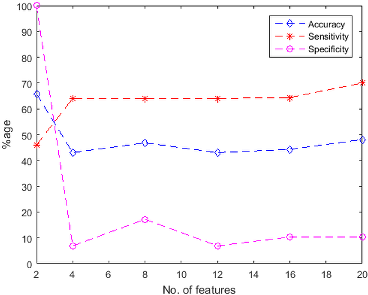
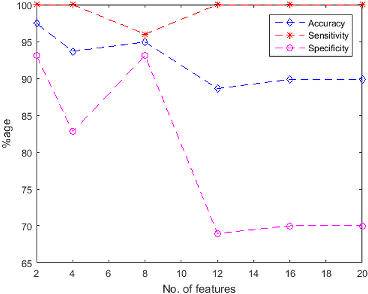
Figure 4: Naive Bayes Classification performance (a) using unsupervised feature ranking (b) using supervised feature ranking.
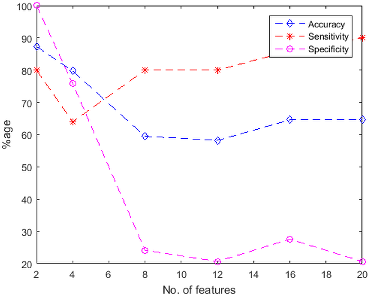
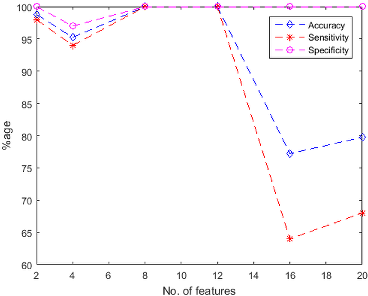
Figure 5: SVM Classification performance (a) using unsupervised feature ranking (b) using supervised feature ranking.
Discussion
The experimental results demonstrate the advantages of supervised feature selection techniques in radiomic analysis. SVM's performance notably surpasses that of Naive Bayes in most scenarios, reaffirming its suitability for complex classification tasks. Conclusions highlight that a careful selection of fewer features can lead to significant classification accuracy, questioning the conventional approach of utilizing larger feature sets.
Conclusion
The study makes significant contributions to radiomic feature selection in lung cancer classifiers, providing insights into optimal feature selection and its critical impact on the performance of machine learning models. Future research may extend these findings by exploring other types of classifiers or varying the number of radiomic features beyond the scope of this study.
These results suggest that optimized feature selection methods, especially supervised ones, can enhance the predictive capabilities of CAD systems used in clinical settings. The implications for future cancer diagnostics are profound, potentially enabling more accurate and early detection of lung cancer through advanced computational techniques.