- The paper presents a novel bidirectional unpaired image translation framework using cycleGAN combined with structural similarity loss to enhance diagnostic image quality.
- It employs ResNet-based generators and multiple loss functions, including cycle, identity, and SSIM losses, to maintain anatomical details between CT and MR images.
- Radiologist validation and metric improvements like SSIM and mutual information scores indicate the framework’s potential for more accurate clinical assessments.
Structurally Aware Bidirectional Unpaired Image Translation Between CT and MR
Introduction
The paper "Structurally aware bidirectional unpaired image to image translation between CT and MR" explores the use of Generative Adversarial Networks (GANs) for transforming Computed Tomography (CT) images into Magnetic Resonance (MR) images and vice versa. CT and MR are essential imaging modalities in medical diagnostics and surgical planning, each offering unique insights into anatomical structures. CT excels in bone imaging, while MR provides detailed soft tissue contrast. This study addresses the challenge of translating images between these modalities, which is particularly difficult due to the inherent differences in their acquisition processes.
The research focuses on employing GAN-based techniques, specifically cycleGAN, to facilitate bidirectional image translation without paired data. By leveraging the cycling consistency and structural similarity of CT and MR images, the paper demonstrates successful image synthesis. The generated images, while not perfectly identical to true counterparts, are validated by radiologists as clinically useful.
Materials and Methods
Data Collection and Pre-processing
The study utilized human pelvis CT and MR 3D volume datasets unpaired and sourced from multiple patients. CT volumes were sourced from the Liver Tumour Segmentation (LiTS) challenge database, and MR images were acquired using a 1.5 Tesla system from Apollo Speciality Hospital in Chennai. The data split consisted of 50 volumes for training and 5 for testing, with average dimensions of (512, 512, 80) for CT and (768, 768, 30) for MR.
Pre-processing involved resampling volumes, normalizing image slices, and resizing using bicubic interpolation followed by random cropping. Data augmentation included flipping and rotating images to enhance robustness during training.
The pipeline employed two generator networks for CT-to-MR and MR-to-CT translations (GCT and GMR), utilizing ResNet architecture with 9 resblocks. Corresponding discriminator networks (DCT and DMR) aimed to distinguish between original and translated images.
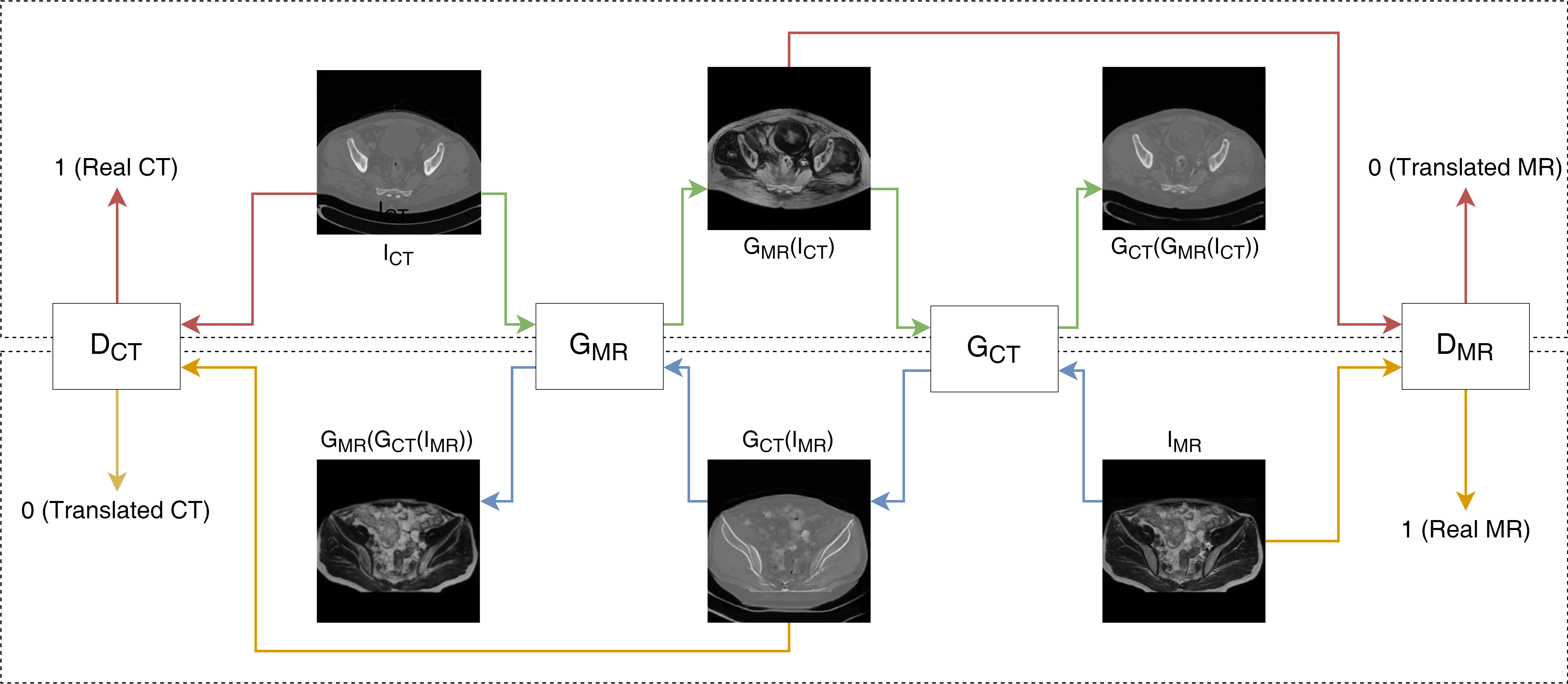
Figure 1: Schematics of our method used inter sequence MRI images.
Loss Functions
The study implemented various loss functions to preserve image integrity during translation:
- GAN Loss: Ensures translated images are not recognized as fake by the discriminator.
- Cycle Loss: Maintains cyclic consistency between input and generated images.
- Identity Loss: Ensures generators recognize input image modality.
- Structural Similarity (SSIM) Loss: Preserves structural integrity between image domains.
Performance metrics included Fréchet Inception Distance (FID), Mutual Information (MI), Structural Similarity Index (SSIM), and pixel-wise accuracy (pixacc).
Results and Discussion
The cycleGAN and cycleGAN-SSIM models were compared based on various metrics across 200 slices. Despite similar performance in most metrics, the cycleGAN-SSIM model demonstrated slight improvements in SSIM and MI scores, indicating better structural integrity in translations.
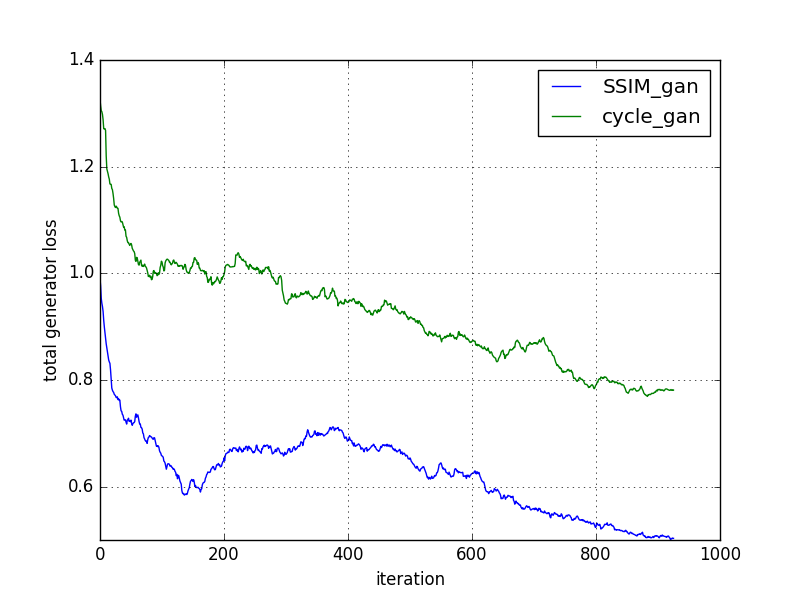
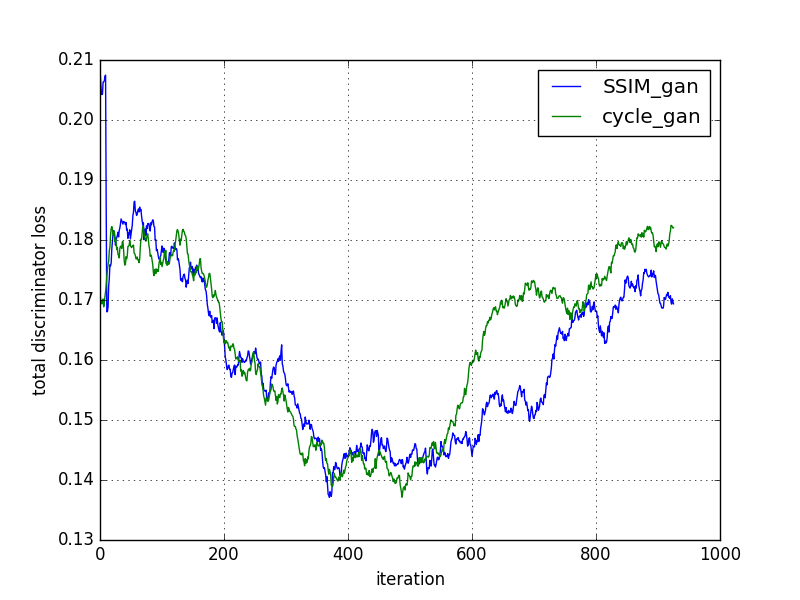
Figure 2: Comparison of models based on loss convergence.
Visual analysis of CT-to-MR and MR-to-CT translations showed enhanced bone detail preservation and textural integrity, especially with cycleGAN-SSIM. Radiologists validated the translated images for practical use, highlighting the cycleGAN model's ability to produce diagnostically useful images, while cycleGAN-SSIM excelled in special diagnostic cases involving structural details.
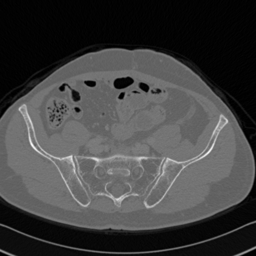


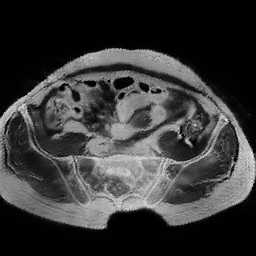
Figure 3: CT to MR conversion. In the above image column (i) corresponds to real CT, (ii) generated MR using cycleGAN (iii) recovery CT using cycleGAN (iv) generated MR with cycleGAN-SSMI (v) recovery CT with cycleGAN-SSMI.
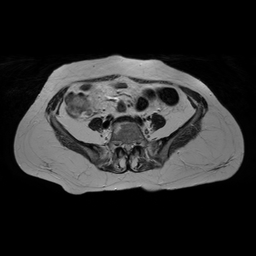


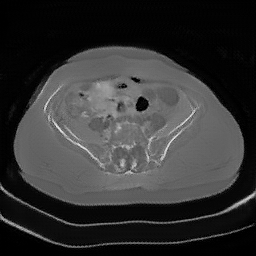
Figure 4: MR to CT conversion. In the above image column (i) corresponds to real MR, (ii) generated CT using cycleGAN (iii) recovery MR using cycleGAN (iv) generated CT with cycleGAN-SSMI (v) recovery MR with cycleGAN-SSMI.
Conclusion and Future Work
The study successfully demonstrated the application of cycleGAN for medical image translation tasks, revealing enhanced learning rates and structurally sound images when integrating SSIM loss. These findings suggest potential improvements in diagnosing fractures and other conditions. Future research aims to extend the methodology to 3D volumes for comprehensive anatomical analysis and to apply the approach in pulmonary diagnostics for nodules and calcifications, as indicated by radiologist feedback.






