- The paper introduces a deep Siamese network that learns local geometric features for enhanced shape correspondence in complex anatomical structures.
- The method integrates supervised learning on geodesic patches with domain adversarial training for unsupervised feature adaptation.
- Results demonstrate significant improvements in precision and generalization over traditional models on both synthetic and clinical datasets.
Learning Deep Features for Shape Correspondence with Domain Invariance
Introduction
In addressing the challenge of generating shape correspondence for complex anatomical structures, this paper proposes a method utilizing deep convolutional neural networks (CNNs) to learn shape features that are conducive to effective correspondence estimation. Such correspondence models are critical in medical imaging for tasks encompassing statistical shape analysis and clinical diagnostics. Traditional models relying on manual landmarking or parameterized transformation often fail to generalize to non-linear anatomical variances due to simplistic assumptions. This paper extends previous work by automating feature extraction using deep learning, thus accommodating complex shape variations through supervised and unsupervised learning paradigms.
Methodology
The approach centers on the extraction of local geometric features directly from surface topology, bypassing the reliance on predetermined descriptors and landmarks. The features are automatically learned via a deep Siamese network, trained on geodesic patches of shapes. This supervised learning method is augmented by domain adversarial training to adapt features for anatomically complex regions without labeled correspondence data.
Features are computed by using geodesic patches around a point on the shape's surface (Figure 1). These patches represent local surface geometry via circular patches formed by rays at various angles relative to principal curvature directions, encoded as pixel distances from the tangent plane.
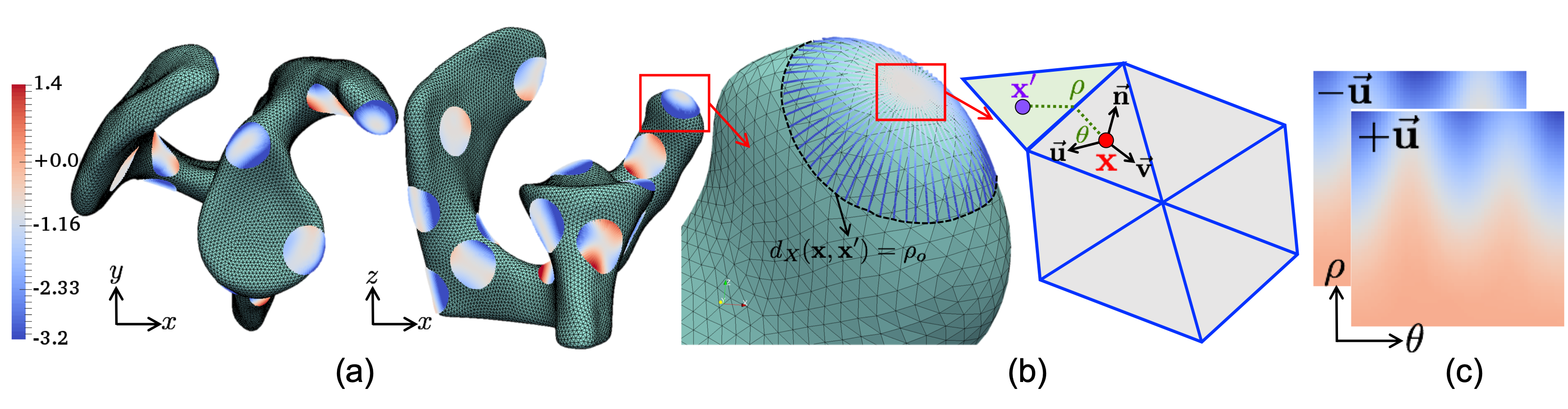
Figure 1: Geodesic patch extraction at a surface point, encoded into two-channel input for CNN.
Supervised Feature Learning
A Siamese network architecture is employed to distinguish between corresponding and non-corresponding point pairs. The CNN learns to map these point-based patches into a feature space where geometrically similar structures are well-aligned. The network's final features are used to assist in optimizing a particle-based shape model (PSM) by incorporating anatomical features beyond mere positional data.
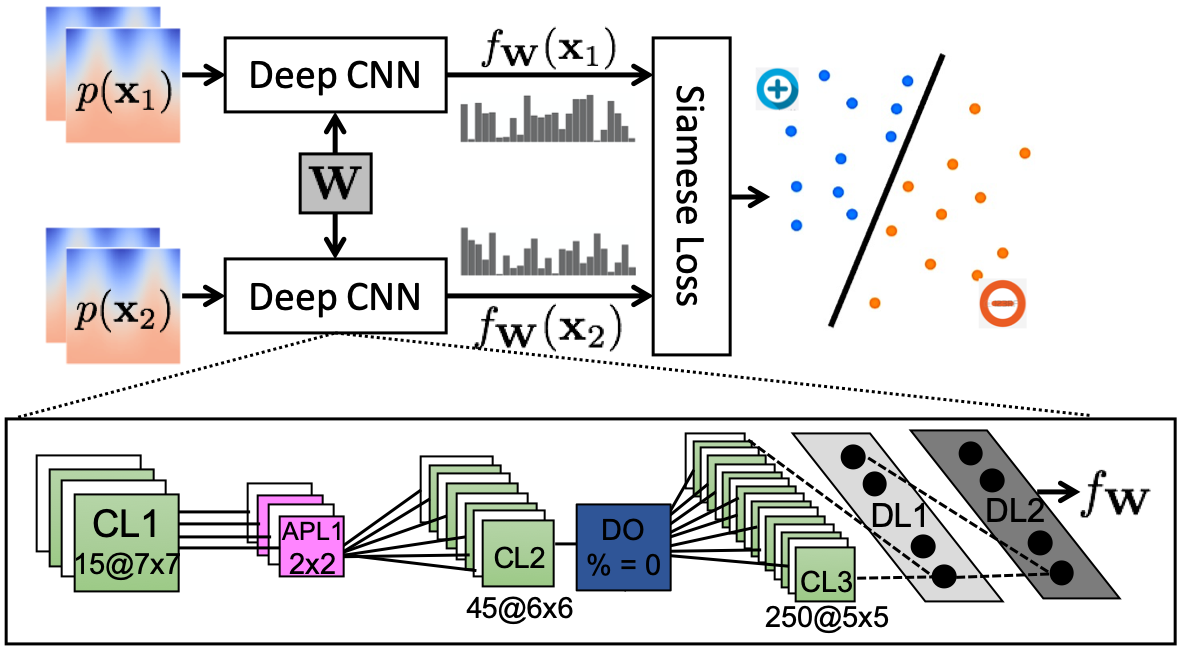
Figure 2: The Siamese network architecture used to learn features for correspondence.
Unsupervised Feature Adaptation
For anatomies where manual correspondences are neither feasible nor available, domain adaptation techniques are utilized. Leveraging adversarial training, the pre-learned features from a simpler anatomy serve as a basis to generate domain-invariant features without requiring target anatomy labels. This method enables effective feature adaptation to anatomies like the pelvis where complex features dominate.
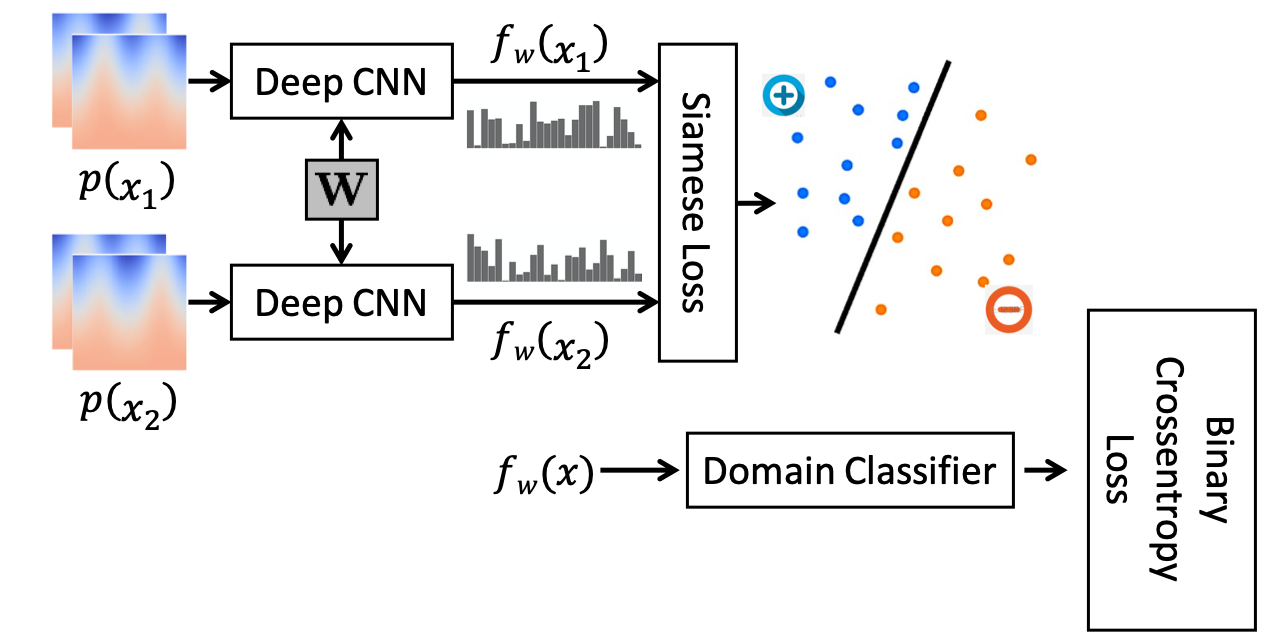
Figure 3: The combined network configuration used for adversarial training of the Siamese network.
Results
Supervised Learning on Synthetic Data: Deep feature PSM outperformed positional PSM in compactness and precision, as verified using synthetic Bean datasets. Evaluation metrics such as variance, generalization, and specificity were improved, indicating a robust feature representation.
(Figure 4-5)
Figure 4-5: Comparison of evaluated metrics showing superior performance of the proposed approach over traditional methods.
Clinical Data Application: Models trained on the scapula dataset demonstrated successful feature generalization with enhanced correspondence accuracy across patient and control groups. The CNN learned to capture relevant anatomical structures, vital for discriminating healthy from diseased specimens.
Feature Adaptation to Complex Anatomies: Utilizing the domain-adapted features, features learned from femur anatomy were successfully transferred to the pelvis structure. The adapted features highlighted anatomical regions of interest with a similar effectiveness to a manually trained pelvis model.
(Figure 5-8)
Figure 5-8: Examples of feature adaptation showing learned characteristics on varying anatomical structures.
Conclusion
The paper positions deep learning as a transformative tool in anatomical shape correspondence modeling. By leveraging both supervised and unsupervised learning, it addresses the computational and manual challenges associated with complex anatomical shapes. With further improvements, such as incorporating image data and optimizing computational efficiency, the techniques proposed promise to enhance the accuracy and applicability of statistical shape models in medical imaging.


