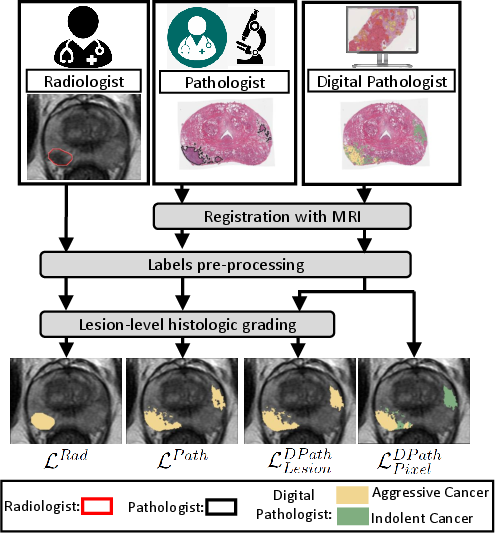
- The paper demonstrates that using digital pathologist labels improves deep learning model accuracy and distinguishes between aggressive and indolent prostate cancers.
- It compares deep learning architectures such as SPCNet, U-Net, branched U-Net, and DeepLabv3+ trained on radiologist, pathologist, and digital pathology labels.
- The findings show that models trained with digital pathologist labels achieve higher Dice overlaps and ROC-AUC scores, indicating potential for more standardized diagnostics.
Bridging the Gap Between Prostate Radiology and Pathology Through Machine Learning
Introduction
Prostate cancer remains a significant clinical problem, ranking as the second deadliest cancer among American men. Magnetic Resonance Imaging (MRI) plays a crucial role in the detection and localization of prostate cancer, often guiding targeted biopsies. However, the high false positive and negative rates, coupled with challenges like inter-reader variability, limit the utility of MRI. This research investigates the optimization of prostate cancer detection on MRI using machine learning models, emphasizing the role of different training label types and introducing comparisons among them.
Methodology
Study Design
The study compared different labeling strategies to evaluate their impact on machine learning model performance for prostate cancer detection. Four deep learning models—SPCNet, U-Net, branched U-Net, and DeepLabv3+—were trained with datasets consisting of various label types, including pathology-confirmed radiologist labels, pathologist labels, and digital pathologist labels at both lesion and pixel levels.
Radiologist and pathologist labels were mapped onto MRI using automated registration platforms. The digital pathologist labels, derived from deep learning algorithms validated on histopathology images, offered a novel training path distinct from the globally utilized manual labeling.
Datasets and Labels
Data was acquired from 315 patients divided into radical prostatectomy and biopsy cohorts. The labeling included aggressive versus indolent cancer distinctions, essential for evaluating the machine learning models' precision in cancer identification.
Label Types
Results
The SPCNet model consistently demonstrated superior performance across various label types. Digital pathologist labels enabled models to achieve higher or comparable accuracy compared to traditional radiologist or pathologist label-trained models. These models also uniquely facilitated the differentiation between aggressive and indolent cancer types, paving the way for more nuanced prostate cancer diagnostics.
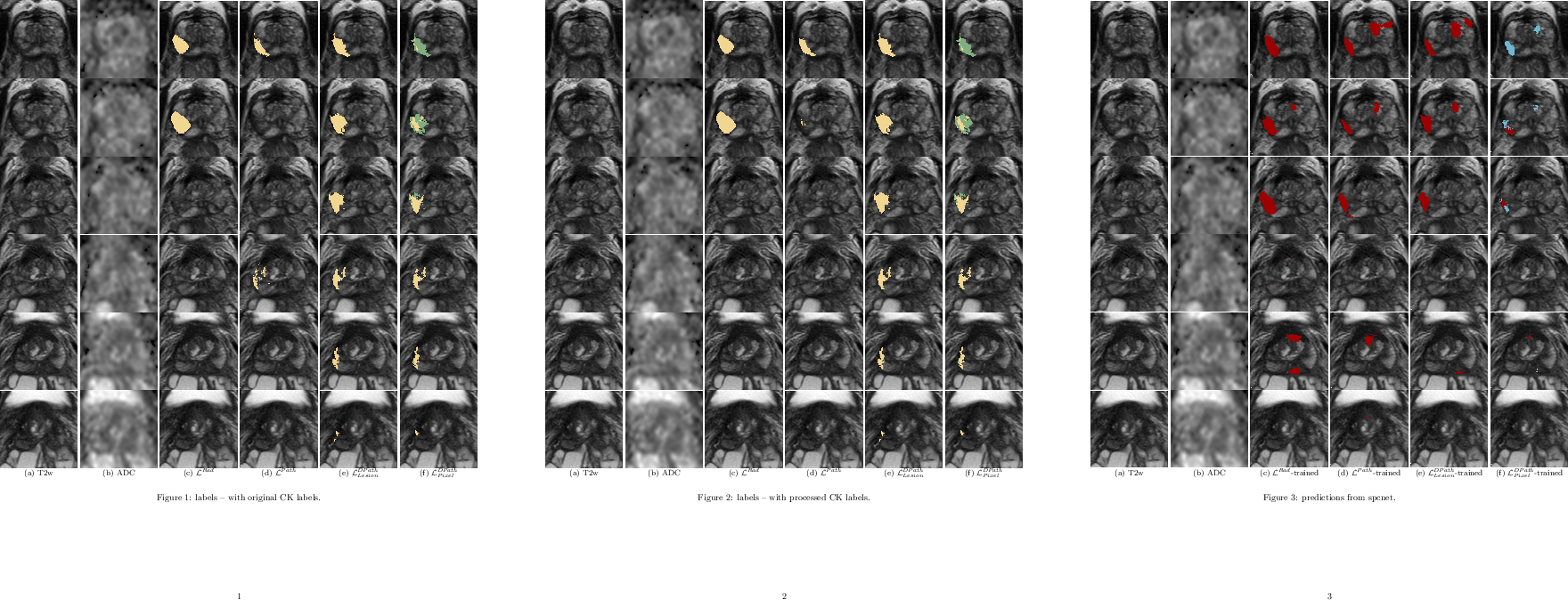
Figure 2: Differences in labeling strategies in a typical patient in cohort C1-test (aggressive cancer - yellow, indolent cancer - green) showed on (a) T2w images and (b) ADC images...
Comparison of Labeling Strategies
Models trained on digital pathologist labels achieved superior Dice overlaps and lesion-level ROC-AUCs compared to radiologist-trained models. The pixel-level digital pathologist label models excelled at detecting mixed lesions within aggressive and indolent cancer contexts, showcasing an unprecedented discriminative ability not feasible with human annotations alone.

Figure 3: Predictions from SPCNet trained with different label types of a typical patient from cohort C1-test (same as Figure 2).
Discussion
The use of digital pathologist labels provides a robust methodological innovation. It not only challenges but potentially surpasses the conventional human expert label practices in prostate cancer detection via MRI. The findings suggest that these models can standardize and potentially enhance diagnostic accuracy while limiting the constraints of labor-intensive pathology labeling.
Furthermore, digital pathologist labels tackle several challenges, such as inter- and intra-reader variability and the intensive labor associated with human-generated annotations. They present a pathway to refine prostate cancer care by reliably distinguishing between aggressive and indolent forms, thus materially influencing treatment strategies.
Conclusion
Digital pathologist labels represent a significant advancement for training deep learning models in prostate cancer detection. This approach enhances model reliability and expands the potential for accurate, standardized prostate cancer diagnostics. Such innovations hold promise for improved clinical decision-making and patient outcomes in prostate cancer management.
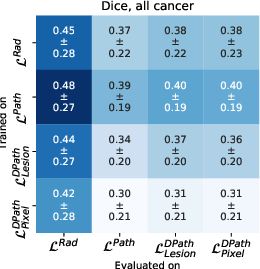
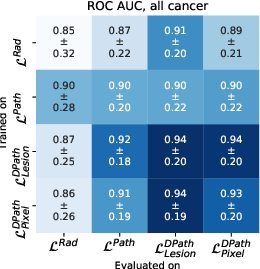
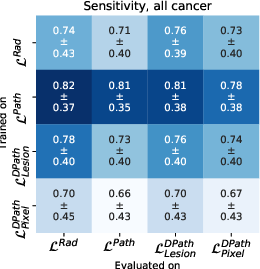
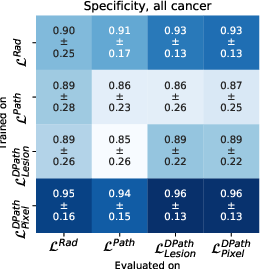
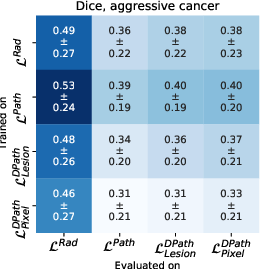
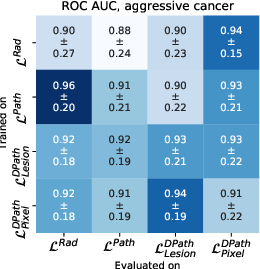
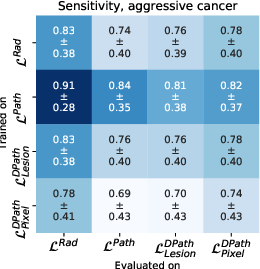
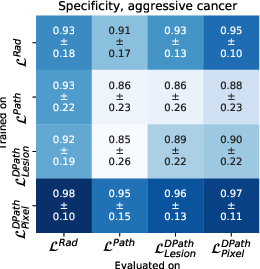
Figure 4: Quantitative comparison between digital radiologist (SPCNet) predictions when trained and evaluated using different label types in cohort C1-test.