- The paper introduces a dynamic online pseudo labeling method using momentum networks to stabilize training.
- It leverages a teacher-student framework with only 20% labeled data, achieving an average Dice Score of 84.1%.
- The method advances medical image segmentation by approaching fully-supervised performance on several polyp datasets.
Online Pseudo Labeling for Polyp Segmentation with Momentum Networks
Introduction
The paper "Online pseudo labeling for polyp segmentation with momentum networks" (2209.14599) addresses a critical challenge in the field of medical image diagnosis, particularly concerning the segmentation of polyps. Traditional deep learning models for semantic segmentation require large annotated datasets, which are expensive and labor-intensive to produce, especially for medical images requiring pixel-level annotations. The study leverages semi-supervised learning (SSL) techniques to refine the segmentation process without relying extensively on labeled data. This is accomplished by introducing an innovative online pseudo labeling strategy combined with momentum networks to improve the quality of pseudo labels during training.
Methodology
The authors propose a multi-stage semi-supervised training framework. Initially, a teacher model is trained on a limited labeled dataset. This model then generates pseudo labels for a student model trained on a combination of labeled and unlabeled data. Unlike previous methods, which treat the teacher model as static during student training, this approach dynamically updates the teacher model using a momentum strategy. The momentum teacher model is essentially a slow-copy version of the original model, maintained via an Exponential Moving Average (EMA) to stabilize updates and provide smoother pseudo labels.
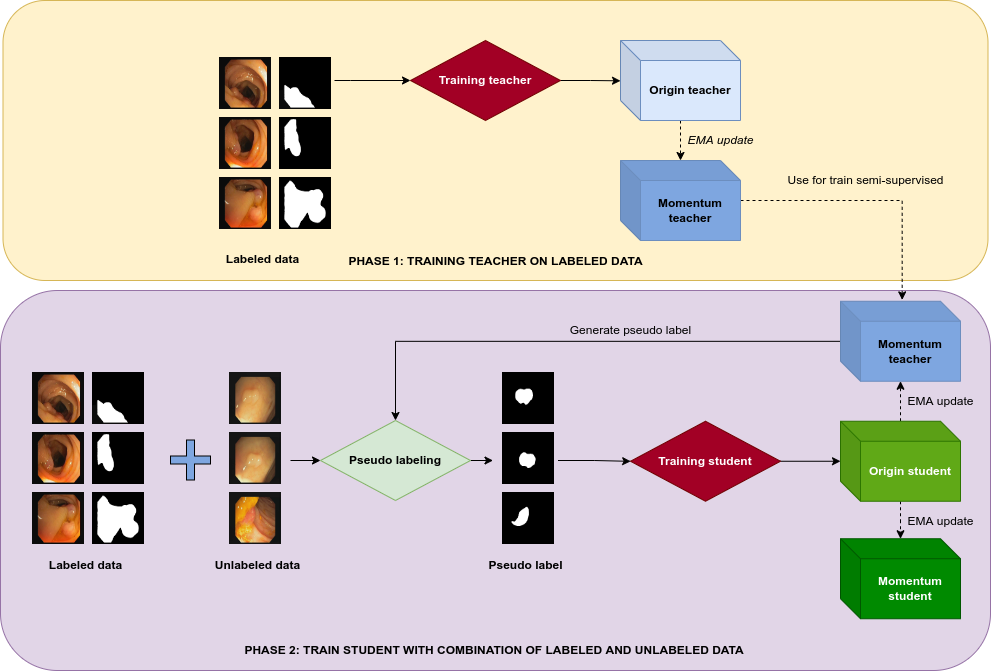
Figure 1: Overview of online pseudo labeling with momentum network pipeline.
Experimental Setup and Results
The experiments were conducted on five prominent datasets: Kvarsir, CVC-ClinicDB, ETIS-LaribPolypDB, CVC-ColonDB, and CVC-300, using only 20% of the data as labeled. The study reports a notable improvement in the Dice Score metrics, achieving an average of 84.1%, which surpasses previous common practices by 3%. The method even approaches fully-supervised performance on some datasets, highlighting the efficacy of the proposed online pseudo labeling combined with momentum networks.
Qualitative Analysis
The effectiveness of this approach is also demonstrated qualitatively. Comparisons between offline and online pseudo labeling showcase the superior segmentation quality achieved through the proposed method. Figure 2 elucidates the visual differences, underscoring the enhancements in label quality and subsequent segmentation performance attributable to the momentum network's influence during online pseudo labeling.
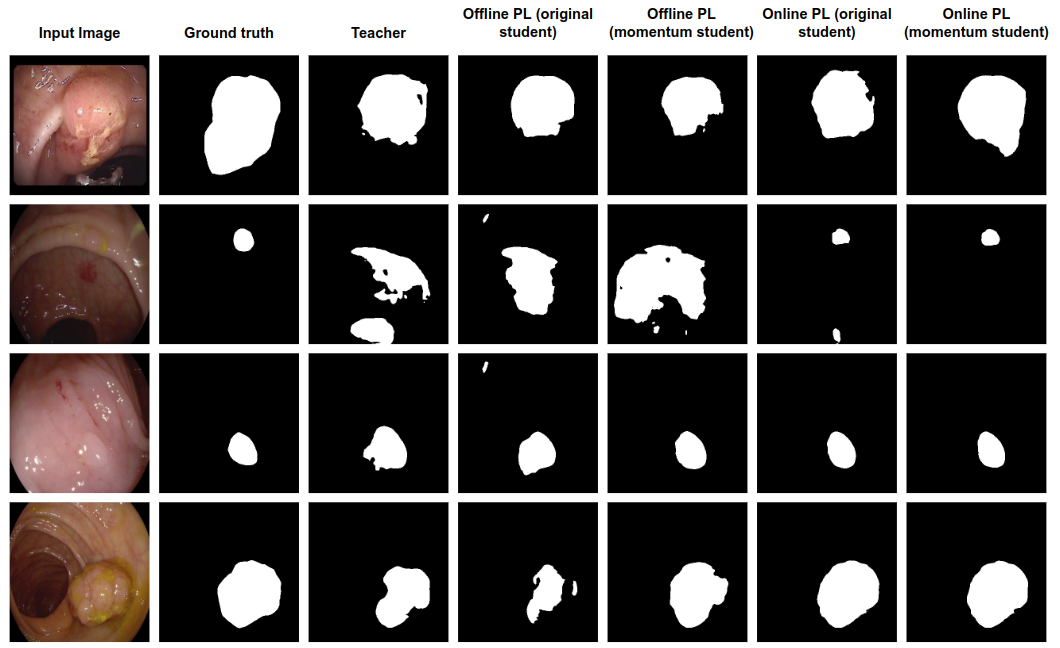

Figure 2: Qualitative result comparison between offline pseudo labeling and online pseudo labeling, with/without using momentum network.
Implications and Future Directions
The method demonstrates significant potential for reducing dependency on fully labeled datasets in medical image analysis, promoting the application of SSL in practical healthcare scenarios. By enhancing pseudo label accuracy, this approach could streamline the development of robust medical diagnostic tools. Future research could explore adapting this framework to a broader range of medical imaging tasks, extending beyond polyp segmentation to areas like tumor or lesion detection in other organs.
Conclusion
The study presents a compelling semi-supervised learning framework that effectively combines online pseudo labeling with momentum networks to improve medical image segmentation, showing both practical and theoretical innovation in SSL methodologies. This advancement not only highlights a pathway for more efficient use of unlabeled data in medical imaging but also sets a foundation for further research in enhancing semi-supervised approaches across various domains.

