- The paper introduces a three-level alignment strategy that aligns images and reports at pathological region, instance, and disease levels.
- Experimental results show significant improvements in classification, detection, and segmentation tasks across multiple medical imaging datasets.
- The MGCA framework demonstrates data efficiency by learning discriminative visual features even with limited labeled data.
Multi-Granularity Cross-modal Alignment for Generalized Medical Visual Representation Learning
The paper introduces the Multi-Granularity Cross-modal Alignment (MGCA) framework, aiming to improve medical visual representation learning by utilizing semantic correspondences across medical images and radiology reports. It adopts a three-level alignment approach—pathological region-level, instance-level, and disease-level—to enhance the representation capabilities for downstream tasks like image classification, object detection, and semantic segmentation. The framework demonstrates superior performance in these contexts, particularly with limited labeled data.
Introduction
Traditional deep learning and contrastive learning approaches have been effective in medical imaging tasks but struggle to incorporate the small yet crucial pathological areas in medical images and align them adequately with corresponding radiology reports. This paper addresses these limitations by proposing an MGCA framework that leverages multi-granularity semantic correspondences. With instance-wise, token-wise, and disease-level alignments, the MGCA framework aims to facilitate more efficient and generalizable visual learning, especially in environments with limited annotated data.
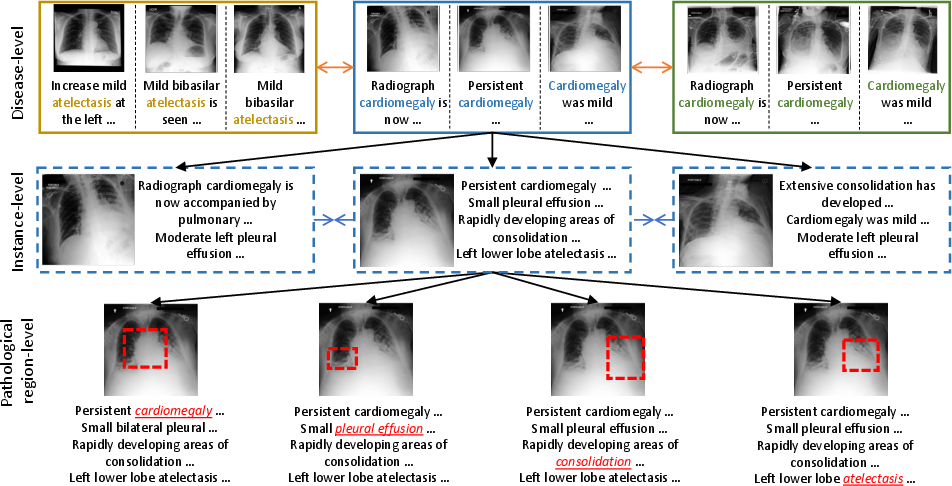
Figure 1: The multi-granularity (disease-level, instance-level, and pathological region-level) semantic correspondences across medical images and radiology reports.
Method
Multi-granularity Alignment Framework
MGCA employs a hierarchical vision-language semantic alignment across three levels:
- Instance-wise Alignment (ITA): Uses InfoNCE losses to differentiate true medical image-report pairs from random pairs based on cosine similarity, focusing primarily on aligning image-report pairs globally.
- Token-wise Alignment (CTA): Implements a bidirectional cross-attention mechanism to learn fine-grained token correspondences within image-text pairs, optimizing mutual information at the token level and providing better localized performance.
- Disease-level Alignment (CPA): Introduces a cross-modal prototype mechanism that exploits high-level semantic consistency between images and reports, thereby capturing disease-level relationships across modalities.
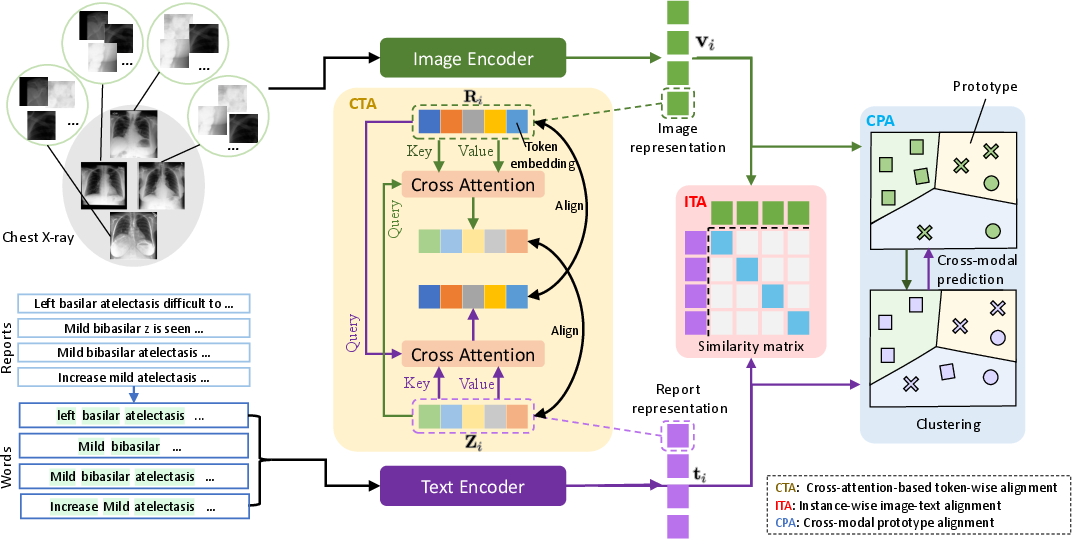
Figure 2: Illustration of our proposed multi-granularity cross-modal alignment framework. CTA, ITA, and CPA represent token-wise alignment, instance-wise alignment, and prototype (disease)-level alignment respectively. The green arrow represents information flow of visual features, while the purple arrow represents information flow of textural features.
Experiments
Pre-training and Evaluation
MGCA was pre-trained using the large-scale MIMIC-CXR dataset, and its effectiveness was evaluated across seven datasets representing various tasks in medical imaging. The representation learning was fine-tuned on medical image classification, object detection, and semantic segmentation tasks.
Results
MGCA demonstrated substantial performance improvements over comparable frameworks such as GLoRIA and ConVIRT, particularly in environments with limited labeled data. For instance, the classification tasks on the COVIDx dataset showed significant accuracy improvements, asserting the framework's data-efficient capabilities. The object detection and segmentation tasks further exemplified MGCA’s ability to learn discriminative and localized representations.
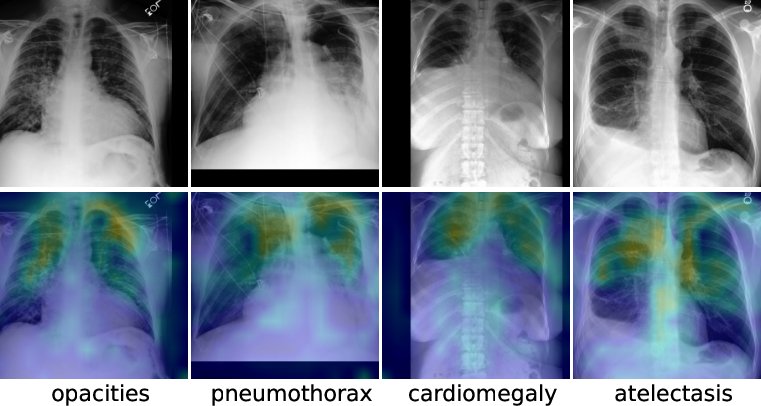
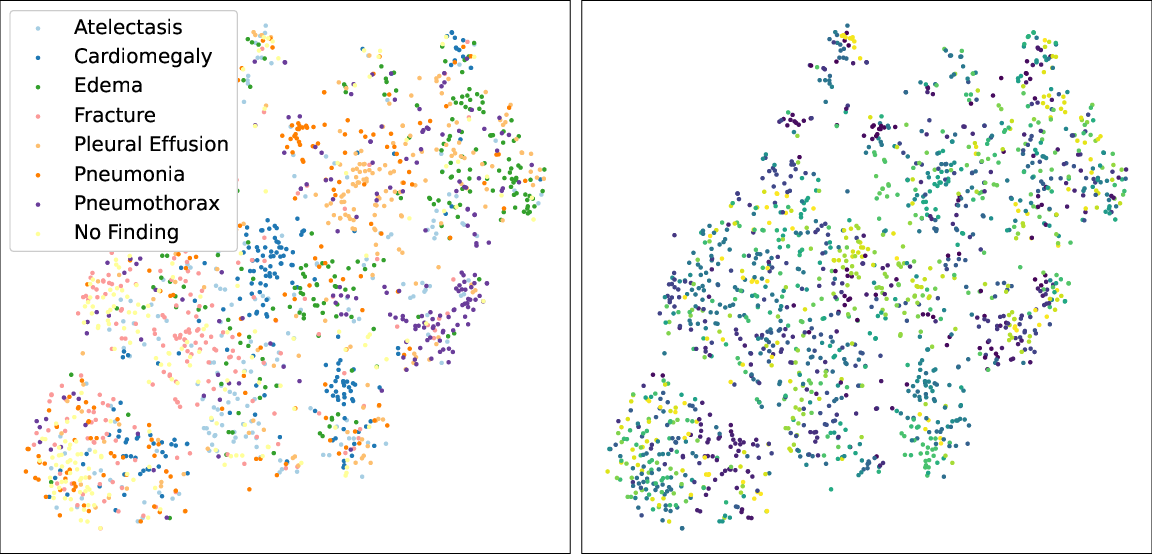
Figure 3: Visualization of learned token correspondence by our MGCA. Highlighted pixels represent higher activation weights by corresponding word.
Conclusion
MGCA effectively exploits the inherent semantic structures between medical images and radiology reports across multiple levels, offering a robust alternative for generalized medical visual representation learning. Despite its promising application in medical diagnosis with reduced annotation requirements, future work could explore integration of discrimination and generation-based pre-training methods, as well as holistically leveraging the multi-granularity correspondences.
The proposed framework demonstrates strong potential for reducing radiologist workload and enhancing diagnostic efficiency, although caution should be exercised regarding usage with data containing personal information. Overall, MGCA represents a step forward in the application of AI to medical imaging, with significant implications for future research and healthcare improvement strategies.



