- The paper introduces GeoLDM, a novel latent diffusion model that combines variational autoencoding with equivariant networks to enhance 3D molecule generation.
- It leverages a compact latent space respecting SE(3) symmetry, leading to significant improvements in molecule validity and stability.
- GeoLDM supports controllable molecule design by integrating conditional molecular properties, advancing applications in drug discovery and material science.
Geometric Latent Diffusion Models for 3D Molecule Generation
Introduction
The paper presents Geometric Latent Diffusion Models (GeoLDM), a novel approach to 3D molecular geometry generation utilizing latent diffusion models. GeoLDM addresses limitations of existing molecular generative models by leveraging a compact latent space to enhance generation capacity, particularly for complex molecular structures. This is achieved through a combination of variational autoencoding and diffusion processes, incorporating both invariant and equivariant features to maintain roto-translational equivariance essential for 3D molecule modeling. The method demonstrates significant improvements in generating chemically plausible molecules and supports controllable generation based on desired molecular properties.
Methodology
GeoLDM models are structured as variational autoencoders (AEs) where molecular structures are encoded into a latent space characterized by both invariant scalars and equivariant tensors. The encoder compresses raw atomic features into a lower-dimensional latent space, which DMs then manipulate to generate new molecular structures. This transformation reduces the complexity of modeling intricate molecular data while maintaining the geometric properties essential for realistic molecule generation.
Latent Representation and Equivariance:
The key innovation in GeoLDM is the introduction of equivariance in the latent space. The latent space is configured to respect the SE(3) group symmetry, crucial for physical simulations involving 3D rotations and translations. This is done by designing encoder and decoder networks using equivariant graph neural networks (EGNNs), which ensure that latent representations and their geometric transformations remain consistent across transformations such as rotations.
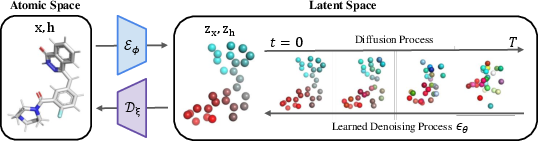
Figure 1: Illustration of GeoLDM. The encoder ϕ encodes molecular features into equivariant latent variables, and the latent diffusion transitions q(zt) add noise until convergence to a Gaussian.
Experimental Results
Molecular Modeling and Generation
GeoLDM was evaluated on multiple benchmark datasets, including QM9 and GEOM-DRUG. It showed superior performance compared to state-of-the-art models, achieving higher atom and molecule stability percentages and validity rates. Notably, GeoLDM achieved up to a 7% improvement in validity for large biomolecules, a significant advancement over existing methods.
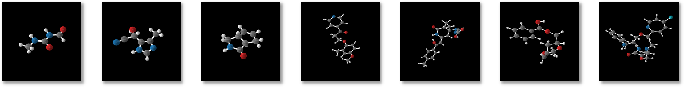
Figure 2: Molecules generated by GeoLDM trained on QM9 (left three) and DRUG (right four).
Controllable Generation
GeoLDM extends beyond unconditional molecule generation by enabling controllable generation based on molecular properties such as polarizability and heat capacity. The model seamlessly integrates conditional information into the generative process, achieving lower mean absolute errors (MAE) in property predictions compared to other approaches. This capability is particularly important for drug design applications where specific molecular attributes are desired.
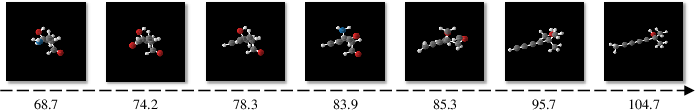
Figure 3: Molecules generated by conditional GeoLDM. Controllable generation is demonstrated by interpolating different polarizability values.
Theoretical Analysis
The authors provide a comprehensive theoretical foundation illustrating that GeoLDM's training objectives are variational lower bounds of log-likelihoods and invariant under SE(3) group transformations. This ensures that the latent distributions and generated outputs maintain essential geometric properties, which is vital for modeling physical systems accurately.
Conclusion and Future Work
GeoLDM represents a significant step forward in 3D molecular generation, offering enhanced flexibility and control through an innovative use of latent diffusion processes. Its ability to maintain geometric constraints while operating in a reduced-dimensional latent space opens new possibilities for efficient and precise molecular design, particularly in the realms of drug discovery and material science. Future work may focus on further optimizing the training process and extending GeoLDM's capabilities to encompass larger and more complex molecular structures, potentially incorporating protein and peptide generation tasks.




