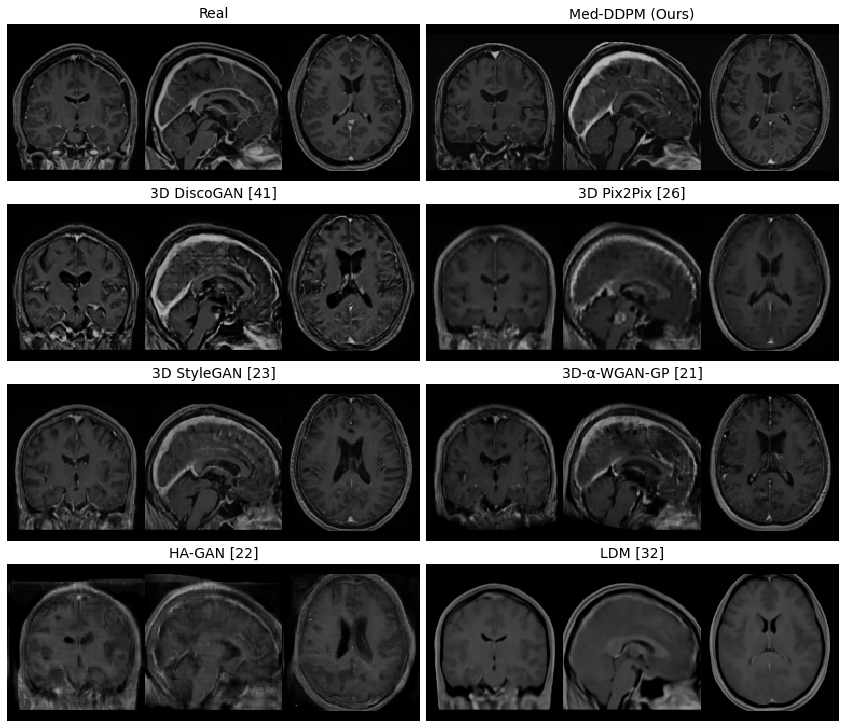
- The paper introduces Med-DDPM, a conditional diffusion model that leverages semantic masks to generate high fidelity 3D brain MRI images closely resembling real data.
- It extends traditional DDPM with a novel pixel-level conditioning mechanism and employs a cosine noise schedule to achieve dice scores near those of real MRIs.
- The method offers robust data augmentation and privacy-preserving applications, validated by both quantitative metrics and clinical assessments.
Conditional Diffusion Models for Semantic 3D Brain MRI Synthesis
Introduction
The introduction of generative models marks a significant milestone in addressing challenges associated with data scarcity and privacy concerns prevalent in medical imaging. The emergence of the Med-DDPM (Medical Denoising Diffusion Probabilistic Model) signals a critical advancement by integrating semantic conditioning into diffusion models for 3D brain MRI synthesis. Unlike traditional 2D image generation methods, Med-DDPM harnesses the power of semantic conditioning, wherein a conditioning image is concatenated channel-wise with the model input. This configuration provides a controlled, high-fidelity synthesis process. Med-DDPM excels in generating diverse, anatomically realistic images with high visual fidelity, bringing it close to real-world datasets in segmentation tasks.
Methodology
Med-DDPM extends the conventional Denoising Diffusion Probabilistic Models (DDPM), grounding its model architecture on the architectural innovations of generative models while implementing a novel pixel-level conditioning mechanism. The forward diffusion and denoising processes in Med-DDPM are defined by specific mathematical formulations with notable augments such as utilizing a cosine noise schedule and segmenting the mask integration via a channel-wise concatenation method to create a meaningful noise-augmented image x~t.
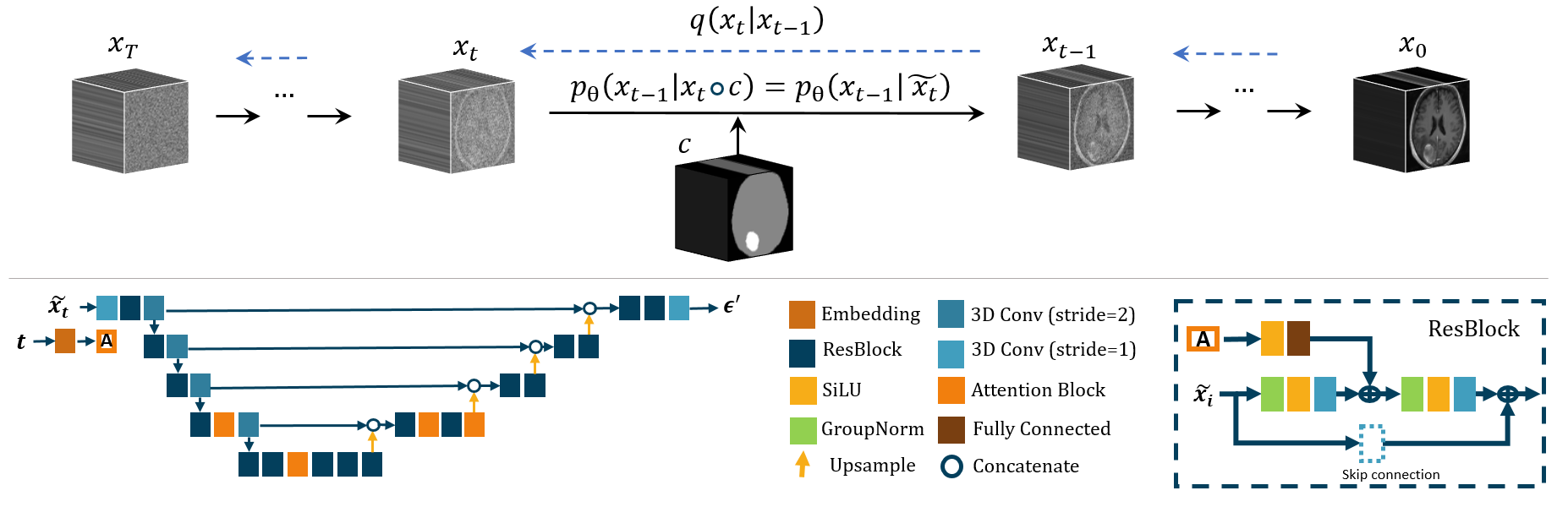
Figure 1: Architecture of the proposed method: The top row of the diagram demonstrates the conditioning mechanism.
- The forward diffusion process introduces Gaussian noise into image samples, establishing an equilibrium that Med-DDPM exploits for a streamlined reverse diffusion, effectively enriching the resultant synthetic image's quality.
- The semantic conditioning approach adapts dynamic control to the synthesis process by drawing on segmentation masks, opening up applications in privacy-preserving data anonymization and controlled pathological synthesis.
Experiments and Results
Data and Setup
The study leveraged clinical brain MRI data, handling 1,500 preprocessed images using a comprehensive image preprocessing protocol that included registration, cropping, and intensity normalization. Performance metrics for segmentation tasks employed the BraTS2021 challenge dataset, emphasizing the robustness of Med-DDPM in synthesizing all four MRI modalities (T1, T1CE, T2, and Flair) from segmentation masks.
Quantitative Evaluation
Med-DDPM demonstrates superior quantitative evaluation results, as evidenced by the experiments. Among notable results:
Qualitative Assessment
In independent qualitative assessments conducted by neurosurgeons, synthetic images crafted through Med-DDPM were challenging to distinguish from actual MRIs. This reaffirms its high detail fidelity vis-à-vis anatomical continuity and pathology representation.
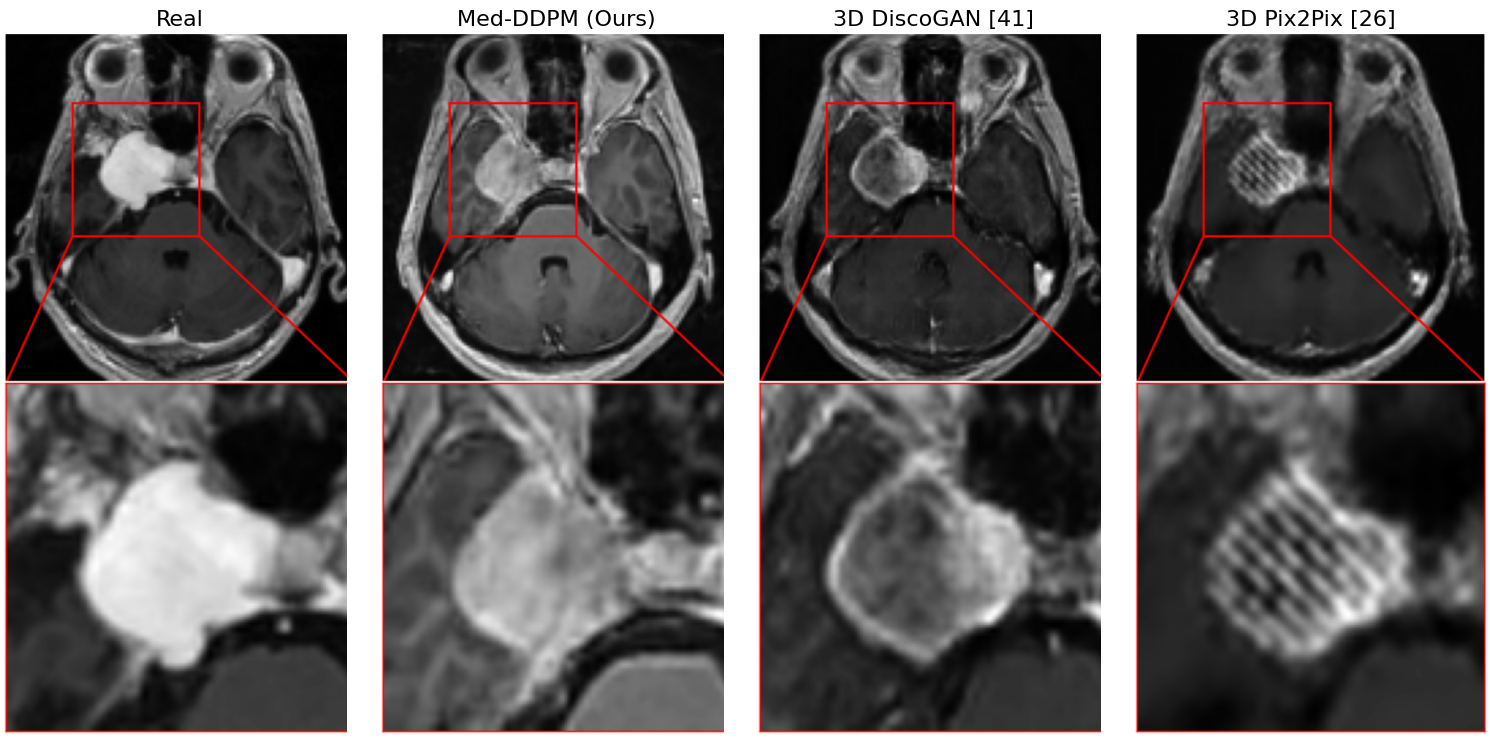
Figure 3: Zoomed visual comparison of tumor areas in real and generated samples.
Discussion
Med-DDPM marks a distinctive turn in semantic 3D brain MRI synthesis, underlining diffusion models' efficacy in surpassing traditional GAN-based approaches in image quality and stability. While engineered segmentation masks grant control, enabling customizable synthesis, certain modeling facets still necessitate advancement, particularly the accurate assimilation of structural features like vessel continuity around tumor regions.
Memory efficiency remains an area ripe for optimization—scrutinizing resource consumption juxtaposed with conditional GAN approaches delineates avenues for improvement.
Conclusion
Med-DDPM represents a substantial step forward in precision-driven medical data synthesis, offering nuanced image quality and segmentation accuracy that approaches real-world capabilities. It confirms the role of conditioned diffusion models in high-stakes applications like data augmentation and image anonymization. Future research must pivot towards refining architecture scalability within clinical settings, promoting extensive uptake across diverse medical imaging applications.
(Figure 4)
Figure 4: Center-cut axial slices of generated samples, showcasing the output diversity of Med-DDPM for a single input mask.