- The paper introduces customized EvoformerAttention kernels that reduce memory usage by 13x for structural biology models.
- It details a tiled computation strategy with on-the-fly broadcasting that improves memory efficiency without sacrificing accuracy.
- The paper also enhances the Megatron-DeepSpeed framework, boosting genomic model sequence length support by up to 14x.
DeepSpeed4Science: Enabling Large-Scale Scientific Discovery Through Sophisticated AI System Technologies
The paper introduces the DeepSpeed4Science initiative, which aims to extend the capabilities of AI systems beyond the generic support for LLMs to accelerate scientific discoveries by addressing unique challenges in various scientific domains. The paper details how DeepSpeed4Science is harnessing AI system technology innovations specifically tailored for structural biology and genomic scale models.
Addressing Memory Explosion in Structural Biology Models
DeepSpeed4Science tackles the memory explosion issue that arises when using Evoformer-centric models for structural biology tasks, such as protein structure prediction with OpenFold. The original OpenFold model suffers from excessively large activation sizes, particularly in the attention logit calculations, which can exceed 12GB in half precision for just one variant. This limits sequence lengths and MSA depths during training.
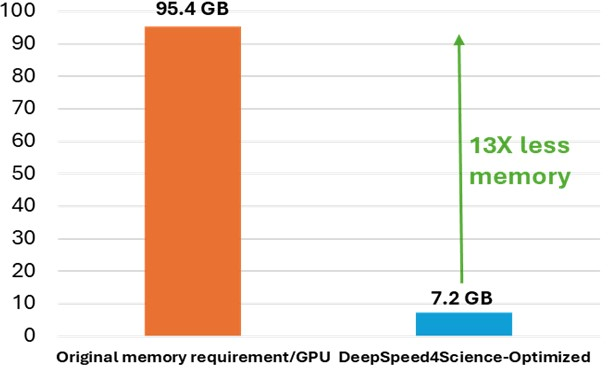
Figure 1: Peak memory requirement for training variants of the MSA attention kernels (with bias). It demonstrates the significant memory reduction achieved by DS4Sci_EvoformerAttention.
The initiative introduces DS4Sci_EvoformerAttention kernels to drastically reduce memory requirements by 13x without sacrificing accuracy. This is achieved by designing customized, efficient attention kernels that use sophisticated fusion and tiling strategies. These improvements allow larger and more complex models to be trained on broader hardware bases, boosting training efficiency and scaling capabilities (Figure 2).
Methodological Enhancements
The paper details the methodology employed to optimize EvoformerAttention, integrating exact attention kernels offloading computations in a highly efficient manner. The innovative approach involves calculating attention logits in tiles, enhancing memory efficiency, and fusing computation steps to minimize memory usage. It addresses challenges such as bias term broadcasting, necessary for backward compatibility of logits, through on-the-fly broadcasting techniques instead of relying on PyTorch's built-in functions (Figure 3).
Enabling Very Long Sequence Support for Genomic Scale Models
GenSLMs Framework Optimization
The paper describes optimizations in the Megatron-DeepSpeed (MDS) framework to support very long sequences, essential for exploring SARS-CoV-2 evolutionary dynamics with the GenSLMs model. MDS successfully combines tensor, pipeline, and sequence parallelism with memory optimization techniques to handle sequence lengths beyond those supported by traditional large model training frameworks like Megatron-LM.
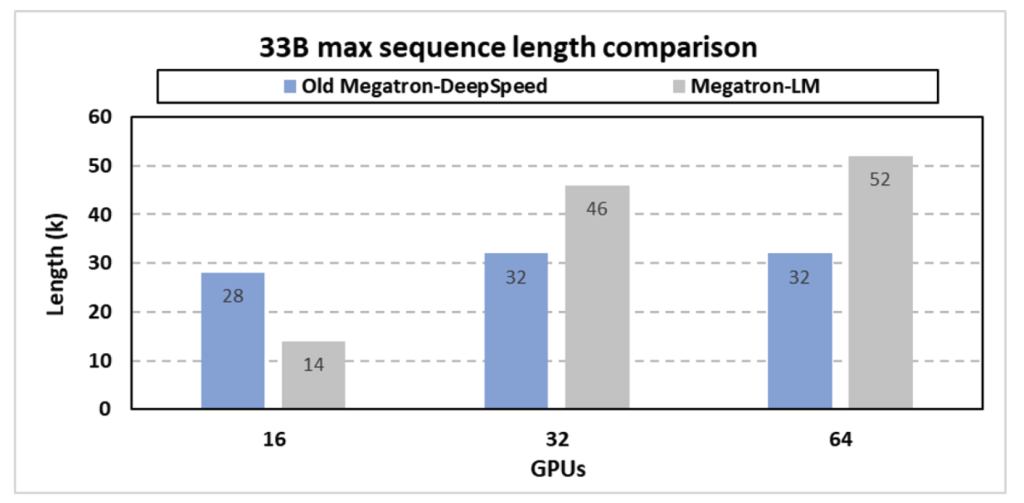
Figure 4: Maximum sequence length support for the 33B GenSLM model.
Enhanced Memory Optimization
DeepSpeed4Science introduces innovative memory-saving strategies such as efficient generation of attention masks and parallelization of position embeddings to scale sequence lengths significantly without encountering memory bottlenecks. By adapting these techniques, the new Megatron-DeepSpeed framework improves maximum sequence lengths by up to 14x compared to previous versions (Figures 8 and 9).
Conclusion
The DeepSpeed4Science initiative presents a transformative approach to harnessing AI technologies for scientific advancements. By addressing specific challenges in structural biology and genomic-scale model training, it optimizes memory usage and scaling capabilities, enabling researchers to tackle complex scientific problems with larger, efficiently trainable models. The initiative sets a foundation for future AI4Science technologies, promising deeper scientific insights across diverse domains.

