- The paper introduces X-LoRA, which employs a dynamic mixture of low-rank adapter experts to enhance scientific reasoning in large language models.
- It details a dual-pass inference and token-level gating mechanism that dynamically selects specialized adapters for tasks in protein mechanics and molecular design.
- Evaluation shows improved quantitative performance in protein property prediction (R²=0.85) and enhanced knowledge recall, outperforming larger, monolithic models.
X-LoRA: A Mixture-of-Low-Rank Adapter Experts Framework for Scientific LLM Specialization
Introduction and Motivation
The paper "X-LoRA: Mixture of Low-Rank Adapter Experts, a Flexible Framework for LLMs with Applications in Protein Mechanics and Molecular Design" (2402.07148) presents a novel approach for integrating diverse specialized capabilities into LLMs using a modular, efficient, and flexible mechanism. The X-LoRA architecture extends the concept of LoRA-based adaptation by orchestrating a dynamic, task-specific mixture of several independently fine-tuned LoRA adapters corresponding to distinct expertise domains. This enables simultaneous access to heterogeneous or overlapping scientific knowledge, efficient resource utilization, and deep integration of expertise even in models with relatively small parameter footprints.
The approach is directly motivated by the increasing demand for science- and engineering-focused LLM applications, such as in biomaterials and molecular design, where rigorous domain-specific reasoning, property prediction, and generative design tasks are paramount. The X-LoRA framework's architecture and training pipeline are influenced by biological paradigms, emphasizing reusability and hierarchical organization of computational blocks.
Architecture and Methodological Innovations
X-LoRA's architecture decouples the base LLM, the set of LoRA adapters, and a dedicated scaling head responsible for token- and layer-wise gating of the adapters. After standalone training of each LoRA adapter on respective scientific tasks (e.g., bioinspired materials, chemistry, protein mechanics, chain-of-thought), the scaling head is trained to compute dynamic mixing coefficients. These coefficients (scalings) are predicted from the base model's hidden states and applied to the adapters' outputs at each layer and token position.
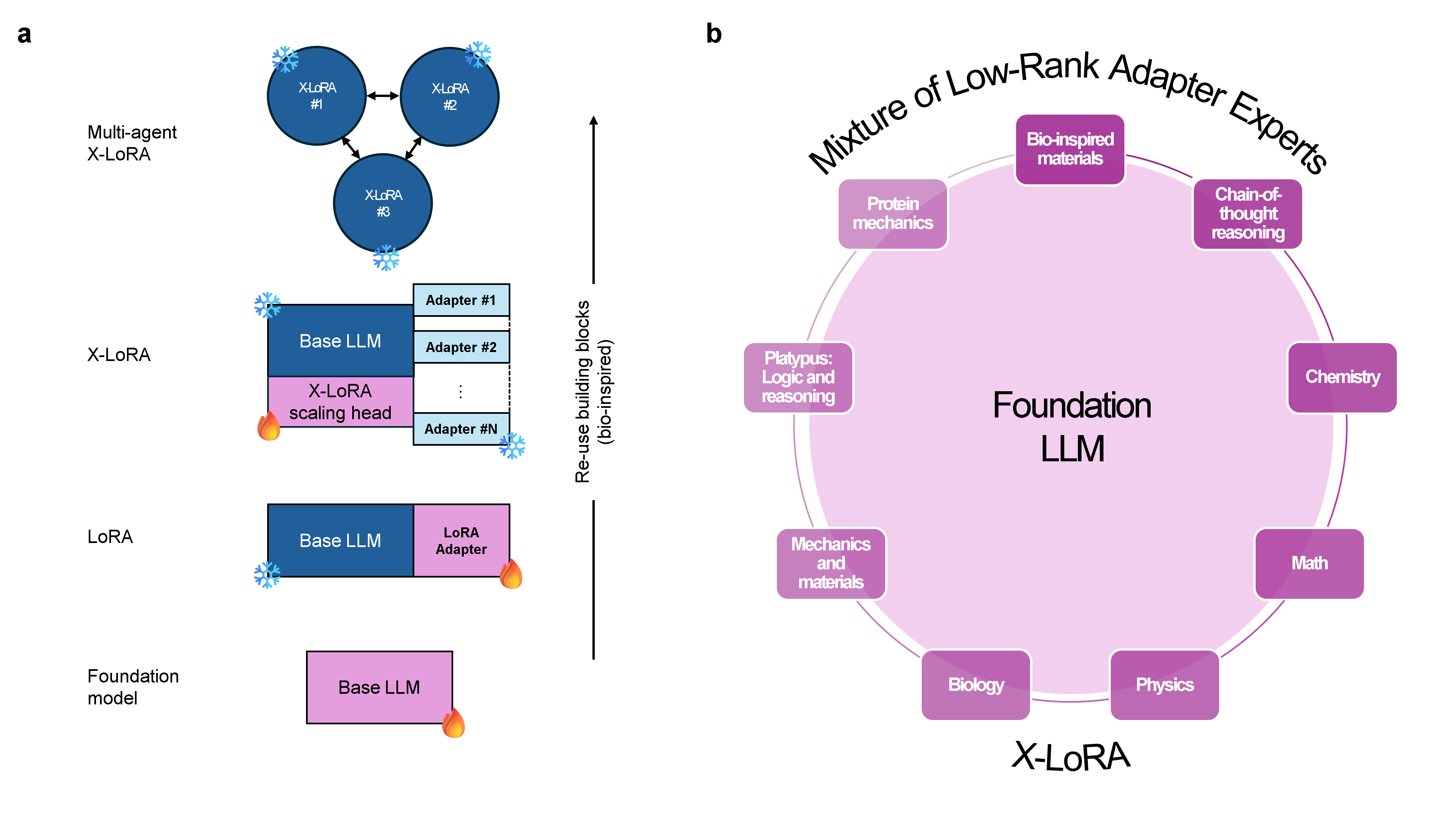
Figure 1: Multi-hierarchical design principle, highlighting frozen base model weights and trainable adapter layers spanning diverse scientific domains.
Formally, the forward computation at each adapted layer is:
h=W0x+i=1∑nBiAi(x⋅λi)αi
where λi are scaling factors predicted for each adapter by the scaling head using current hidden states, and αi are native LoRA scaling parameters.
The dual-pass inference protocol involves an initial forward pass to calculate the hidden states used by the scaling head, followed by a second pass with the computed scaling coefficients to produce the final logits. This meta-reasoning mechanism endows the model with rudimentary structural introspection during generation.
Evaluation: Domain Reasoning, Knowledge Recall, and Quantitative Modeling
Question Answering and Domain Reasoning
X-LoRA's performance is evaluated against the base model (Zephyr-7B-β) across a diverse array of multi-disciplinary scientific queries in mechanics, protein science, materials, and chain-of-thought logic. The scaling patterns observed demonstrate highly structured, context-specific gating of adapters with pronounced layerwise heterogeneity.
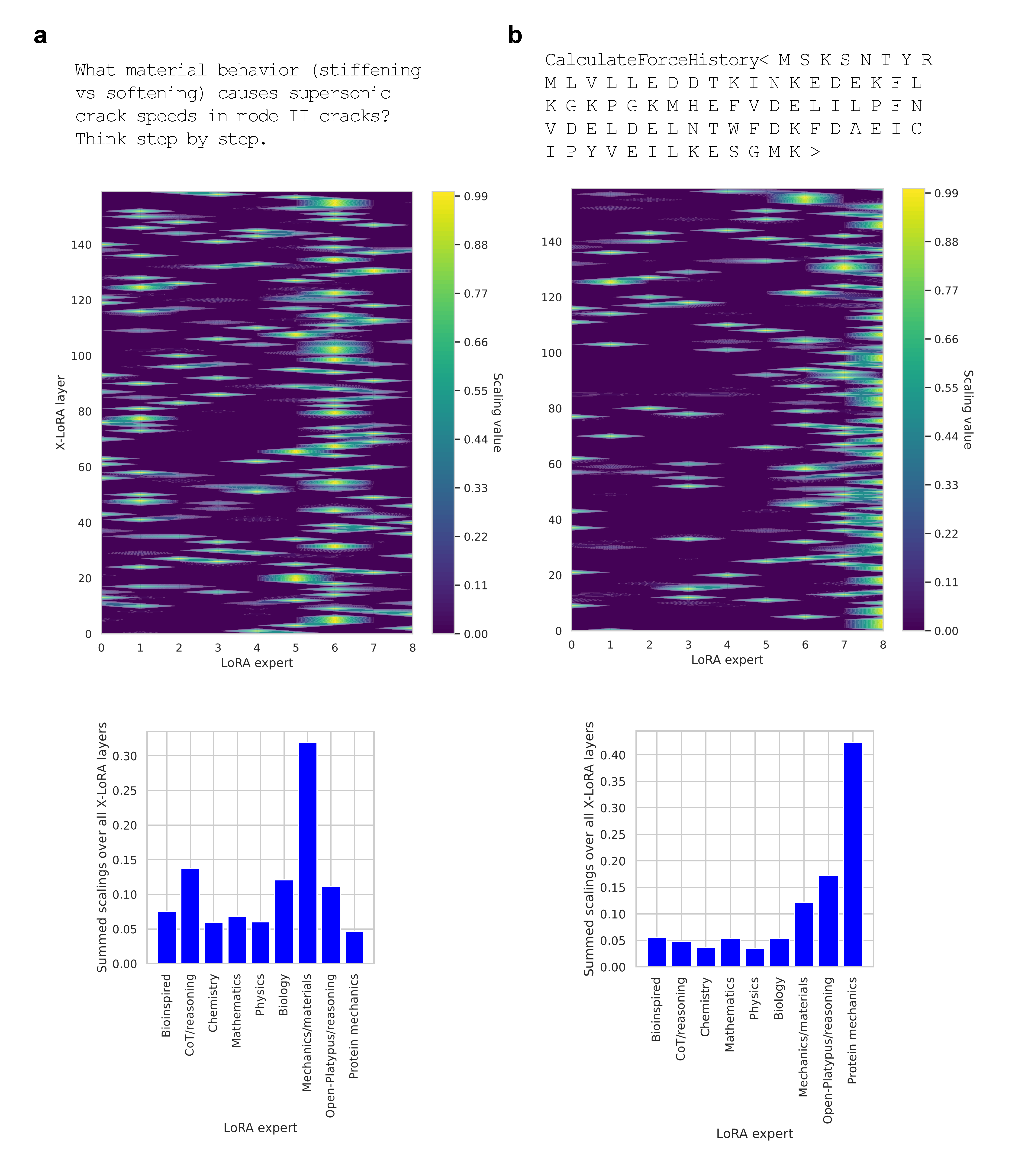
Figure 2: Task-dependent scaling weight distributions show selective activation of pertinent expert adapters, e.g., mechanics for fracture questions and protein mechanics for sequence-based tasks.
Generalization to complex, cross-domain questions is marked by heterogeneous mixing of experts, with notable sparsity. Some queries elicit activation from several specialized adapters across different layers, reflecting the model's ability to blend reasoning modes and scientific domains as required.
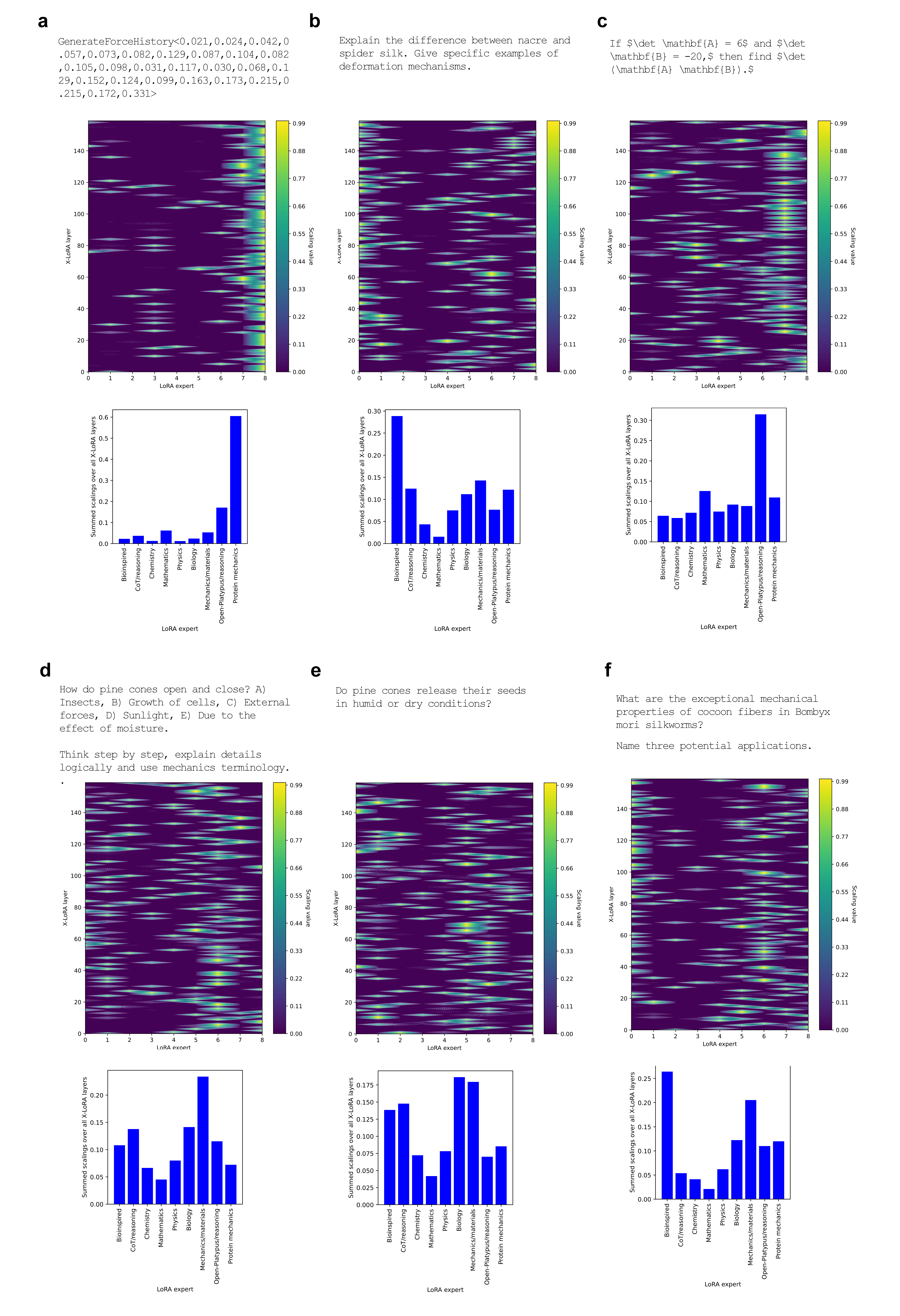
Figure 3: Multiple task examples across different domains, illustrating fine-grained, dynamic gating of expert adapters depending on prompt content.
Token-wise analyses reveal adaptive shifts in adapter utilization during multi-turn reasoning and iterative problem-solving, particularly in long sequences and agentic conversation flows.
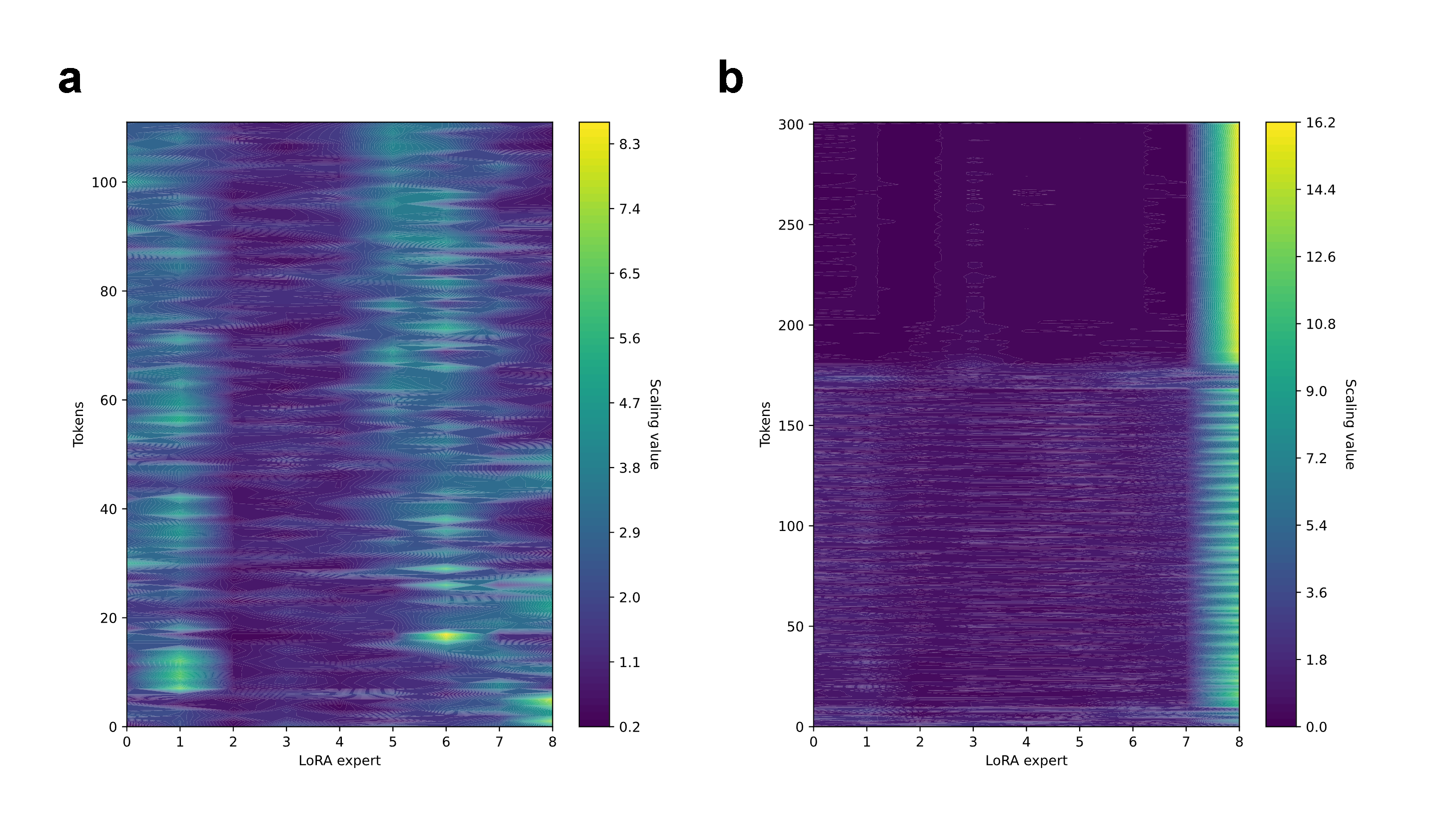
Figure 4: Temporal evolution of adapter usage; notable shifts in expert prominence as the context changes (e.g., during an extended protein design session).
Knowledge Recall Benchmarks
In direct comparison with prior domain-specialized LLMs, X-LoRA achieves superior knowledge recall in a bioinspired materials exam despite using a much smaller parameter base (7B vs 13B parameters). Further, X-LoRA's performance subsumes that of the base model in mechanics/materials knowledge and custom QA evaluations.
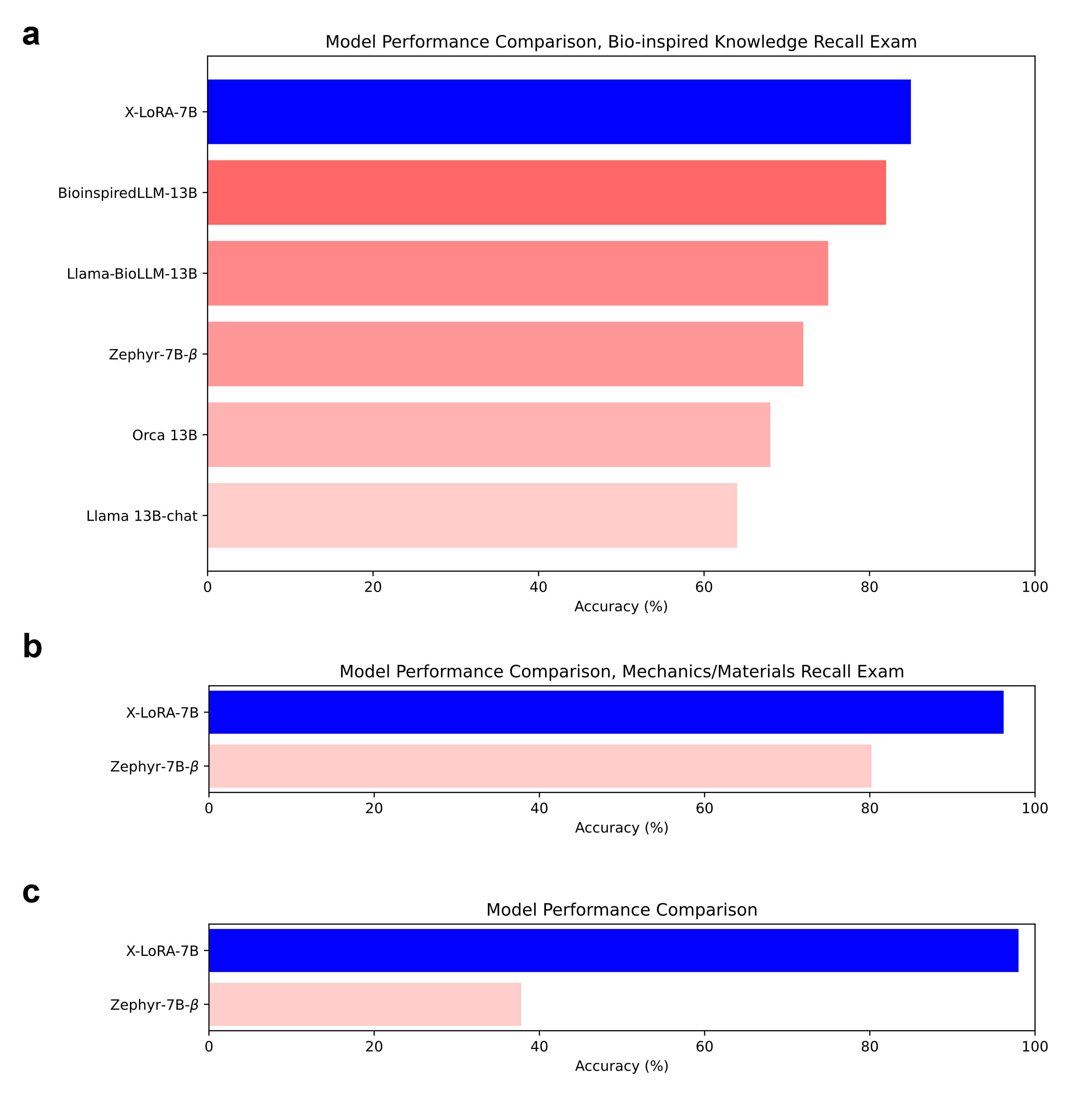
Figure 5: Knowledge recall comparisons indicate that X-LoRA outperforms larger, monolithic models and its own base model in targeted scientific assessments.
Scientific Text-to-Image and Generative Design
When tasked with generating DALL-E prompts for scientific image synthesis, X-LoRA produces precise, focused input resulting in more accurate downstream image generation than the foundation model.

Figure 6: X-LoRA-generated DALL-E prompts yield concise, accurate image synthesis aligned to task constraints, highlighting improved domain task understanding.
Quantitative Protein Property Prediction and Design
X-LoRA exhibits strong quantitative predictive accuracy in protein mechanics: it predicts force-extension profiles, unfolding force, and energy from sequence with R2=0.85, outperforming prior models and rivaling direct molecular simulation pipelines.
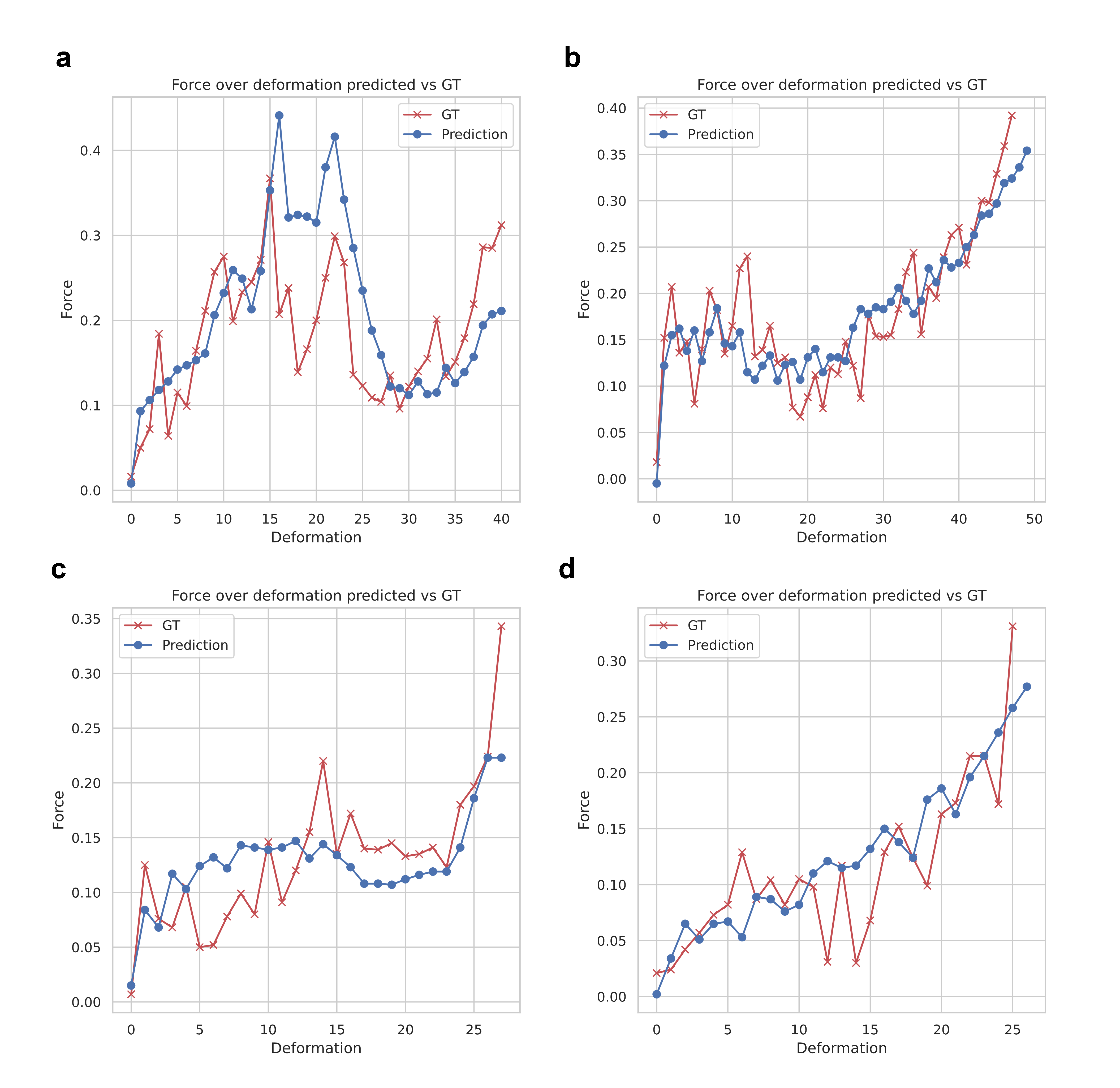
Figure 7: Accurate prediction of protein force-deformation behaviors directly from sequence input for multiple targets.
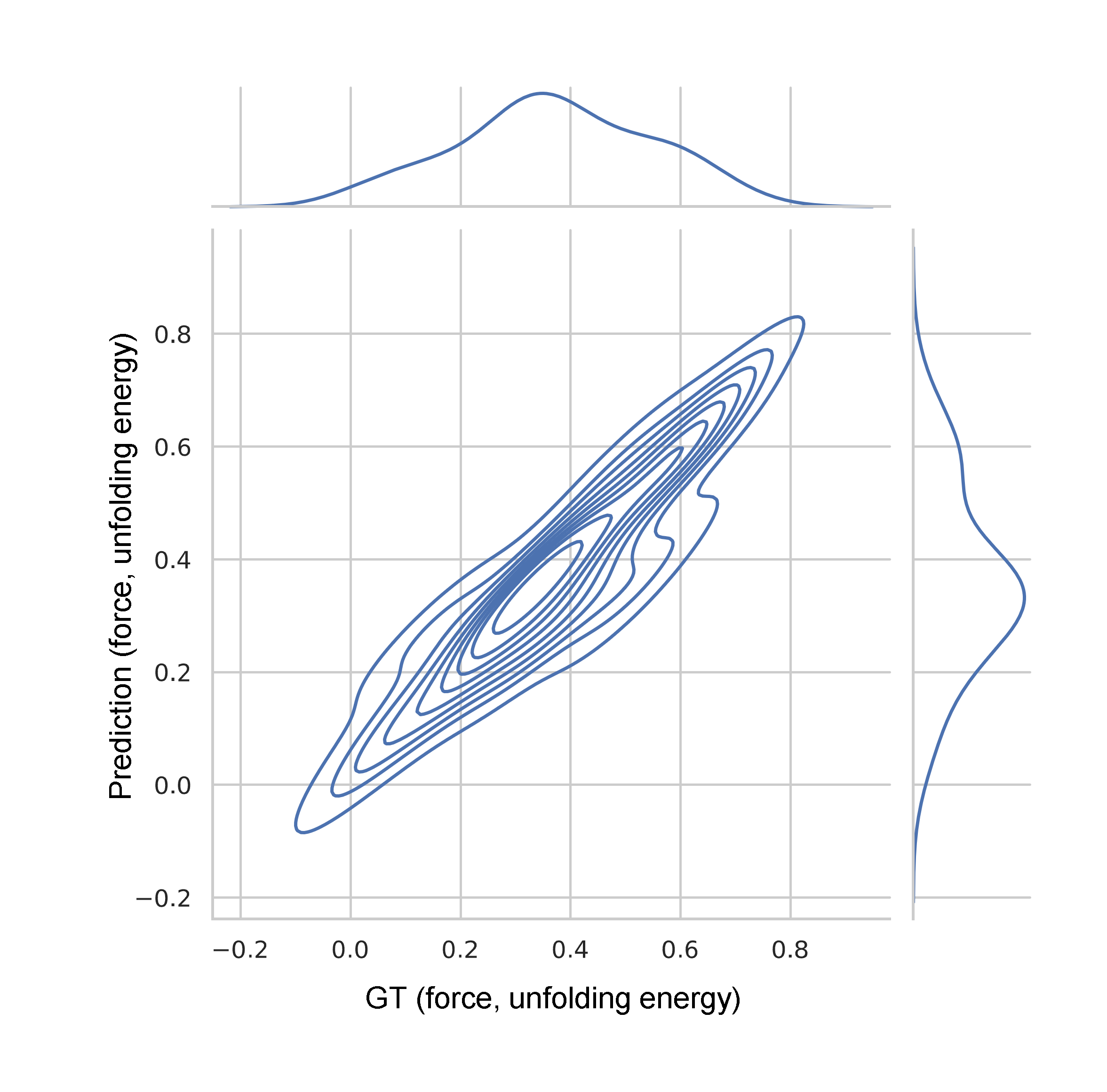
Figure 8: Test set correlation of predicted vs ground truth unfolding force and energy; R2=0.85.
The model also demonstrates robust cycle-consistency in inverse design: prompting for a protein sequence with a specified mechanical profile, X-LoRA generates candidates whose predicted properties closely match the supplied conditions.
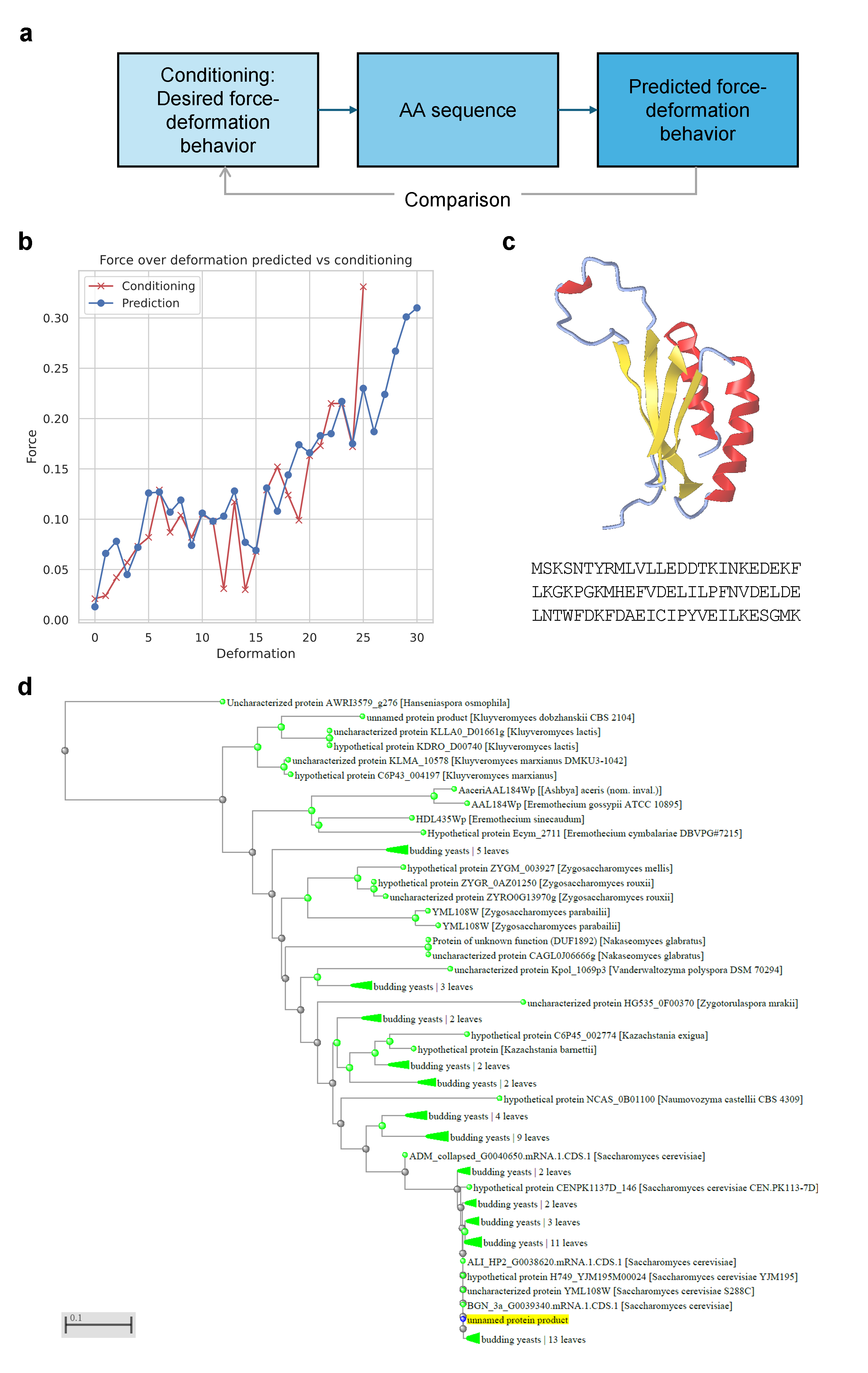
Figure 9: Inverse protein design (targeted force-deformation curve), direct evaluation of generated sequence, folded structure (AlphaFold), and sequence homology analysis.
Multi-Agent, Agentic, and Knowledge Graph Modeling
A distinctive feature is X-LoRA's support for complex agentic workflows: e.g., multi-expert conversational reasoning, protein design analysis, and ontological knowledge-graph construction through staged adversarial dialogue.
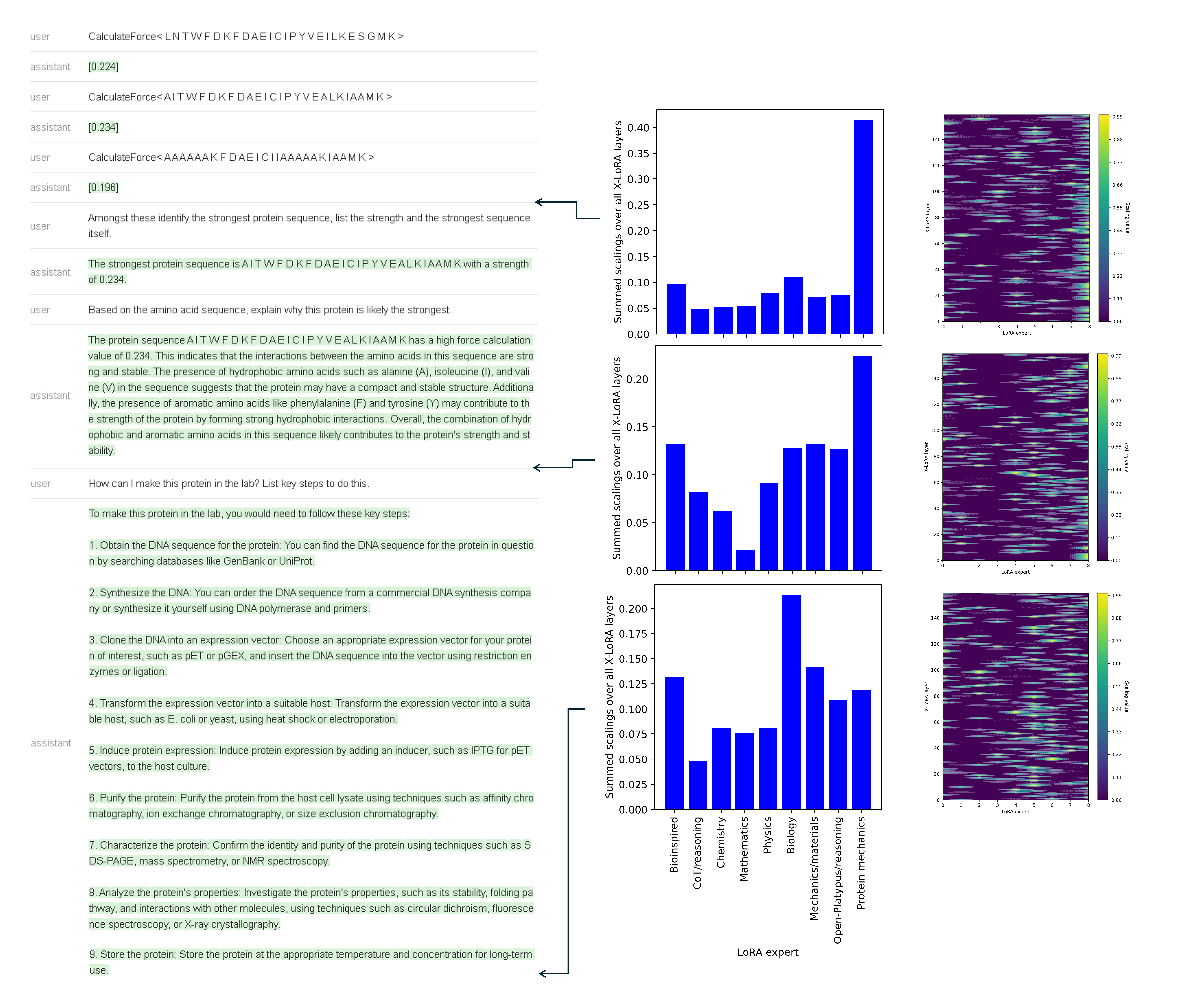
Figure 10: Layerwise expert composition dynamically shifts between protein mechanics, reasoning, and materials/biology adapters as a conversation evolves through design, analysis, and manufacturing steps.
Coherence between model-predicted protein stability and AlphaFold structural confidence is directly observed, confirming the model's mechanistic insight into sequence–structure–property relationships.
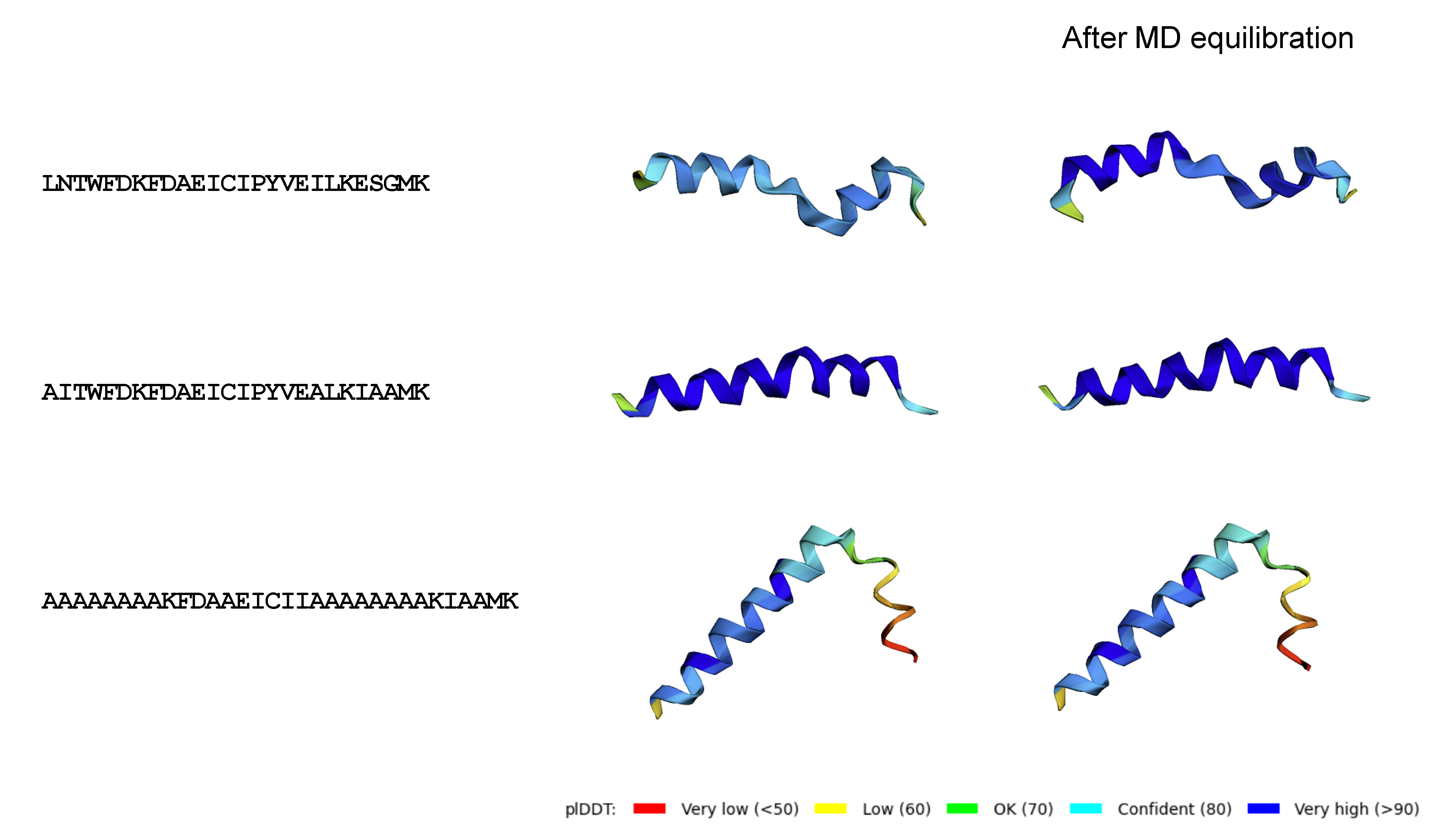

Figure 11: Generated protein 3D structures confirm model predictions regarding stability and secondary structure motifs.
Extended conversations exhibit a shift in the relative dominance of adapters—e.g., reasoning and logic adapters become more active during aggregate explanation and higher-order synthetic or analytic tasks.
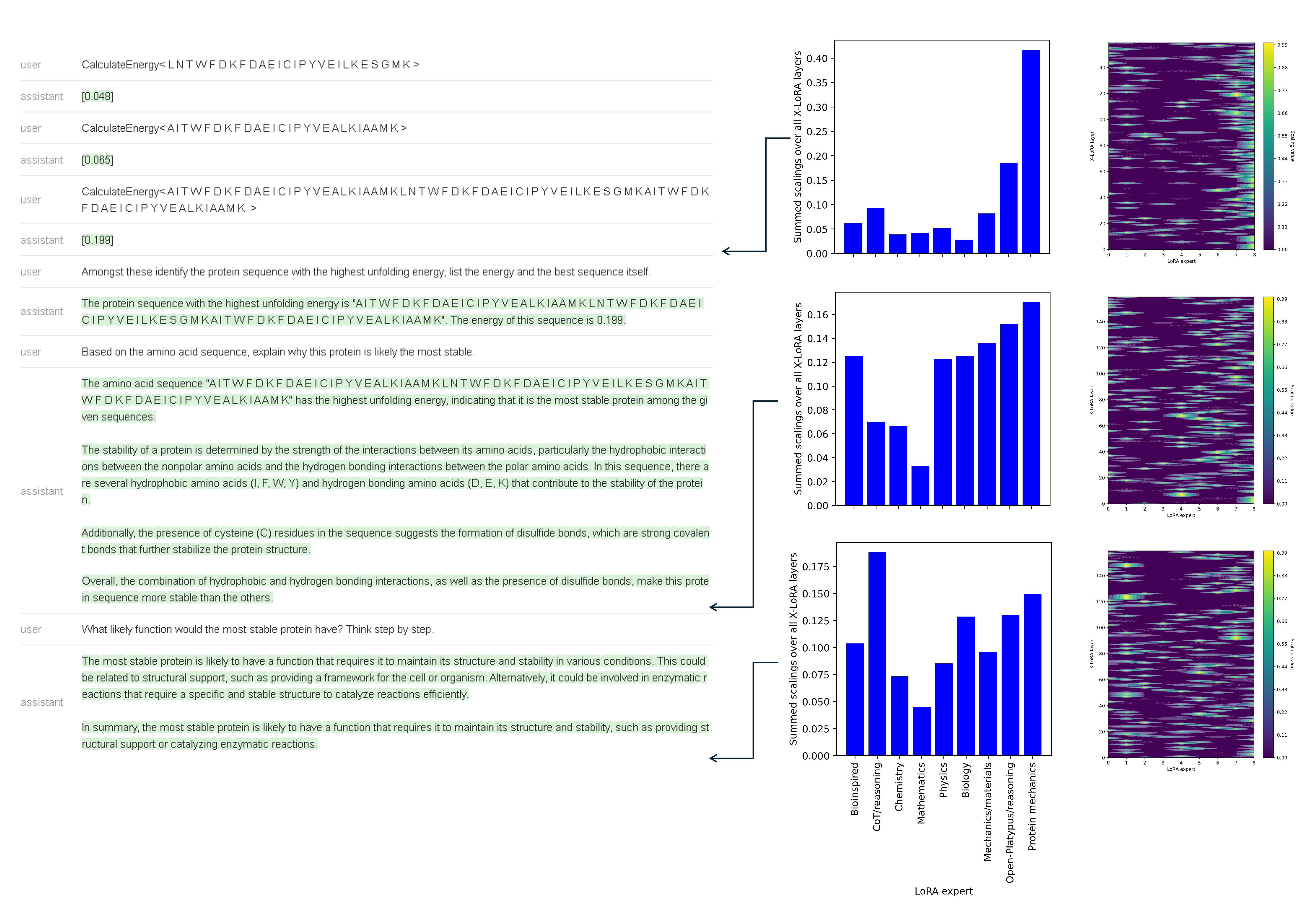
Figure 12: Trajectory of expert adapters as the conversation shifts from analysis to integrative reasoning and hypothesis generation.
Extension to Other Architectures and Property Domains
The methodology generalizes to other backbone architectures. A Gemma-7B-based X-LoRA variant, with adapters spanning protein mechanics and quantum molecular property prediction (QM9), achieves high correlation (R2=0.96) in quantum property regression and demonstrates rapid adaptation to molecular design tasks.
Implications, Limitations, and Future Directions
X-LoRA enables practical assembly of highly task-specialized, cross-domain scientific LLMs with minimal compute overhead and without retraining the foundation model. It provides a scalable path to synthesis of multi-domain expert systems, with direct applicability in protein engineering, scientific QA, molecule design, and other forward/inverse prediction tasks.
However, dual forward passes introduce additional inference cost compared to standard single-pass protocols, and the efficiency of the scaling head depends on careful dataset selection and coverage at the mixing stage. The choice and construction of adapter sets, compositional strategies, and nuanced interactions between adapters represent active areas for further investigation.
From a theoretical perspective, the dynamic, contextually-aware reconfiguration of internal representations via adapter scaling could inform the development of self-modifying or meta-reasoning LLMs and may help clarify principles underlying modularity and functional specialization in large neural systems.
Practical extensions include (a) support for larger/multimodal base models, (b) automated adapter expert discovery, (c) hierarchical and multi-agent coordination, and (d) coupling with high-fidelity simulation or experimental pipelines for closed-loop scientific discovery.
Conclusion
X-LoRA operationalizes a modular, dynamic, and computationally efficient framework for mixing expert specializations within LLMs, establishing strong baseline results in scientific reasoning, structured knowledge recall, protein and molecular design, and agentic integration workflows. It provides a flexible methodology for future specialization, interpretability, and deployment of scientific LLMs across a broad range of data-driven research domains.











