- The paper demonstrates how LLM-driven generative knowledge extraction constructs robust ontological graphs from scientific literature.
- It employs graph-theoretical tools and multimodal synthesis to uncover novel, cross-domain relationships and actionable hypotheses.
- The framework enables automated biomaterials design through algorithmic graph traversal and isomorphic mapping of interdisciplinary knowledge.
Generative Knowledge Extraction and Graph-Based Multimodal Reasoning for Scientific Discovery
This work presents an integrated generative AI framework for the automated extraction, relational structuring, and multimodal reasoning over scientific knowledge with an emphasis on biological materials. The authors stitch together large-scale LLM-driven generative knowledge extraction, formation of a comprehensive ontological knowledge graph, and multimodal graph-centric reasoning to establish a data-driven, transdisciplinary discovery paradigm. Of particular technical emphasis is the use of graph-theoretical tools—embedding, community structure, centrality, and isomorphism detection—coupled with LLM-based reasoning and prompting to transition from basic “information” (facts, relationships, mechanistic summaries) to actionable, synthetic “knowledge,” capable of hypothesis generation, interdisciplinary design, and prediction.
The workflow meticulously addresses the inaccessibility of deep procedural or mechanistic knowledge in the scientific literature (the “how”), as opposed to mere factual retrieval (“what,” “where,” “when”). The authors demonstrate that graph reasoning, traversals, and isomorphism mapping, when combined with LLM-based multimodal synthesis, enable the derivation of nontrivial, sometimes cross-domain, analogies and innovation pathways in automated materials design and theory formation.
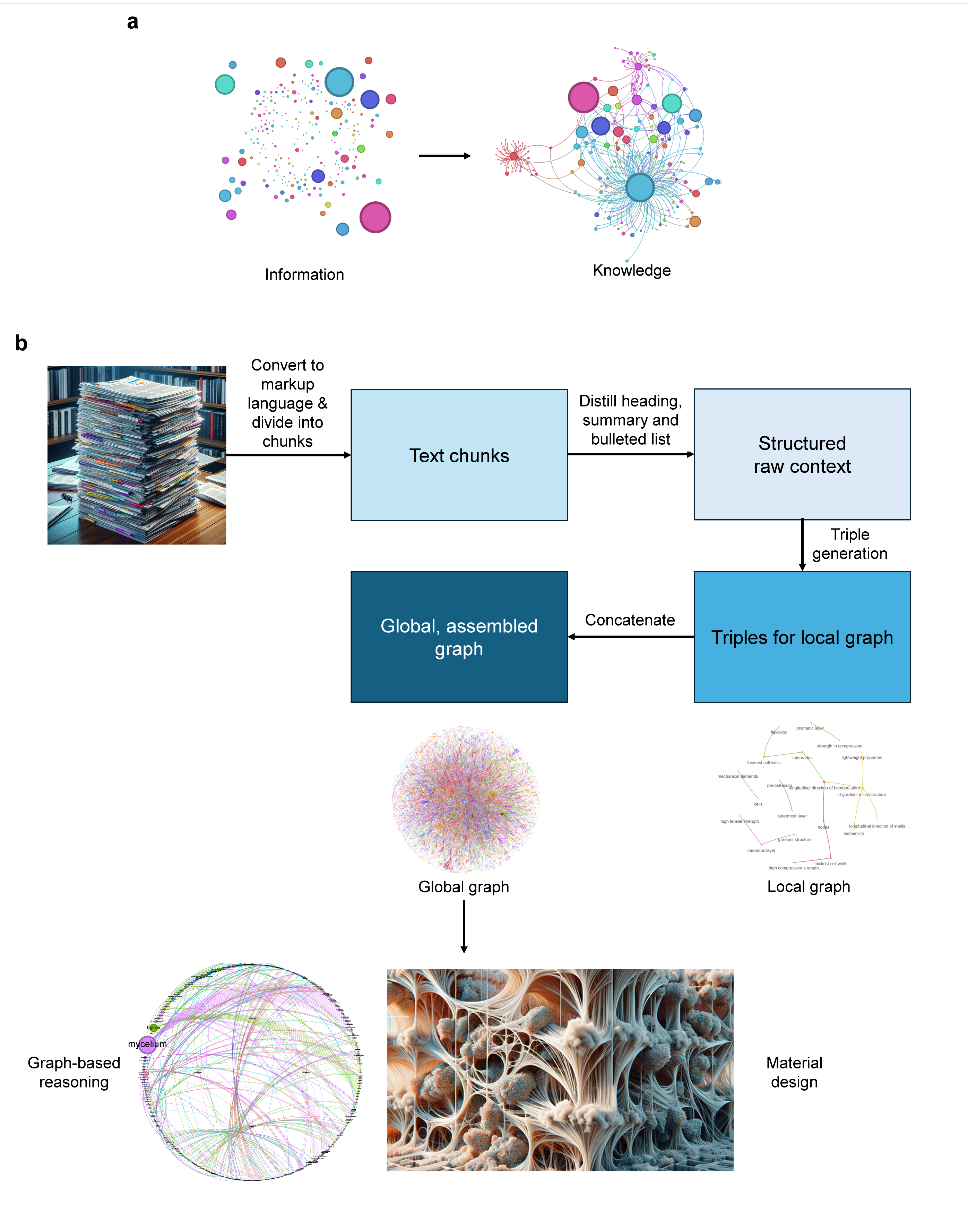
Figure 1: Overview of the graph-based scientific knowledge extraction and reasoning pipeline, bridging the gap from raw information to high-level, transferable knowledge representations suitable for generative reasoning and design.
Construction, Analysis, and Statistical Properties of the Scientific Knowledge Graph
The pipeline processes 1,000+ scientific papers in biomaterials, extracting contextual triples (subject-predicate-object) to construct a global ontological knowledge graph, subsequently refined through node embedding-driven semantic merging, component filtering, and community detection algorithms. This framework results in a large, sparse, highly connected and structured knowledge network suitable for both symbolic and embedding-based reasoning.
Structure analysis demonstrates the network is in the highly connected, low-density regime, and features scale-free properties, as reflected by a power-law degree distribution with exponent α=2.88 (standard error =0.07; log-likelihood ratio R=4.15 versus exponential; p=3.29×10−5). Modularity analysis uncovers 80-109 strongly defined community clusters with high within-community edge density, corresponding to specialized conceptual domains (collagen, biomineralization, mechanical strength, etc.).

Figure 2: Multi-scale visualization of the global knowledge graph, demonstrating both broad and localized connectivity with highlighted central concept nodes.
High-degree hub nodes (e.g., “collagen microfibrils,” “mechanical properties,” “hydroxyapatite,” “biological materials”) serve as bridges for long-range information flow and enable low-diameter navigation and inference. Degree and clustering coefficient analyses indicate network heterogeneity, with select communities showing tight local coupling, while betweenness centrality analysis identifies critical cross-domain connector nodes. The scale-free architecture is algorithmically exploited for efficient traversal and combinatorial path sampling.
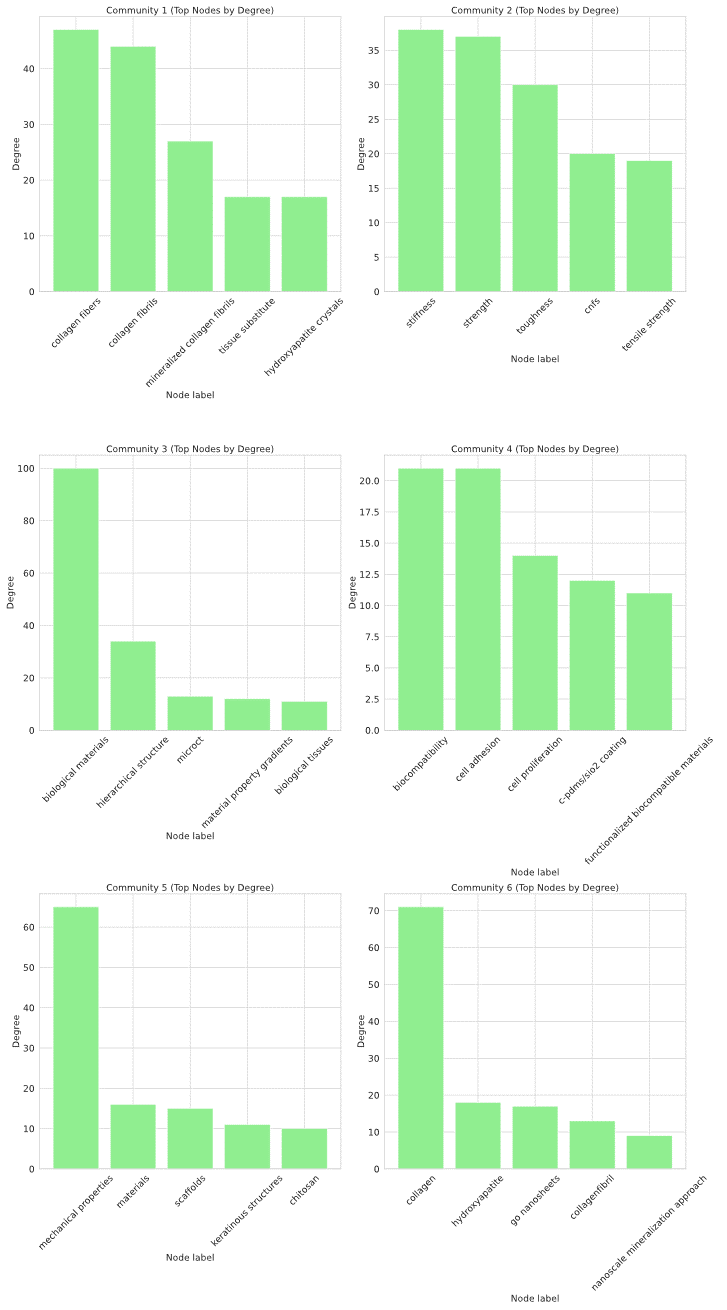
Figure 3: Community-level structural statistics, including node degrees, clustering coefficients, and modularity, illustrating the emergence of hub-centric, modular knowledge clusters.
Generative Path Sampling and Graph Traversal for Hypothesis Generation
The generative framework leverages deep LLM embeddings (e.g., BAAI-bge-large-en-v1.5) for node representation and similarity search, enabling combinatorial path sampling between otherwise unrelated scientific concepts. Cosine similarity-based node ranking and path finding algorithms uncover both direct and high-order, transitive mechanistic chains, facilitating the transition from factual linkage to hypothesis generation.
For example, the work demonstrates discovering multi-step connective pathways from “graphene” to “silk” (via “mechanical strength” and “biological materials”) and from “inkjet-based bioprinting” to “spider silk protein” (traversing through multi-hop semantic and relational transitions). This approach enables the detection of implicit, literature-spanning relationships not present in any single primary source.
Multimodal Reasoning and Query-Driven Design via Graph-LLM Interactions
A key innovation is the use of combinatorial traversal-derived context as input for complex, prompt-based LLM reasoning. By augmenting prompts with specific ranked path samples, the system induces models (including open-source and proprietary LLMs such as X-LoRA, Mixtral, GPT-4) to critically reason about scientific relationships, extrapolate technical synthesis/design hypotheses, and propose novel, testable materials concepts.
In a notable demonstration, LLMs were queried across four combinatorially distinct paths connecting “a flower” (morphological inspiration) to "nacre-inspired cement" (bio-composite system), with models synthesizing design hypotheses involving integration of chitosan, PEGDMA, and hierarchical nanostructuring, sometimes referencing concrete chemical mechanisms (intermolecular interactions, self-assembly, supramolecular inclusion complexes, etc.).
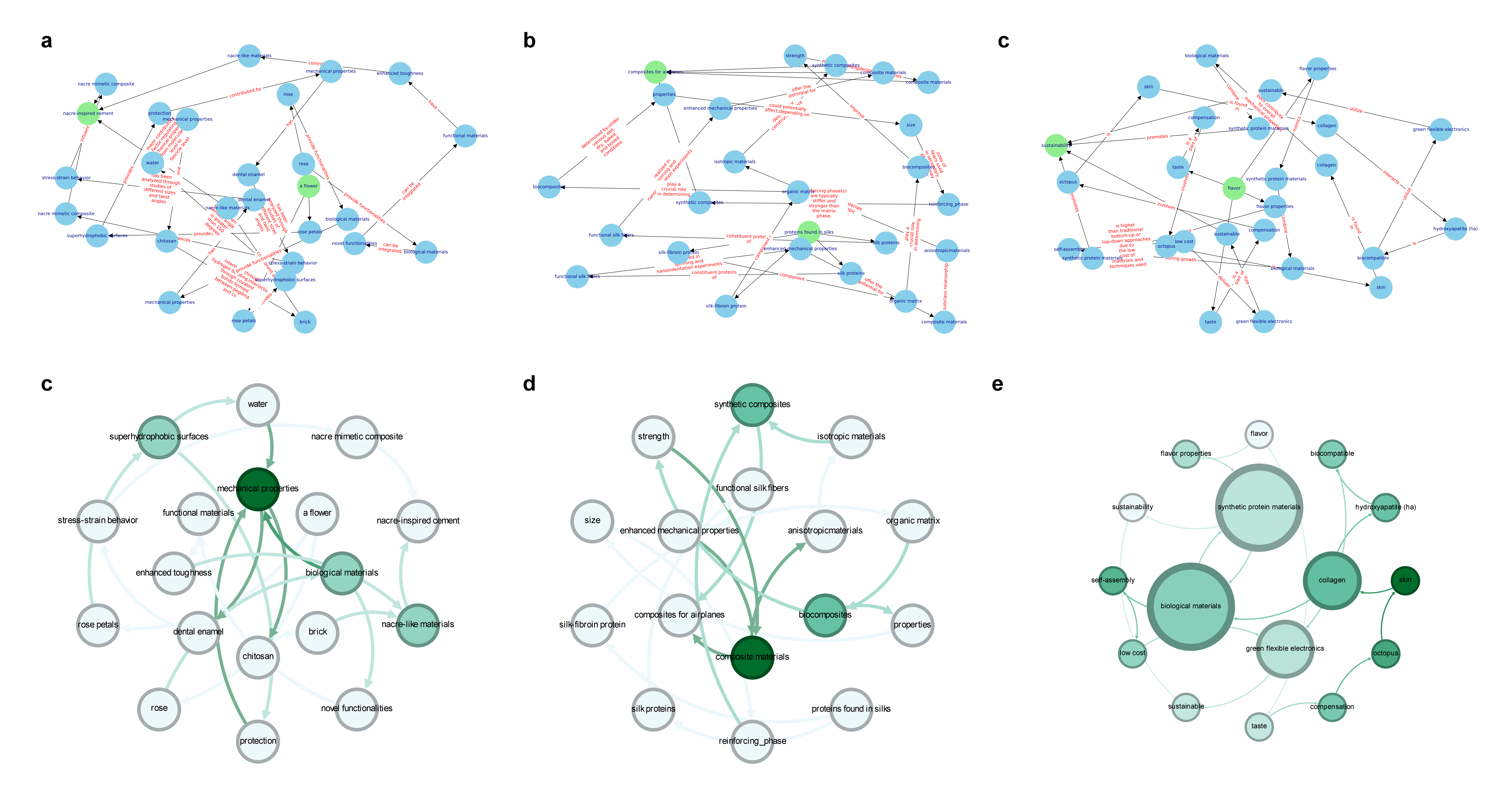
Figure 4: Visualization of multi-path graph sampling and merged traversals between conceptually distant nodes, highlighting emergence of high-degree, functionally relevant hubs and novel synthetic pathways.
Isomorphic Mapping and Cross-Domain Knowledge Transfer
A strong and bold assertion of the paper is the use of isomorphism analysis on subgraphs derived from disparate scientific and artistic corpora, including biological materials and the formal structure of Beethoven’s 9th Symphony. By mapping structurally isomorphic graph motifs, the authors claim that “deep” analogies or design heuristics can be extracted even when no node or entity overlap exists, moving beyond simple analogy toward algorithmic, mechanism-independent transfer of functional principles. The semantic mappings produced span structural organization (e.g., adhesive force ↔ tonality), failure modes ↔ musical resolution, and system-wide flow properties.
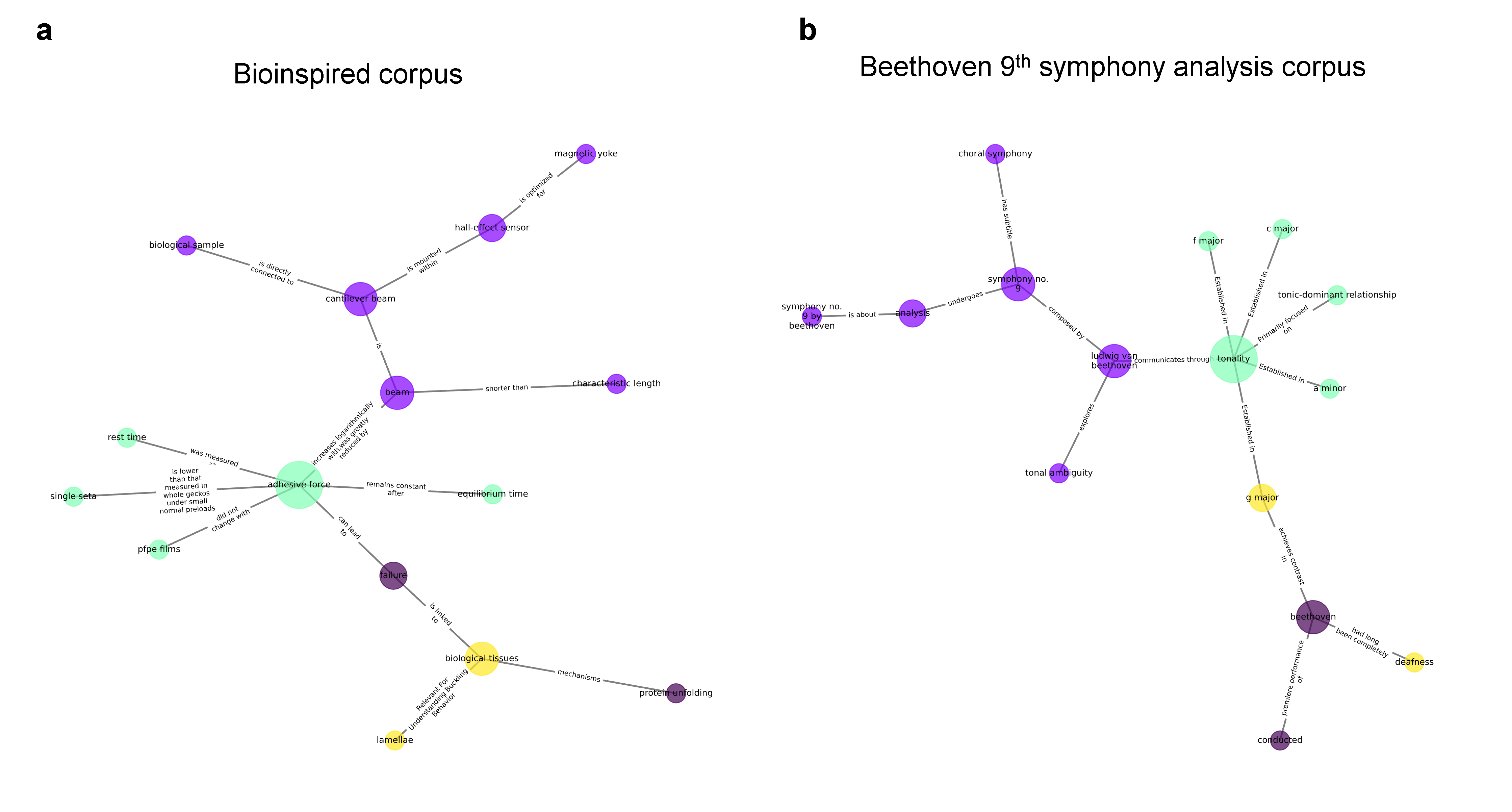
Figure 5: Structural isomorphism detected between biological materials and Beethoven’s 9th Symphony, purportedly supporting cross-domain analogy and abstraction in scientific reasoning.
Augmentation via Self-Generated, Agentic, and Literature-Ingested Knowledge
The pipeline supports both continual, autotelic knowledge graph augmentation—via:
- Model-self-generated experimental results (e.g., LLM-driven protein stability prediction).
- Agentic, adversarial multi-agent systems performing open-ended question–answer cycles (e.g., food/mechanics intersections, safety/sustainability/texture/flavor of synthetic protein foods).
- On-demand ingestion of external literature or direct experimental pipelines.
For example, two X-LoRA agents iteratively explore the design, safety, and societal/functional implications of introducing synthetic protein mechanics into food systems. The resulting adversarially-generated dialog is ingested for graph augmentation, and subsequent graph traversal reveals machine-identified cross-domain consensus research opportunities.
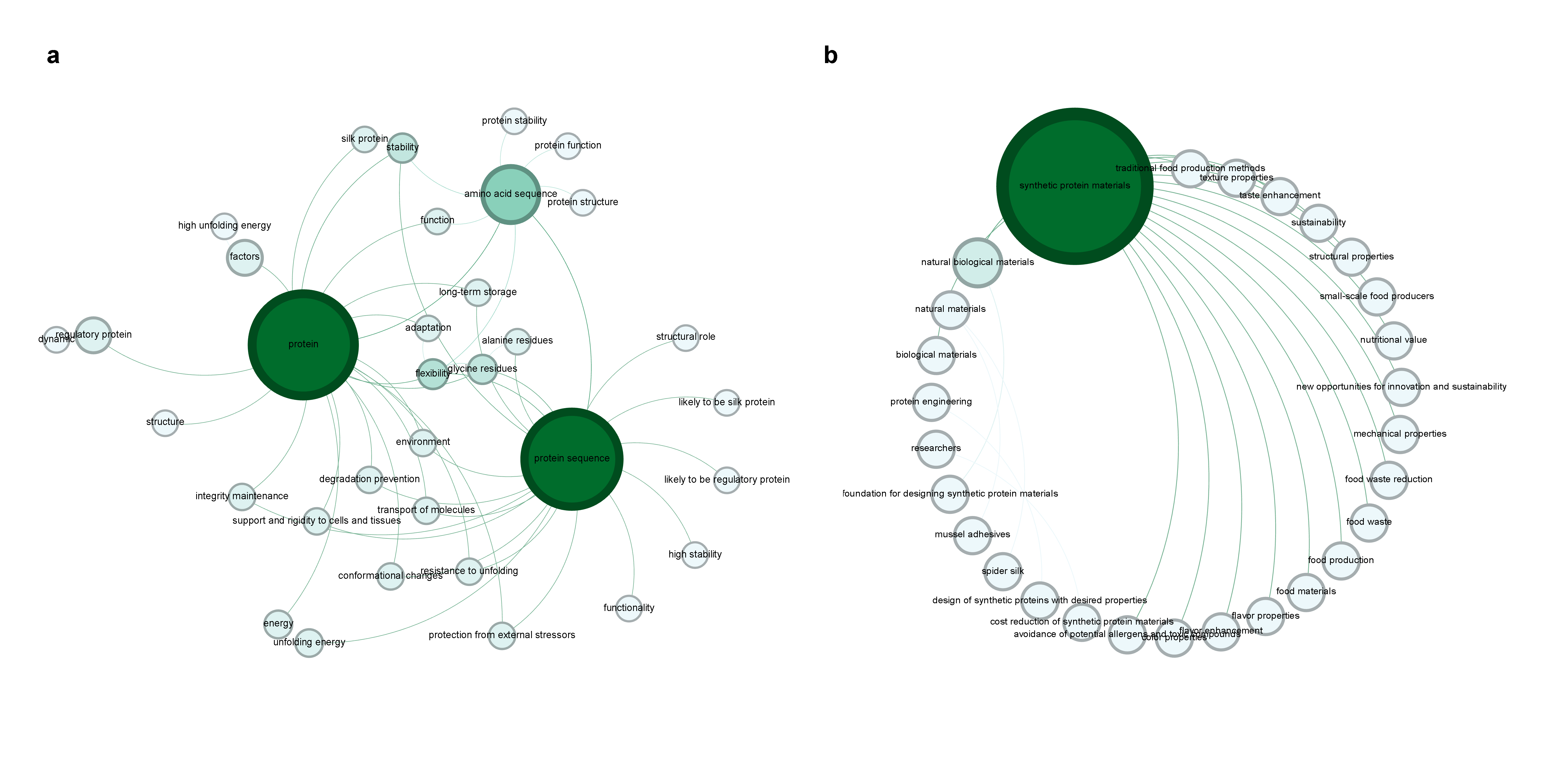
Figure 6: Expanding the graph through new data sources, with autonomous agentic modeling and self-generated protein/mechanics predictions facilitating knowledge base growth.
Concrete AI-Driven Materials Design Examples: From Scientific Synthesis to Artistic Integration
A technically ambitious sequence uses the graph-traversal-derived context and multimodal models to jointly interpret abstract paintings (e.g., Kandinsky’s “Composition VII”) and synthesize entirely new material microstructures—e.g., hierarchical mycelium-collagen composites with specified chemical, nanostructural, and mechanical features. The generative pipeline couples GPT-4V’s vision-language reasoning with generative image synthesis models (DALL·E 3), translating aesthetic, spatial, and organizational cues from the artwork into detailed, explicitly actionable material design blueprints.
Implications, Limitations, and Prospects for AI-Augmented Scientific Discovery
The approach demonstrates that LLM-powered, graph-structured reasoning achieves three key objectives:
- Transcending domain boundaries (e.g., music → mechanics, food → protein assembly) via structural and semantic isomorphism.
- Algorithmic generation of experimental hypotheses and novel material blueprints from “in-the-wild” literature and model-generated data.
- Multimodal design and reasoning, capturing both quantitative and qualitative features from diverse data sources (scientific literature, visual art, numerical simulation).
The key claims are: the graph exhibits robust scale-free properties (power-law exponent α=2.88 with statistically significant model comparisons); graph-centric reasoning enables the discovery of nontrivial scientific relationships not present in isolated literature sources; and multimodal integration expands the generative capacity of AI for scientific discovery well beyond conventional retrieval or QA.
Limitations include constraints imposed by the computational scale of graph generation, node disambiguation, embedding model adequacy, and current context/intelligence limitations of LLMs (including hallucinations, lack of domain-specific knowledge, or erroneous analogies). Nonetheless, the architecture is modular, permitting rapid expansion to new corpora, materials domains, and reasoning tasks as model capacity and data quality improve.
Conclusion
This work articulates an comprehensive, modular pipeline for the automated, large-scale, LLM-driven extraction, structuring, and reasoning over scientific knowledge graphs. The combination of scalable, structurally precise graph-theoretic analysis and advanced LLMs for multimodal reasoning and generation enables step-changes in autonomous hypothesis generation, cross-domain design, and knowledge transfer—facilitating a data-driven, transdisciplinary approach to scientific discovery and materials design. The approach is extensible to a diversity of scientific areas, with direct implications for open-ended hypothesis automation, theory formation, and AGI-augmented scientific practice.





