- The paper presents a CNN-based solution using 2D image representations for GCMS data, achieving a competitive second-place log loss score.
- It employs HRNet-w64 with innovative mass and time normalization and dynamic time resizing to enhance feature extraction across varying temperature ramps.
- Grad-CAM++ visualizations improve explainability by highlighting key mass-to-charge regions related to hydrocarbon and mineral detection.
"Mars Spectrometry 2: Gas Chromatography - Second Place Solution" Summary
Introduction
The paper outlines the solution for the "Mars Spectrometry 2: Gas Chromatography" challenge. The objective was to develop a model capable of processing gas chromatography-mass spectrometry (GCMS) data. The task was framed as a supervised multi-label classification problem, with nine binary non-exclusive target labels per sample. The performance metric used for evaluation was log loss, optimizing for lower values. This essay provides an in-depth examination of the methodology applied to attain the second-best score in the competition.
Methodology
The approach involved leveraging two-dimensional, image-like representations of GCMS data samples. Convolutional Neural Networks (CNNs) served as the backbone for the solution, specifically adapted to handle the data's image-like format. Pre-trained CNN models from the timm package, particularly HRNet-w64, were integral to the success of the solution. Importantly, the models employed both mass normalization and time normalization techniques to enhance feature extraction from 2D images.
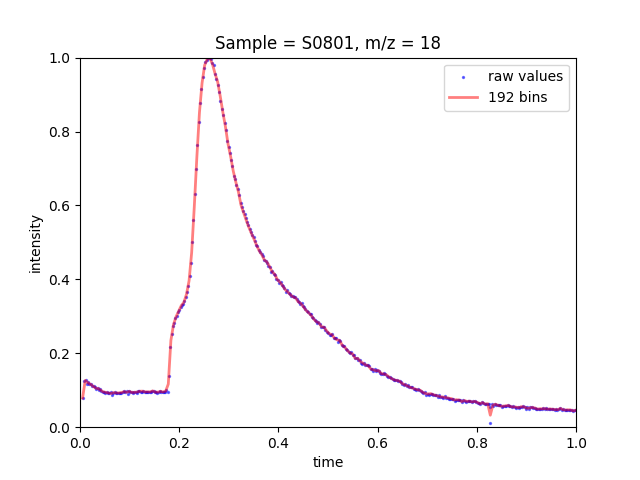
Figure 1: Raw intensity values for sample S0801 at m/z=18.
Image Conversion and Model Training
Data samples were initially converted from CSV files into 2D images with distinct image channels representing mass and time dimensions. This conversion enabled the application of advanced image processing techniques. The model’s innovation lay in dynamic time resizing during both training and inference, facilitating the construction of a robust predictive framework adaptable to varying temperature ramp rates across samples. The incorporation of test-time augmentation (TTA) further augmented model capabilities to handle variations in the time axis effectively.
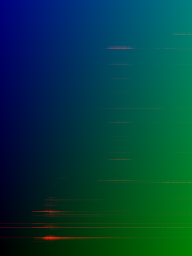
Figure 2: Sample S0801 converted to 2D representation and saved into the red channel, where y-axis is mass rows (0≤m≤255) and x-axis is time columns (0≤t≤191).
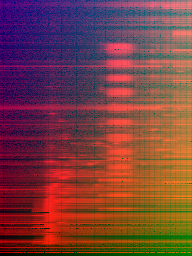
Figure 3: Same as in Fig.~\ref{fig2} but divided by maximum column values (mass-normalization).
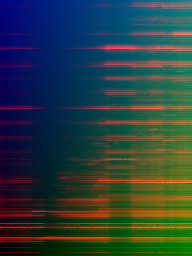
Figure 4: Same as in Fig.~\ref{fig2} but divided by maximum row values (time-normalization).
A critical adjustment during preprocessing was to average time dimensions selectively, which minimized the loss of temporal data granularity crucial for identifying mass spectrometry patterns. The overall model architecture prioritized retaining existing structural components while integrating custom heads for enhanced classification outcomes.
Interpretation and Explainability
Explaining model decisions was facilitated using the Grad-CAM++ method, which provided visualizations indicative of model attention across sample dimensions. Such transparency in model reasoning underscored the model's predominant focus on specific mass-to-charge ratios when predicting outcomes, particularly highlighting regions of hydrocarbon and mineral compound presence.
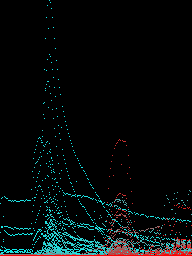
Figure 5: Grad-CAM++ visualisations of samples showing regions of interest for certain compounds.
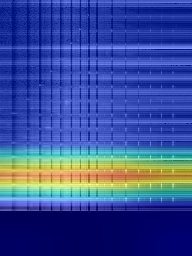
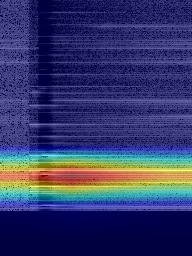
Figure 6: Grad-CAM++ visualisation of two samples containing hydrocarbon compounds.
Implications and Future Directions
The research highlights the importance of incorporating temperature data for improved model accuracy and suggests enhancements like time warping could further refine prediction fidelity. Advances in understanding variabilities across temperature ramps remain critical for achieving progress in this research domain. Additionally, adopting more comprehensive multimodal approaches may enhance future iterations of GCMS data processing within planetary exploration frameworks.
Conclusion
The devised solution for the Mars Spectrometry 2 challenge emphasizes an integral fusion of deep learning with domain-specific GCMS data representation. The rigorous adaptation to GCMS data intricacies, coupled with strategic ensembling and model innovations, delineates a path forward in automatic mass spectrometry data analysis, offering valuable insights for subsequent studies in both terrestrial and extraterrestrial applications.






