- The paper introduces BrainFounder, a two-stage pretraining model that leverages large-scale healthy neuroimaging data to improve segmentation accuracy.
- It employs vision transformers with a SwinUNETR architecture to execute self-supervised learning in 3D brain imaging, achieving superior Dice coefficients.
- The study demonstrates potential clinical impact by effectively generalizing across imaging modalities and handling limited annotated data conditions.
BrainSegFounder: Towards 3D Foundation Models for Neuroimage Segmentation
Introduction
The paper "BrainSegFounder: Towards 3D Foundation Models for Neuroimage Segmentation" (2406.10395) presents an innovative approach in neuroimaging analysis by developing a foundational model for brain image segmentation. Leveraging vision transformers, this study provides a framework for handling medical data, primarily focusing on enhancing the diagnostic accuracy for MRI-based tasks. The central contribution is a two-stage pretraining methodology that utilizes a large-scale dataset from generally healthy individuals to refine model accuracy and segmentation capabilities, surpassing the current state-of-the-art methods.
Methodology
Two-Stage Pretraining Process: The proposed model, referred to as BrainFounder, adopts a comprehensive pretraining regimen that starts with a large-scale anatomical understanding phase using the UK Biobank (UKB) dataset, encompassing 41,400 participants. This dataset provides a diverse array of healthy brain imaging modalities that help form a robust baseline model. The second stage advances this baseline to specialize in pathological data by employing datasets like BraTS and ATLAS v2.0. These stages (Figure 1) are crucial for developing the model's anomaly detection skills, focusing on the distinct geometrical and spatial configurations associated with brain tumors and stroke lesions.
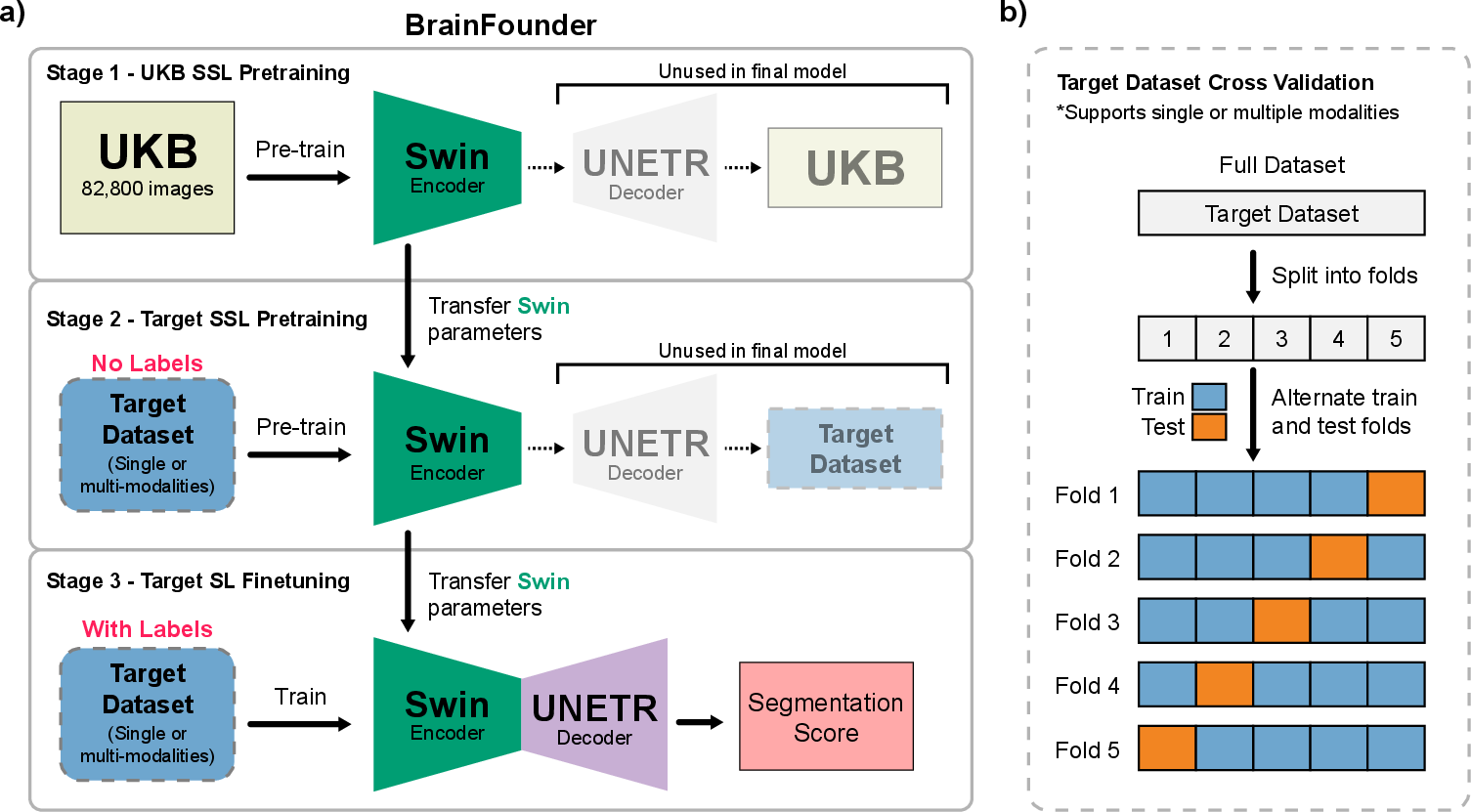
Figure 1: Overall Study Design. a) The two-stage pretraining process using Swin Transformer decoders and encoder. Initially, the model is pretrained on the UKB dataset (Stage 1), followed by the downstream task dataset (Stage 2). b) This is succeeded by fine-tuning on each downstream dataset, with transfer learning applied between each stage.
Architecture: BrainFounder employs a vision transformer-based encoder coupled with an up-sampling decoder, building on the SwinUNETR architecture. This model framework allows effective self-supervised learning across 3D image volumes (Figure 2). The encoder's novel structure facilitates a unique dual-phase pretraining approach that prioritizes the encoding of normal anatomical structures and then refines anomaly detection via additional data encoding, significantly improving predictive capabilities.
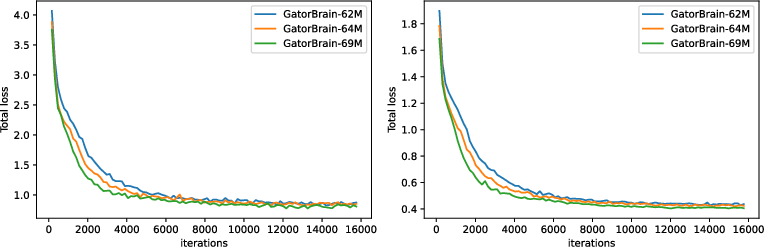
Figure 2: Training (left) and validation (right) loss of Stage 1-pretraining three different scale of BrainFounder models on UKB.
Evaluation and Results
Performance Metrics: The evaluation of BrainFounder against datasets such as BraTS highlights its superior segmentation performance, with the small model (64M parameters) achieving the highest Dice coefficient relative to both the baseline SwinUNETR model and other variants (Table 1). This performance showcases the model's ability to generalize better across various data constraints, achieving significant strides in segmentation accuracy even under limited data conditions (Figure 3).
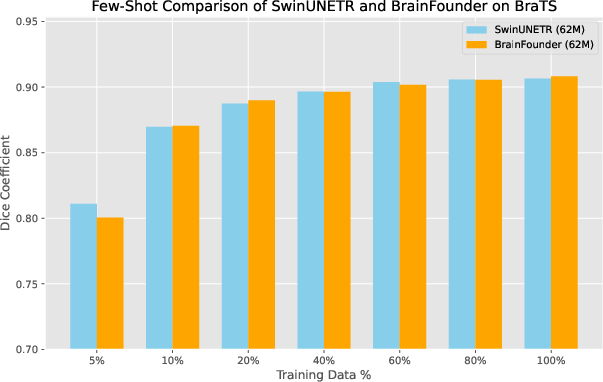
Figure 3: Dice coefficients for baseline (SwinUNETR) and our model across different levels of training data availability.
Modality and Generalization: The study also emphasizes BrainFounder's flexibility across various imaging modalities. Tests involving modality restrictions revealed that BrainFounder outperforms existing models even when constrained to single-modal imaging, validating the framework's extensibility and robustness in diverse clinical scenarios (Table 2).
Discussion
The study posits that BrainFounder represents a substantial development in neuroimaging AI, particularly regarding its ability to maintain high precision and generalization capabilities across disparate datasets. The foundation model structure allows for efficient training, reducing the typical reliance on extensive annotated datasets. This capability suggests potential for widespread adoption in clinical settings, particularly for tasks involving segmentation of brain tumors, stroke lesions, and potentially other complex neurological conditions.
Conclusion
BrainFounder sets a new benchmark in 3D neuroimage segmentation models, combining foundational pretraining on healthy brain imaging with specialized pathological training to deliver a robust, versatile solution. Future research could expand on this foundation model approach to explore broader neurological applications, potentially leading toward a unified AI-driven methodology for comprehensive brain health analysis.


