- The paper introduces a CNN-BiLSTM pipeline that leverages explicit plaque surface edge images for robust detection of the dynamic ‘Jellyfish sign’.
- Using template matching and advanced preprocessing, the method isolates the plaque region to enhance surface-focused features in ultrasound video analysis.
- Experimental validation shows that integrating temporal modeling via BiLSTM boosts performance beyond 80% accuracy, underscoring its clinical potential in stroke risk assessment.
Automated Classification of the Jellyfish Sign in Carotid Plaques Using Deep Convolutional and Recurrent Neural Architectures
Introduction
The presence of dynamic instability in carotid plaques, referred to as the "Jellyfish sign," has notable clinical significance due to its association with increased risk of cerebral infarction. The Jellyfish sign is characterized by surface undulations synchronized with arterial pulsation, as identified in B-mode ultrasound video imaging. Conventional assessment for the sign relies primarily on subjective visual evaluation, leading to variability and suboptimal early detection. This paper addresses the challenge by presenting a method that leverages spatial-temporal deep learning for objective, automated classification of the Jellyfish sign using carotid ultrasound videos.
Methodology Overview
The proposed pipeline integrates spatial feature extraction and temporal dynamics modeling via a two-stage approach: advanced preprocessing, followed by a CNN-BiLSTM based classifier. Key innovation is the explicit use of plaque surface edge images, traced and binarized by a sonographer, and concatenated with plaque video frames to guide the model's focus on the lesion surface.
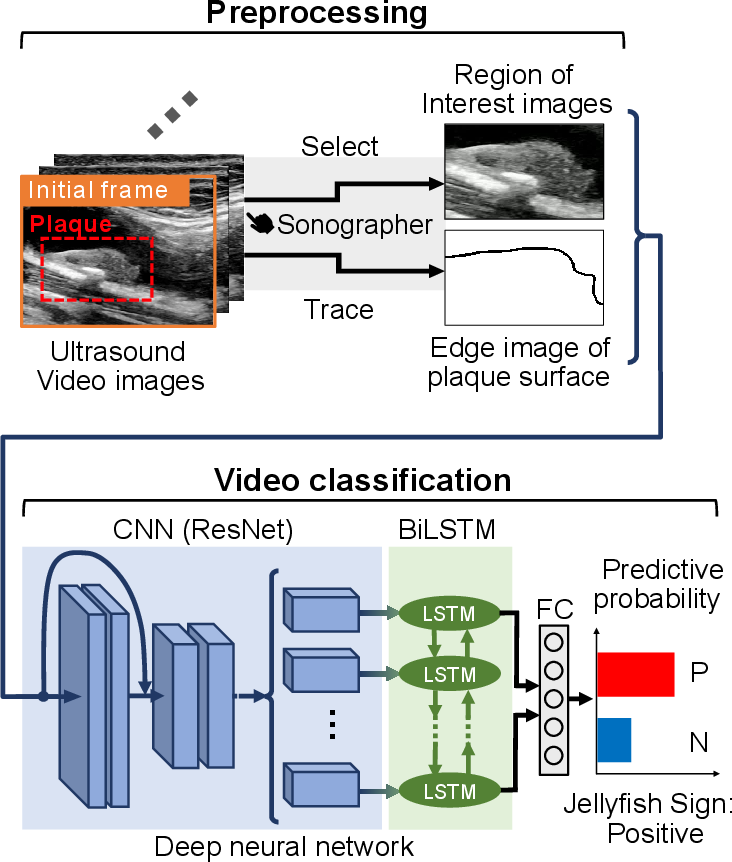
Figure 1: Overview of the proposed method integrating preprocessing and CNN-BiLSTM classification for the Jellyfish sign.
Preprocessing and Template Matching
To enable robust isolation of plaque-specific motion from confounding factors such as vascular pulsation and respiratory movement, template matching is employed. The region of interest (ROI) set by the sonographer serves as the template, and normalized cross-correlation identifies the region in each frame with maximal correspondence to the template, effectively stabilizing the plaque region through the sequence. The plaque surface is emphasized using manual edge tracing and binarization, providing explicit localization of the surface boundary.
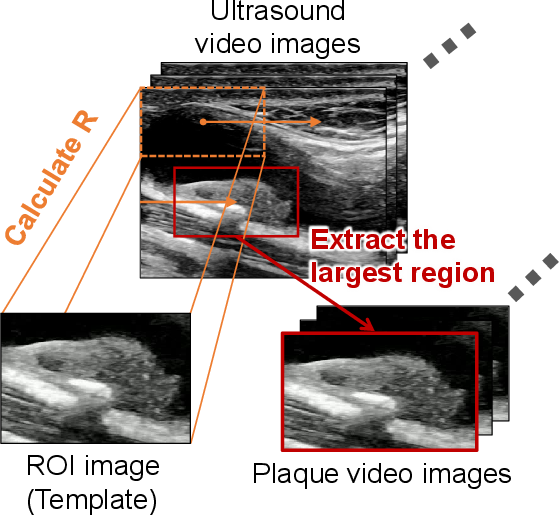
Figure 2: Schematic diagram of the template matching process for plaque-focused video extraction.
Deep Neural Network Architecture
Input videos are structured as two-channel sequences: each frame comprises both the ROI video data and the corresponding binary surface edge mask. The architecture utilizes a ResNet-18 backbone for spatial feature extraction, with output features temporally modeled via bidirectional LSTM cells. ReLU-activated fully connected layers precede a final sigmoid output, providing a probability for Jellyfish sign presence per case. Training optimizes the cross-entropy loss via SGD, with both spatial and temporal data augmentations for regularization.
Experimental Validation
Dataset and Protocol
A balanced dataset of 200 clinical cases (100 positive, 100 negative for the Jellyfish sign) was curated, with videos covering approximately two heartbeats and annotated by specialists. The network evaluation employs 5-fold cross-validation, reporting the mean values across three random initialization seeds for accuracy, precision, and recall.
Ablation Studies
A thorough ablation strategy quantifies contributions from plaque surface image integration and temporal modeling. Performance was evaluated for: (1) input without plaque surface image, (2) CNN only (spatial features, no temporal modeling), (3) CNN-LSTM (unidirectional recurrence), and (4) the full CNN-BiLSTM model.
Impact of Plaque Surface Image
Incorporation of explicit edge images improved all metrics, with test accuracy rising from 77.8% (without edge) to 80.5% (with edge), precision from 78.5% to 81.0%, and recall from 77.3% to 80.2%. This supports the hypothesis that targeted surface features enhance identification of the Jellyfish sign, which is fundamentally a surface-level phenomenon.
Impact of Temporal Modeling
Temporal modeling is critical given the sign's pulsatile dynamics. The CNN-FC baseline, lacking temporal context, is outperformed by CNN-LSTM (accuracy: 78.7%) and by the bidirectional CNN-BiLSTM (accuracy: 80.5%). BiLSTM extends modeling to both past and future context, maximizing capture of periodic motion essential for classification.
Interpretability and Class Activation Visualization
Grad-CAM++ analysis of the ResNet features reveals strong localization of informative regions near the plaque surface in many positive cases, demonstrating alignment with expert-annotated lesion regions. Nonetheless, non-surface interior regions sometimes show notable activations, indicating that clinically relevant dynamics extend beyond the plaque surface, meriting further investigation.
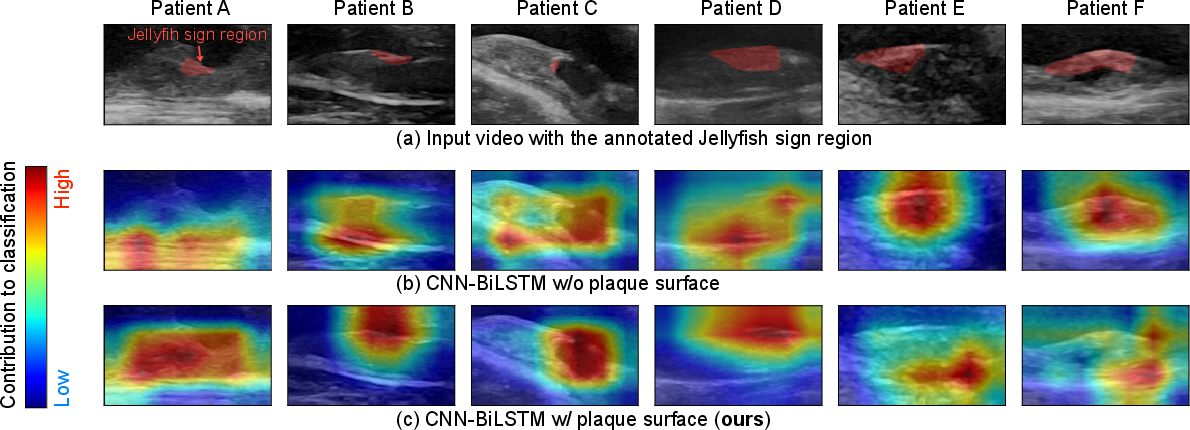
Figure 3: Visualization of activation maps, with strong correspondence to sonographer-annotated Jellyfish sign regions.
Discussion and Implications
The integration of convolutional and bidirectional recurrent neural architectures, guided by explicit surface boundary information, demonstrates substantial performance gains for video-based detection of dynamic pathological signs. The strong numerical results highlight the utility of combining domain-specific annotations with deep learning, moving beyond purely end-to-end approaches.
Clinically, the approach offers a scalable, automated mechanism for early detection of high-risk plaque instability, with potential impact on stroke prevention and surgical planning. The method's modularity allows adaptation to other dynamic ultrasound biomarkers, and its reliance on surface-centric annotations suggests avenues for refining edge extraction with automated segmentation or active learning.
Theoretically, the findings support continued development of hybrid spatial-temporal neural architectures, especially those leveraging expert knowledge for inductive bias. The observed importance of both explicit edge features and bidirectional recurrence underscores the need for architectures that can flexibly encode multi-scale, domain-specific priors.
Future research could explore attention-based feature integration, fusion with physiological signals (e.g., synchronized ECG), and extension to multimodal vascular risk assessment.
Conclusion
The presented method achieves a robust, automated, and quantitative approach for Jellyfish sign classification from carotid ultrasound videos, attaining over 80% accuracy through explicit plaque surface modeling and bidirectional temporal analysis. The clinical utility for risk stratification and theoretical implications for integrating expert annotation into neural architectures are substantial, directing future work toward further feature enrichment and broader dynamic biomarker classification.


