- The paper demonstrates that LSSF-Net integrates conformer-based focal modulation with self-aware and global spatial attention to achieve high segmentation accuracy using just 0.8M parameters.
- The model utilizes innovative components such as CFMA, SAB, and split channel shuffle to enhance both local and global feature extraction in a resource-efficient manner.
- Experimental evaluations on multiple datasets confirm LSSF-Net's robust generalization, computational efficiency, and potential for mobile deployment in computer-aided diagnosis.
LSSF-Net: Lightweight Segmentation with Self-Awareness, Spatial Attention, and Focal Modulation
The paper introduces LSSF-Net, a lightweight deep learning architecture designed for accurate skin lesion segmentation in dermoscopic images, with a focus on mobile deployment (2409.01572). The network integrates conformer-based focal modulation attention, self-aware local and global spatial attention, and split channel-shuffle to achieve state-of-the-art performance with a minimal number of parameters (0.8 million). The efficacy of LSSF-Net is validated across multiple benchmark datasets, demonstrating its potential for computer-aided diagnosis on resource-constrained devices.
Introduction to LSSF-Net
The imperative for early and accurate skin lesion detection, particularly melanoma, is well-established. LSSF-Net addresses the challenges posed by the complex characteristics of skin lesions, such as indistinct boundaries, diverse appearances, and subtle variations in texture and color. The architecture leverages an encoder-decoder structure, incorporating several novel components to enhance feature extraction and segmentation accuracy, with the goal of balancing performance and resource efficiency for mobile deployment.
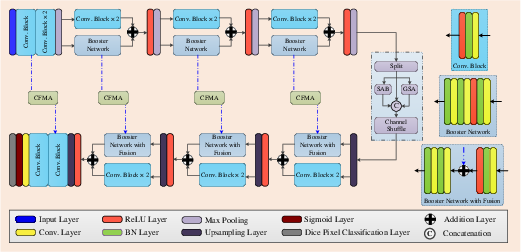
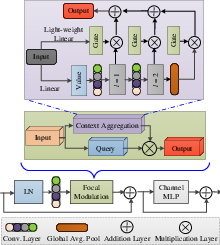
Figure 1: Block diagram of the proposed LSSF-Net. CFMA'' is conformer-based focal modulation attention,SAB'' is the self-attention block, and ``GSA'' is global spatial attention.
Architectural Innovations
LSSF-Net incorporates several key architectural innovations:
Implementation Details
The implementation involves four encoder-decoder blocks with specific convolutional operations, maxpooling, and upsampling techniques. The initial skip connection (so) and encoder block output (Eo) are computed using convolutional operations and maxpooling:
so=l3×3(Xin)
Eo=mp(l3×3(l3×3(so)))
Subsequent encoder block outputs (Ek) are refined using skip connections (sk) and residual learning:
Ek=mp[ℜ{βn(f3×3(βn(f3×3(sk))))+f3×3(l3×3(l3×3(Ek−1)))}]
The decoder stage reconstructs spatial feature maps using CFMA on skip connections:
ℑk=CFMA(sk)+l3×3(up(Dk−1))
The final output (Xout) is obtained through convolutional and sigmoid operations:
Xout=σ(f1×1(l3×(ℑk)))
Experimental Results
LSSF-Net's performance was evaluated on ISIC 2016, ISIC 2017, ISIC 2018, PH2, BUSI, and DDTI datasets. The evaluation metrics included Jaccard index, Dice coefficient, accuracy, sensitivity, and specificity. Ablation studies on the ISIC 2017 dataset demonstrated the contribution of each component to the overall performance (Figure 3). The results showed that the combination of CFMA, SAB, and SCS-SAB, along with transfer learning, yielded the best performance.
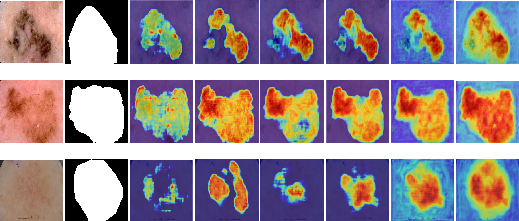
Figure 3: Visual results of ablation study on ISIC 2017 dataset. 1st column shows the color image, 2nd column shows the corresponding ground truth, 3rd column shows the output of baseline network (BN), 4th column shows the output of (BN + CFMA), 5th column shows the output of (BN + SAB), 6th column shows the output of (BN + CFMA + SAB), 7th column shows the output of (BN + CFMA + SCS-SAB), and 8th column shows the output of (BN + CFMA + SCS-SAB + Transfer Learning).
Visual comparisons on the ISIC 2018 dataset (Figure 4) further illustrate the effectiveness of LSSF-Net in accurately segmenting skin lesions, particularly in challenging scenarios with occlusions and low contrast.
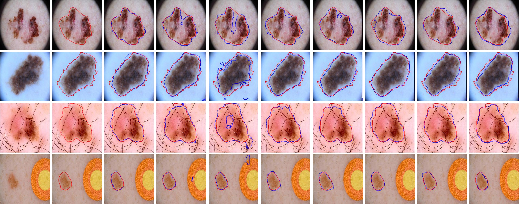
Figure 4: Comparison of the visual performance of the proposed LSSF-Net on ISIC 2018 [codella2019skin] dataset.
Similar visual results on the ISIC 2017 dataset (Figure 5) confirm the superior performance of LSSF-Net in achieving segmentation results that closely align with the ground truth.
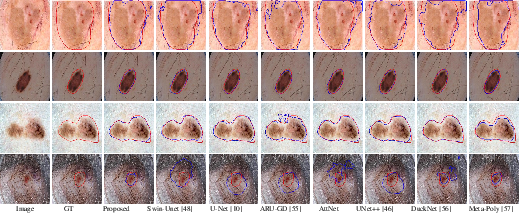
Figure 5: Comparison of the visual performance of the proposed LSSF-Net on ISIC 2017 [codella2018skin] dataset.
The ISIC 2016 dataset (Figure 6) also demonstrates LSSF-Net's ability to handle diverse scales and irregular shapes, consistently achieving optimal segmentation results.
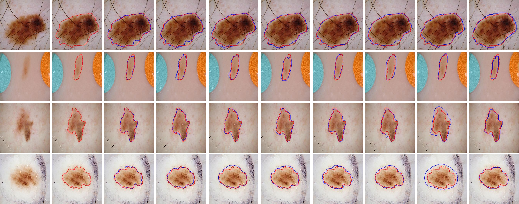
Figure 6: Comparison of the visual performance of the proposed LSSF-Net on ISIC 2016 [gutman2016skin] dataset.
Additionally, the generalization capability of LSSF-Net was validated on the PH2 dataset (Figure 7), where it accurately segmented lesion regions despite the presence of hair, contrast variations, and irregular boundary shapes.
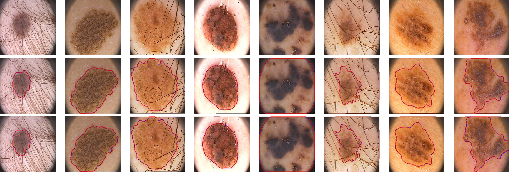
Figure 7: Visual results of the proposed LSSF-Net on the PH2 [mendoncca2013ph] dataset.
The BUSI (Figure 8) and DDTI (Figure 9) datasets further demonstrate LSSF-Net's ability to deliver precise segmentation results, even for images exhibiting diverse sizes and irregular shapes.
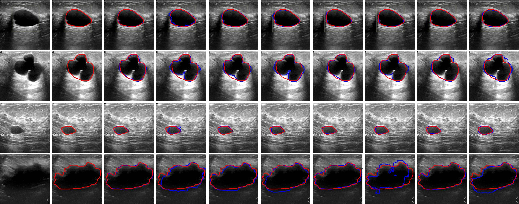
Figure 8: Comparison of the visual performance of the proposed LSSF-Net on BUSI [BUSIdataset] dataset.
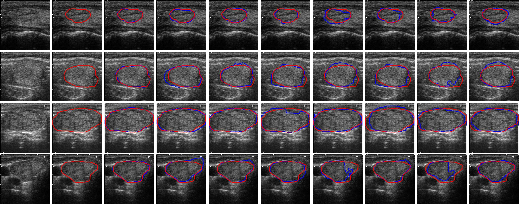
Figure 9: Comparison of the visual performance of the proposed LSSF-Net on DDTI [DDTIdataset] dataset.
Quantitative Analysis
Across the datasets, LSSF-Net consistently outperformed other state-of-the-art methods, achieving higher Jaccard indices and demonstrating robust generalization capabilities through cross-dataset evaluations. The computational complexity analysis revealed that LSSF-Net achieves superior computational efficiency with a minimal number of parameters and reduced inference time.
Limitations and Future Directions
While LSSF-Net excels in binary class segmentation tasks, its lightweight architecture may limit its applicability to more complex problems involving multiple modalities and classes. Future research could focus on extending LSSF-Net to support multiclass segmentation and multimodalities, enhancing its versatility and applicability to various medical imaging scenarios. Additionally, implementing quantization techniques can further optimize LSSF-Net for deployment on resource-constrained devices, making it suitable for CAD systems and mobile devices.
Conclusion
LSSF-Net represents a significant advancement in skin lesion segmentation, offering a balance of accuracy and efficiency suitable for mobile deployment. The architecture's innovative components and robust performance across multiple datasets highlight its potential for real-world applications in computer-aided diagnosis of dermatological conditions. Future research directions include extending the network's capabilities to handle more complex segmentation tasks and optimizing its deployment on resource-constrained devices.