- The paper introduces a novel active learning framework that synergizes machine learning with energy-based Flex ddG computations to enhance antibody screening.
- The methodology improves predictive performance with enhanced Spearman correlation and ROC-AUC scores, reducing dependency on extensive experimental data.
- The approach was validated on Trastuzumab mutants, achieving significant improvements in selecting candidates with lower binding energy values.
Active Learning for Energy-Based Antibody Optimization and Enhanced Screening
Introduction
The paper discusses a novel approach in optimizing protein-protein binding affinity with implications for therapeutic antibody development. This integration addresses the limitations of both machine learning and energy-based methods for predicting binding affinities. Machine learning methods, while adept at large-scale mutant screening, struggle with accurate predictions when limited data exist. Conversely, energy-based approaches like Rosetta's Flex ddG offer more precise predictions but at the cost of high computational demands. The proposed active learning methodology synergizes these approaches, combining the rapid screening capability of machine learning with the accuracy of energy-based models, by integrating Rosetta's Flex ddG with an RDE-Network to achieve efficient antobody optimization and screening.
Materials and Methods
Key to the methodology is an active learning workflow designed for Trastuzumab, an antibody targeting HER2, focusing on complementarity-determining regions (CDR-H). Through random mutagenesis, 100,000 mutants were generated, 98,567 of which were unique. These mutants were assessed using Luo's RDE-Network, enhancing predictive capability through Flex ddG, a surrogate model considering both experimental and computational binding predictions via multitask learning. The workflow involves iterative cycles of selecting mutants predicted to improve antibody binding, achieving a balance between enrichment of learning data and computational efficiency.
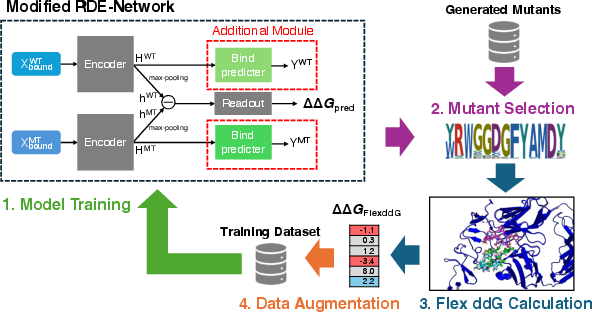
Figure 1: Overview of the proposed active learning workflow.
During the process, 1200 mutants were selected over six cycles, with Flex ddG computations guiding the candidate selection. Combinatorial techniques ensured an optimal balance between exploration and exploitation, aiding in the discovery of beneficial mutant candidates.
Results and Discussion
The results indicate a marked improvement in screening capability with the active learning approach over random selection. Throughout the cycles, the distribution of selected mutants with lower Flex ddG values significantly improved, demonstrating the suitability of active learning in antibody screening processes.
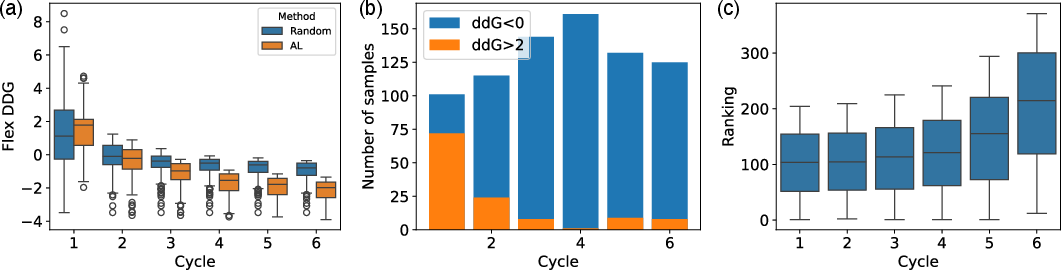
Figure 2: (a) Transition of the calculated top 200 Flex ddG values of the selected mutants at each active learning cycle. (b) Number of selected mutants that bound and unbound based on Flex ddG at each active learning cycle.
Spearman correlation analyses for binding predictions revealed improvements consistent with Flex ddG results, where the proposed method surpassed the baseline model and even Flex ddG itself in certain discriminative metrics. Notably, classifier performance enhanced without experimental data reliance, suggesting potential application in environments with limited initial data.
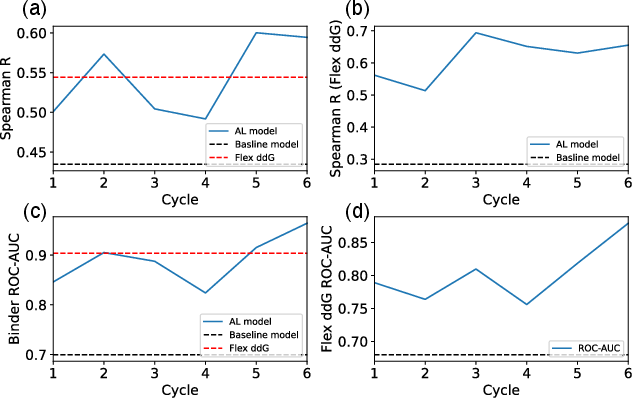
Figure 3: (a)-(d): Transition of Spearman correlation and ROC-AUC scores for the surrogate model's predictions at each active learning cycle.
Conclusions
The presented study validates the capability of an active learning framework not only in optimizing antibodies like Trastuzumab against targets such as HER2 but also in doing so efficiently by leveraging the computational gains of machine learning and the predictive precision of energy-based methods like Flex ddG. The integration achieves improved performance without necessitating exhaustive data, highlighting its utility in early design stages or in optimizing pharmacological properties of variants. This framework is adaptable across models and potentially beneficial in furthering computational antibody design innovation. Future work proposes enriching the active learning framework by combining it with genetic algorithms or more precise affinity prediction methods to enhance modeling accuracy and screening precision.


