- The paper introduces an FFT-ASCA pipeline that transforms chromatographic signals into the frequency domain for robust hypothesis testing despite peak drift.
- It adapts ASCA to operate on complex Fourier coefficients, enabling reliable detection of significant chemical factors under misaligned peak conditions.
- The methodology circumvents traditional time-domain challenges, offering improved data integrity and reproducible results in both synthetic and real datasets.
An Alignment-Agnostic Methodology for the Analysis of Designed Separations Data
Introduction
The analysis of chemical separations data, particularly in chromatography, often presents challenges related to peak alignment and retention time drift. These artefacts can significantly affect data interpretation, leading to false positives or negatives in multivariate analyses. Traditional methods rely heavily on time-domain integration of chromatographic peaks, which can be misrepresentative if the components don't align perfectly across samples. This paper proposes an alternative methodology that adopts a frequency domain approach using Fast Fourier Transform (FFT) to analyze separations data. This approach is alignment-agnostic, aiming to preserve the integrity of chemical information without the pitfalls of retention time inconsistencies.
Frequency Domain Approach
The foundation of the proposed method is the transformation of time-domain chromatographic data into the frequency domain via FFT. This transformation represents each sample as a matrix of complex Fourier coefficients, capturing amplitude and phase information of the chromatographic signals without directly relying on retention time alignment. This frequency domain representation is subjected to a generalized ANOVA-Simultaneous Component Analysis (ASCA), traditionally used for structured multivariate data, modified here to handle complex matrices.
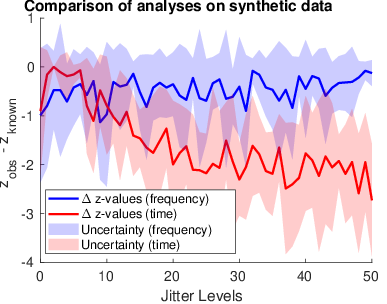
Figure 1: As shown by the results of this analysis, as the jitter in the data increases, the hypothesis testing step in parGLM analysis using the time-domain data becomes much less sensitive. However, following pre-processing using an FFT analysis the results are much more consistent.
Methodological Components
- FFT-ASCA Pipeline: The initial step involves transforming raw GC-FID signals using FFT. This transformation is followed by a generalized ASCA to decompose the data into significant factors.
- Complex Matrix Operations: ASCA is adapted to handle complex matrices, enabling permutation testing on the magnitude of Fourier coefficients to assess statistical significance.
- Data Reconstruction and Interpretation: While frequency domain analysis aids in significance testing, interpreting the results necessitates transforming loadings back to the time domain, facilitating a clear understanding of the chemical phenomena represented by the signal components.
Results
Synthetic Dataset: Demonstration on synthetic GC-FID datasets indicates that FFT-based methods maintain sensitivity in hypothesis testing under peak drift conditions better than time-domain analyses. The method reliably detects significant factors even when peak alignment is inconsistent.
Real Dataset Application: Application to real-world datasets, specifically in the context of experiments involving chemical profile changes in Tribolium castaneum, illustrates comparable data interpretation to conventional peak table analyses. However, the frequency domain approach circumvents issues arising from missing values and unaligned peaks in the peak tables, offering a robust alternative in these scenarios.
Discussion
The introduction of a frequency domain approach presents a paradigm shift in chromatographic data analysis. This method reduces reliance on precise peak alignment, a major source of artefacts in multivariate analyses, thereby preserving the integrity of chemical information. While current findings are promising, further exploration is necessary to integrate this method with existing analytical pipelines and evaluate its performance across a wider variety of samples and experimental designs.
Conclusion
This proposed FFT-ASCA methodology offers a novel way to analyze chromatographic data, sidestepping alignment issues that traditionally hinder data interpretation. By transforming data into the frequency domain, the approach provides a robust framework for analyzing complex chromatographic datasets. Future work may consider extending this approach to other modalities and exploring its integrations with emerging data analysis frameworks. The implementation details and code can be accessed from the provided repository, enabling reproducibility and further exploration in diverse analytical settings.
