- The paper introduces GAFL, a method that integrates geometric algebra with flow matching to generate highly designable protein backbones.
- It leverages Clifford Frame Attention to encode complex geometric relationships, achieving accurate modeling of secondary structure content.
- Results demonstrate improved designability, diversity, and novelty compared to existing models, enhancing potential applications in protein engineering.
Generating Highly Designable Proteins with Geometric Algebra Flow Matching
Introduction
The paper "Generating Highly Designable Proteins with Geometric Algebra Flow Matching" introduces a novel method for designing protein backbones using geometric algebra and flow matching. This approach harnesses Clifford Frame Attention (CFA), which enhances AlphaFold2's invariant point attention (IPA) architecture by embedding geometric features in projective geometric algebra (PGA). The method aims to generate protein backbones that exhibit high designability, diversity, and novelty, reflecting the secondary structure distributions found in natural proteins.
Methodology
Geometric Algebra and Protein Representation
The core innovation lies in representing protein backbone residue frames using projective geometric algebra (PGA). PGA allows for the representation of geometric transformations and features as multivectors, enabling operations like rotations and translations to be expressed algebraically. This choice introduces a geometric inductive bias, beneficial for modeling properties like distances, angles, and incidences in 3D space.
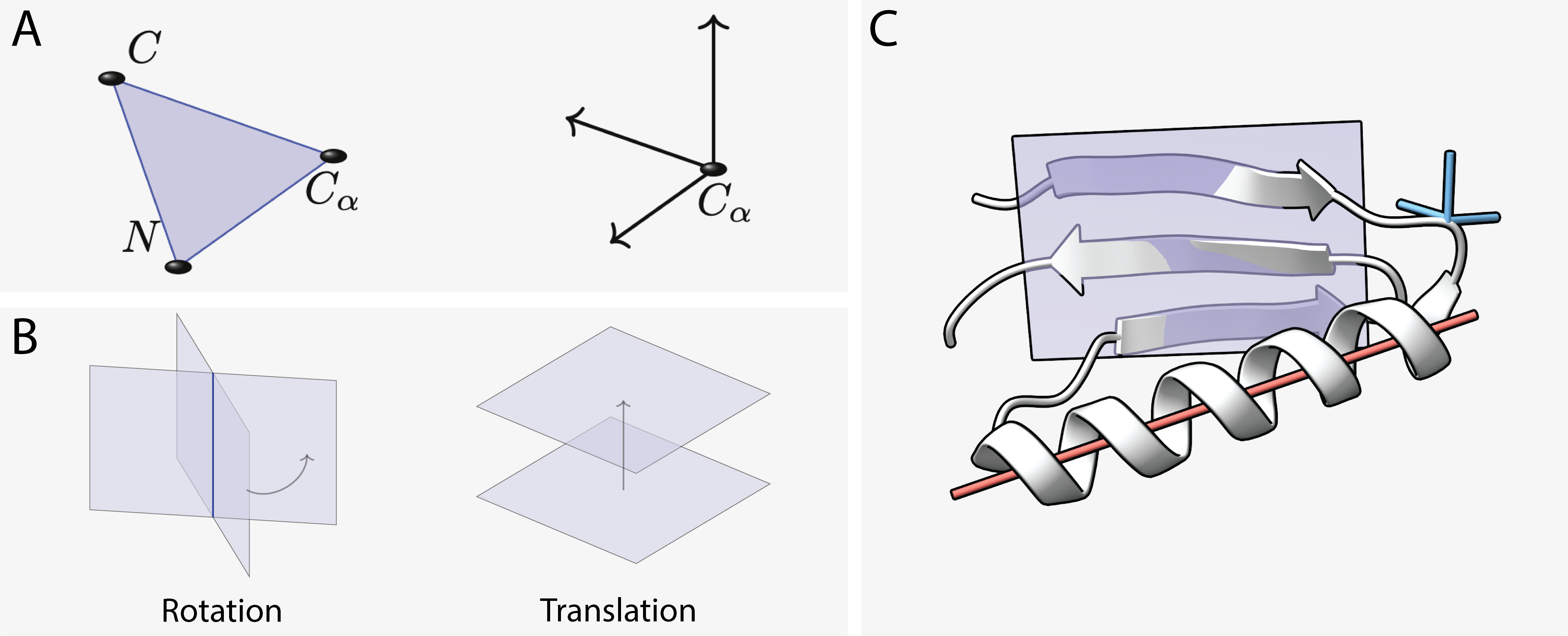
Figure 1: (A) Protein backbone residue with three backbone atoms represented by a coordinate frame. (B) In PGA, a frame can be represented via the geometric product of four planes. Two of the planes parameterize the frame's rotation, while the other two encode translation.
Clifford Frame Attention (CFA)
CFA extends IPA by integrating geometric algebra, using multivectors instead of scalar features for attention values. This allows the construction of higher-order messages that incorporate complex geometric relationships beyond pairwise interactions. The architecture leverages the algebraic properties of PGA to calculate meaningful geometric transformations, enhancing the generation of diverse secondary structures, such as α-helices and β-sheets.
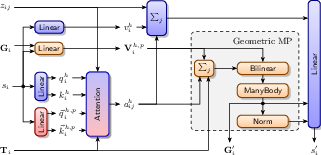
Figure 2: Overview of Clifford frame attention, showcasing layers that handle scalar information, point-valued information, and features in PGA.
Flow Matching Framework
The flow matching approach underpins the protein design process by transforming a prior distribution into a target distribution of frames using continuous normalizing flows. This method trains a vector field conditioned on noise-perturbed data examples, interpolating between an initial and target distribution over protein conformations. The implementation builds on FrameFlow, which already employs a flow matching mechanics, but incorporates the advanced geometric messaging capabilities of CFA.
Experimental Results
The experiments demonstrate that the proposed GAFL model excels in generating protein backbones with desirable characteristics across several metrics:
- Designability: GAFL achieves high designability scores, indicating that generated structures can be converted reliably into stable protein sequences that fold correctly.
- Diversity and Novelty: The model produces a range of structures that maintain low similarity to each other and to known proteins, deviating from the prevalent overproduction of α-helices seen in other models.
- Secondary Structure Content: The method captures the statistical distribution of secondary structures found in natural proteins, even for longer and more complex backbones.
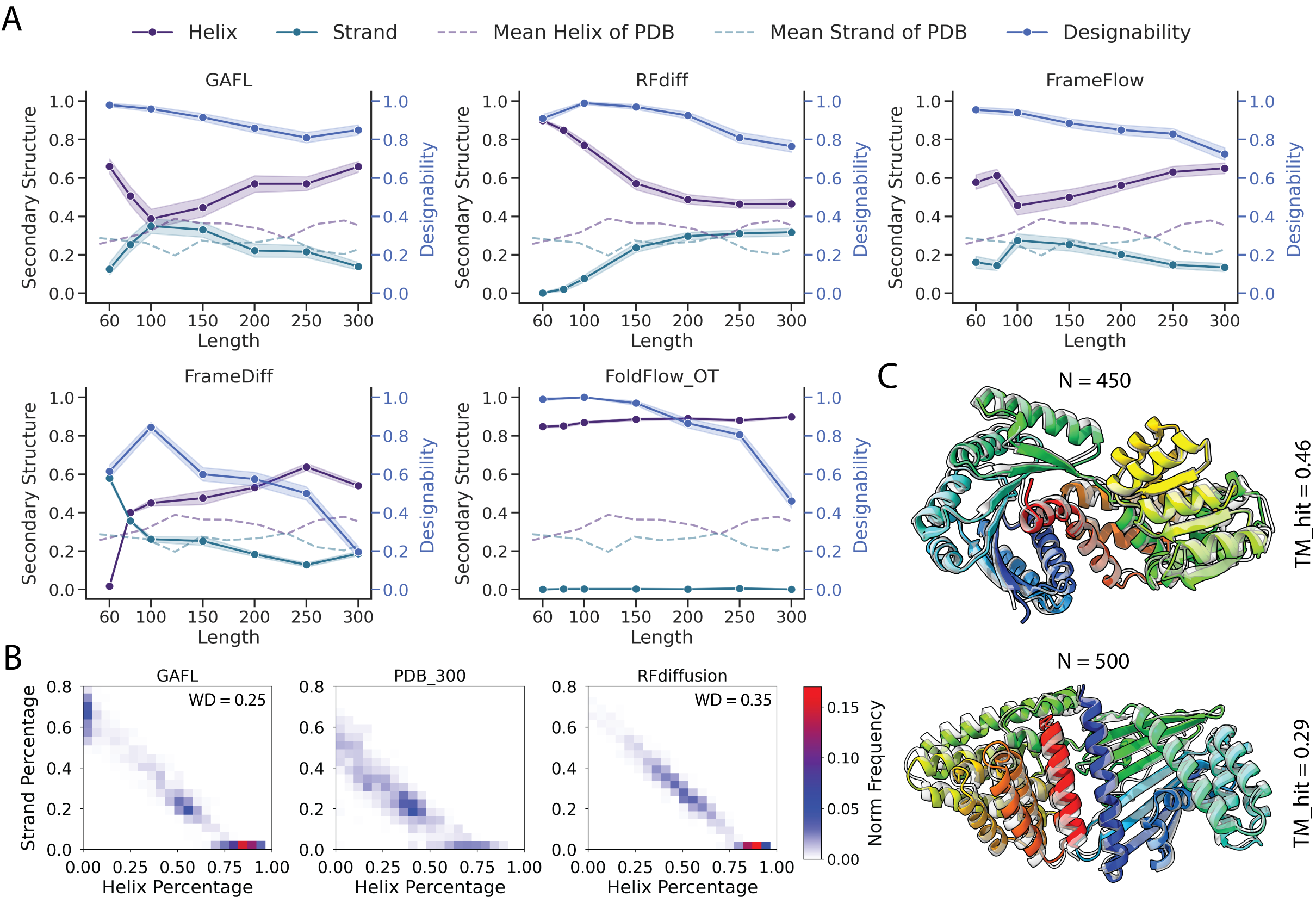
Figure 3: Performance metrics for models evaluated in terms of designability and secondary structure content as a function of backbone length.
Implications and Future Work
The presented approach significantly contributes to the field of de novo protein design. GAFL's ability to accurately reproduce natural secondary structure distributions while maintaining high designability and diversity positions it as an effective tool for protein engineering tasks, from medical therapeutics to novel material design. Future work could focus on extending the methodology to more explicitly incorporate functional constraints, enabling the design of proteins with specific desired activities.
Overall, this research highlights the immense potential of integrating geometric algebra into neural architectures for complex biological applications, paving the way for more sophisticated and efficient generative models in protein design.
Conclusion
The integration of geometric algebra with flow matching and advanced attention mechanisms offers a promising avenue for protein design, as demonstrated by the GAFL architecture. This approach not only meets but exceeds current capabilities in generating feasible, novel, and diverse protein structures, opening exciting possibilities for future developments in computational biology and biochemistry.


