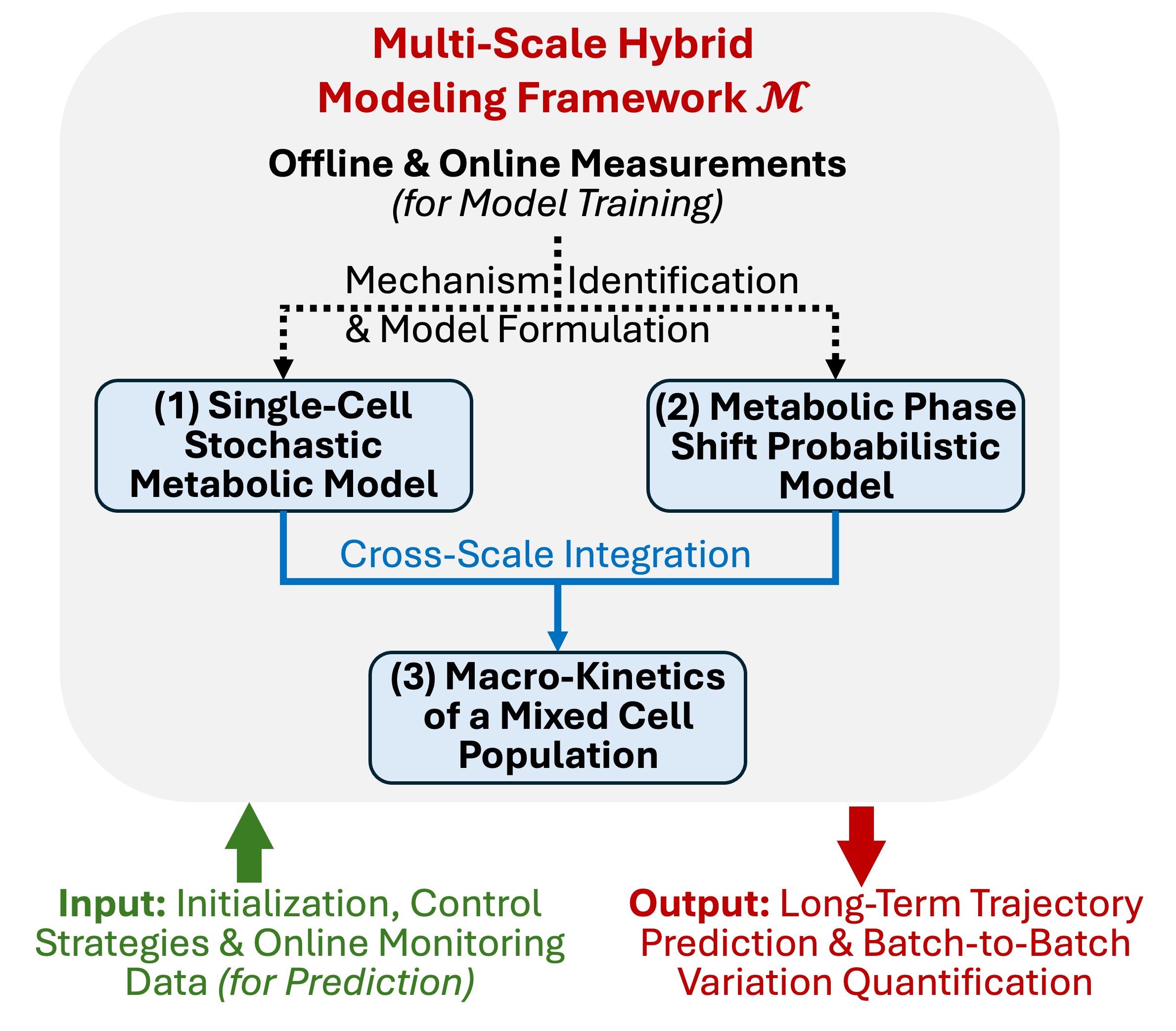
- The paper introduces a hybrid framework that combines stochastic single-cell kinetics, probabilistic phase transitions, and macro-population modeling to predict CHO cell culture processes.
- It demonstrates robust prediction accuracy with metrics such as less than 10% WAPE and 90-95% prediction intervals across multiple forecasting horizons.
- The model offers mechanistic interpretability and supports digital twin applications by integrating heterogeneous online and offline data streams.
Introduction and Motivation
This work presents a rigorous framework for predictive modeling of Chinese Hamster Ovary (CHO) cell cultures by integrating multiscale biological knowledge—capturing cellular stochasticity and inter-phase metabolic regulation—with mechanistically grounded macroscale population kinetics. The primary objective is to improve quantitative prediction and uncertainty quantification of cell culture trajectories, particularly in the presence of asynchronous metabolic phase shifts and process variability, which are critical for robust biomanufacturing and digital twin applications.
Limitations of Existing Models
Prevailing approaches such as Flux Balance Analysis (FBA), Metabolic Flux Analysis (MFA), and mechanistic kinetic models have yielded significant mechanistic understanding and optimization capabilities in microbial and mammalian cell culture processes. However, virtually all multi-scale frameworks to date are deterministic, often do not capture cell-to-cell or batch-to-batch stochasticity, and typically assume homogeneous or synchronized phase transitions at the population level. This results in underrepresented process variability and insufficient support for risk-aware, real-world process optimization and control.
Multi-Scale Hybrid Modeling Framework
The authors introduce a modular, hierarchical model structure composed of three fundamental components: (1) a stochastic single-cell mechanistic metabolic network, (2) a probabilistic phase transition model, and (3) a macro-kinetic population model. The integrated system is designed to flexibly represent process dynamics under diverse bioreactor conditions, map causal interdependencies, and enable reliable prediction with robust uncertainty quantification.
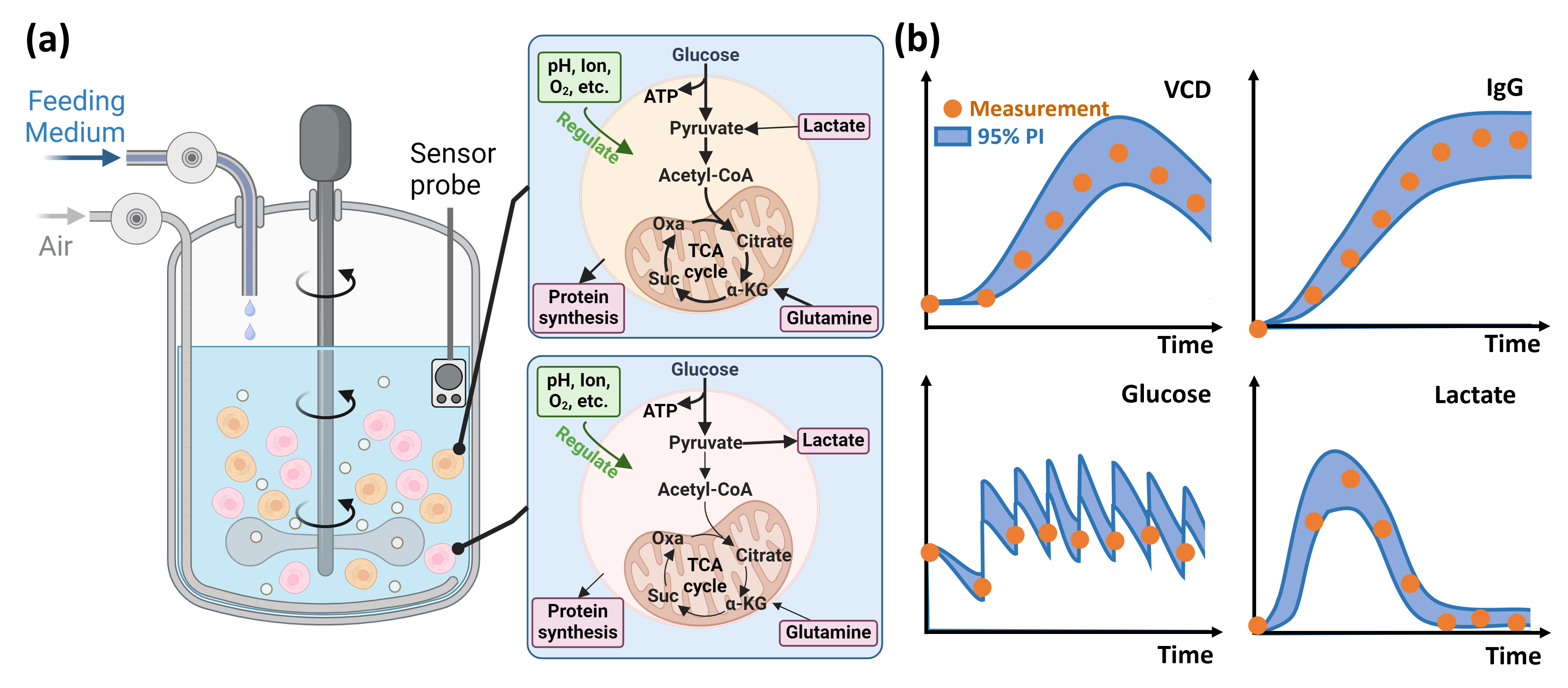
Figure 2: Schematic illustration of the multi-scale hybrid framework and example risk-based predictions for key variables (VCD, IgG, glucose, lactate), including uncertainty estimates.
Mechanistic Integration Across Scales
This system supports flexible data assimilation from heterogeneous measurement modalities, enabling inference of hidden states (e.g., phase distribution, latent fluxes) and prediction of unmeasured critical variables over extended time horizons.
Experimental System and Data Sources
The framework is empirically validated using a recombinant CHO-K1 cell line expressing a monoclonal anti-HIV antibody (VRC01), grown in an advanced 12-vessel ambr250 system under three feeding/pH control strategies (Cases A–C; triplicate for each). Comprehensive, high-frequency online and offline measurements include viable cell density (VCD), viability, metabolite panels (glucose, lactate, glutamine, etc.), product titer, pH, dissolved oxygen, as well as volumetric and environmental process controls.
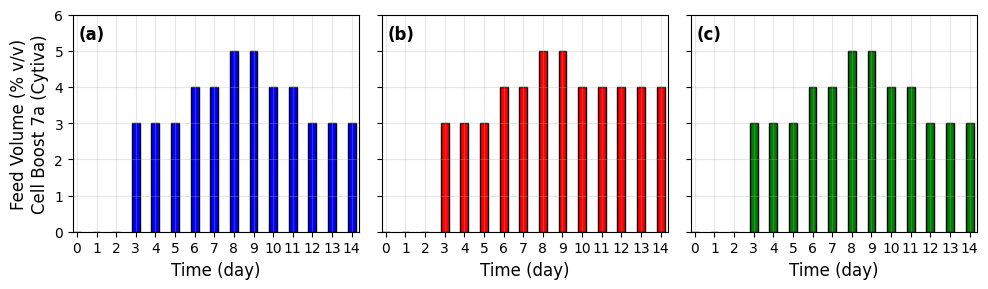
Figure 5: Feeding strategies for three experimental cases illustrate the pyramid and dynamic modifications employed to probe regulatory responses and phase transitions.
Key Model Components
Each cell’s exchange fluxes are modeled as follows:
rt=Nvz[ut]dt+{Nσz[ut]N⊤}1/2dWt
- N: stoichiometry matrix
- vz: M-M flux vector, phase- and context-dependent
- σz: diagonal fluctuation matrix (proportional to flux mean, modulated for phase-specific stochasticity)
- z: cell's metabolic phase (growth, stationary, decline)
- dWt: standard Wiener process
Regulatory dependencies (e.g., substrate/product inhibition, pH-dependent enzymatic activity) are explicitly included at key posts in the metabolic network.
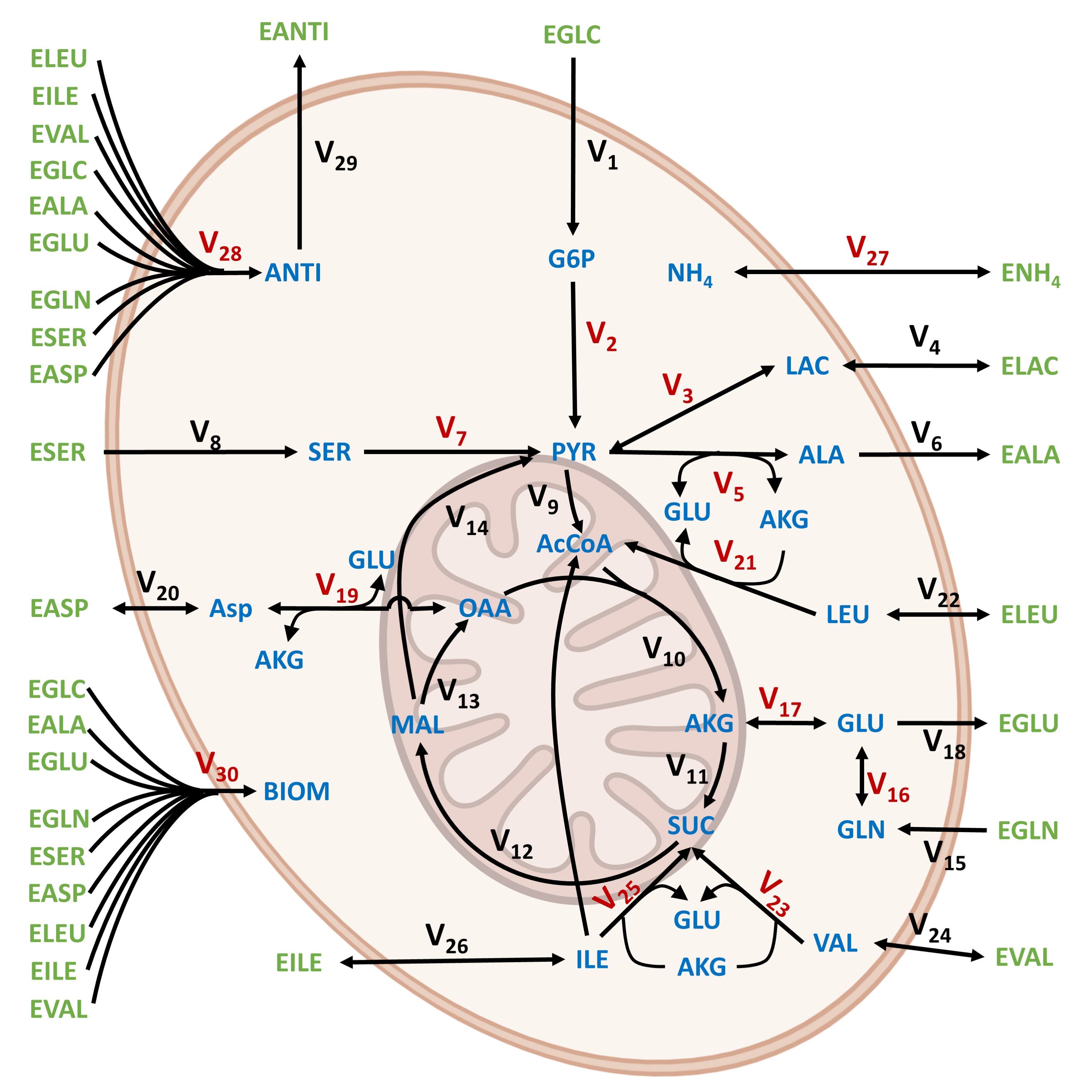
Figure 7: Core CHO metabolic network, annotated to distinguish reactions governed by explicit kinetic models (red) from those handled via pseudo-steady-state closure.
Probabilistic Phase Transition Model
At designated time steps, a sigmoid logistic model computes phase transition probabilities as joint functions of time, qO2, pH, and contextually relevant metabolite rates:
P(zth+1=j∣zth=i,th,qO2,th,pHth)=1+exp(−(w⋅x))1
where x is the feature vector and w denotes learned coefficients. Only the most predictive variables (culture age, qO2, pH) are retained for parsimony and identifiability.
Macro-Kinetic Heterogeneous Population Model
Population vectors for each phase evolve according to:
dXtz=μz[ut]Xtzdt+{σμz[ut]}1/2XtzdWt
with cross-phase transitions governed by the phase transition matrix Pth.
The aggregate extracellular dynamic (for any metabolite) is:
dut=z∑XtzNvz[ut]dt+z∑i=1∑Xtz{Nσz[ut]N⊤}1/2dWt,i
Due to sparse sampling intervals for some readouts, an Expectation–Maximization (EM) algorithm is implemented to interpolate latent transitions, fit model parameters, and account for missing or uncertain data. Prediction accuracy is evaluated using weighted absolute percentage error (WAPE), while forecast uncertainty is quantified by the empirical coverage of prediction intervals (PI), evaluated for multiple look-ahead horizons.
Trajectory Prediction
The model shows robust WAPE (typically <10%), both for 1, 3, and 5-day ahead look-ahead and for cross-batch extrapolation scenarios. PI coverage consistently matches nominal levels (90–95%) even for multi-day prediction horizons, indicating reliable uncertainty quantification. Notably, model median and full credible intervals for cell growth, antibody titer, and major metabolite panels closely parallel the measured batch data.
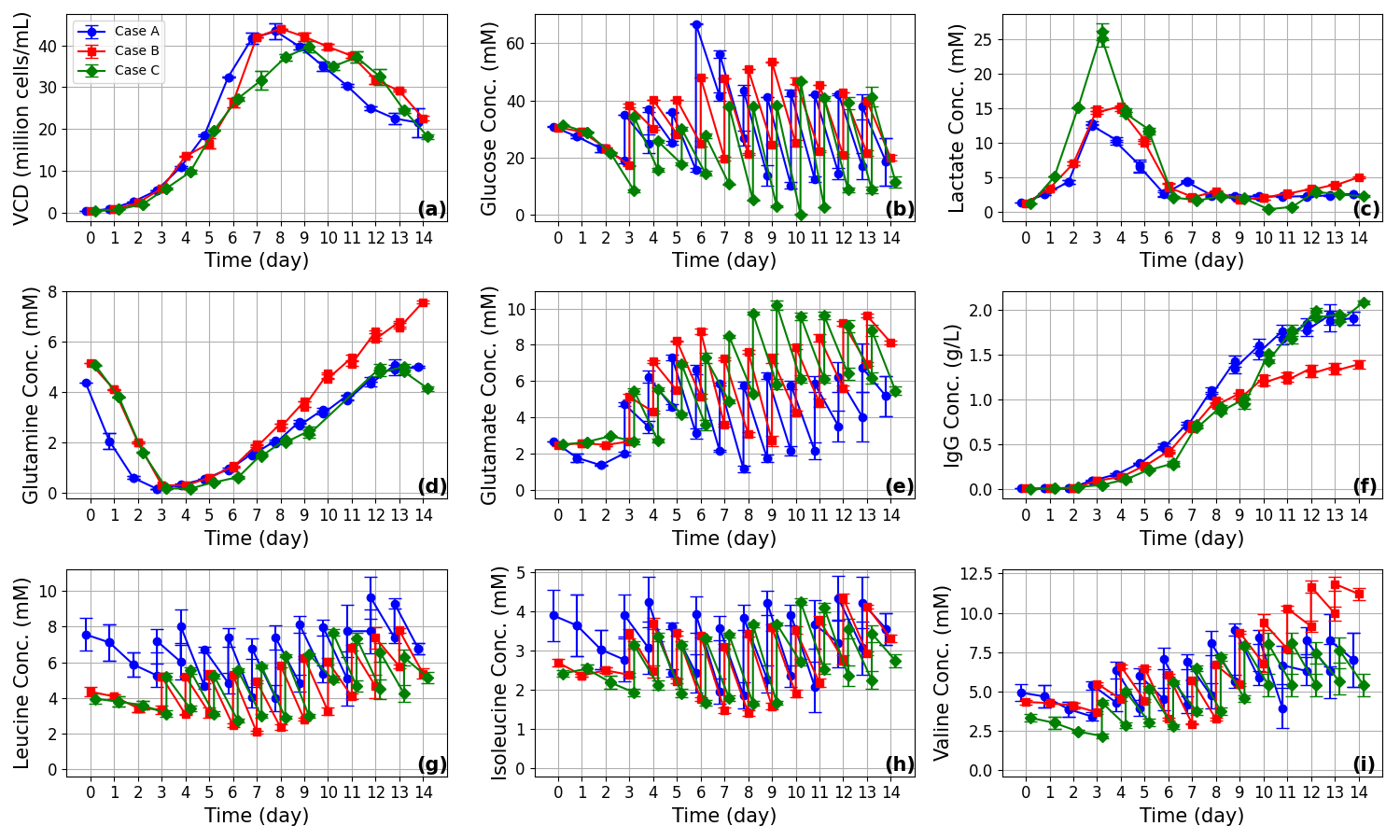
Figure 3: Comparison of measured and predicted time courses for VCD, glucose, lactate, and key amino acids across all three experimental regimes. Training on remaining datasets enables rigorous “leave-one-batch-out” generalization assessment.
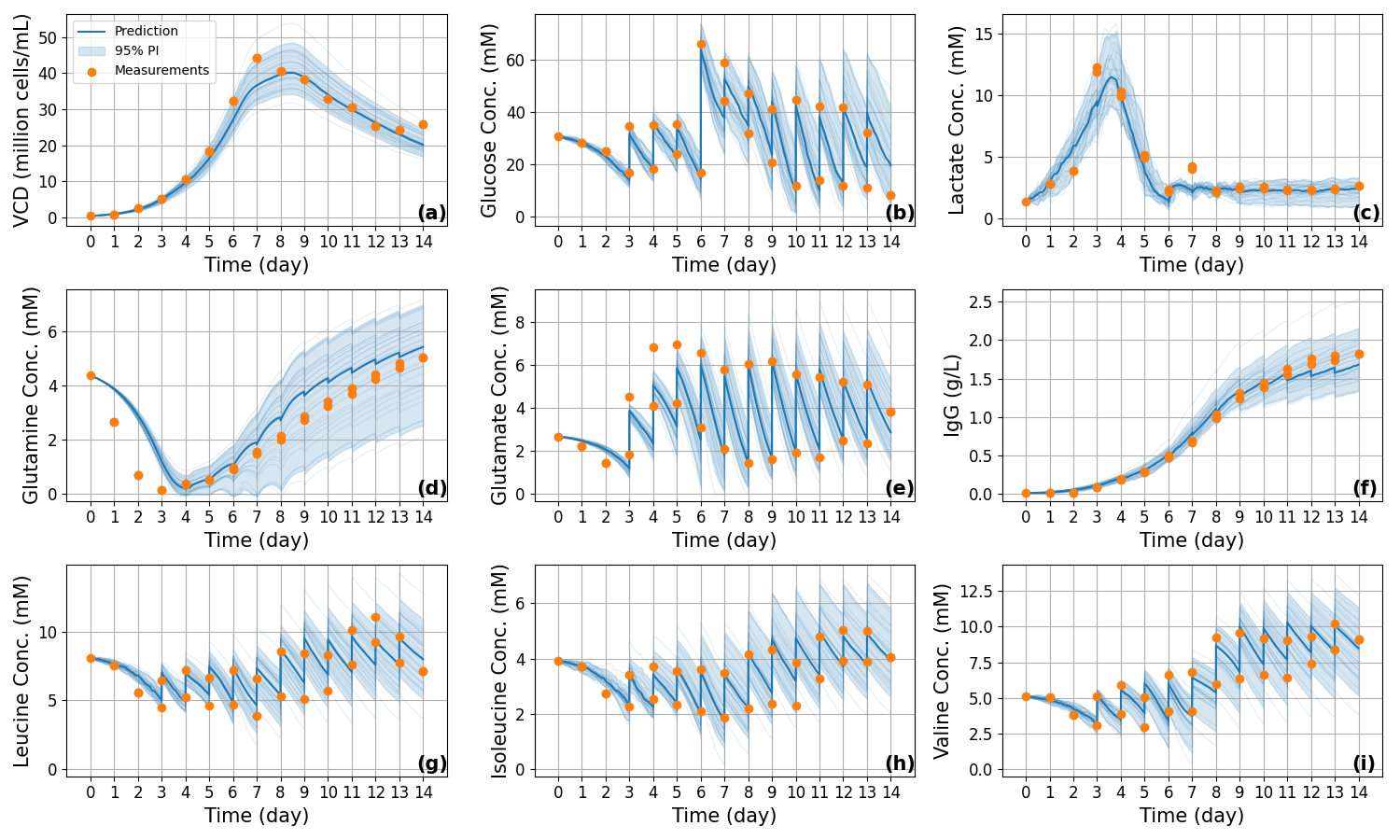
Figure 10: Out-of-sample (Case A, Rep 1) trajectory prediction using only online data and initial conditions; the blue band displays the 95% prediction interval, and orange dots are ground-truth time points.
Mechanistic Interpretability
The model not only enables prediction but provides detailed mechanistic attributions. For example, Case A exhibits heightened IgG synthesis correlated with sustained BCAA abundance and higher qO2 (Figure 11). Case C, with dynamically regulated pH, shows late-stage productivity driven by combined pH and ammonia uptake effects, in agreement with both fluxomic and transcriptomic theory.
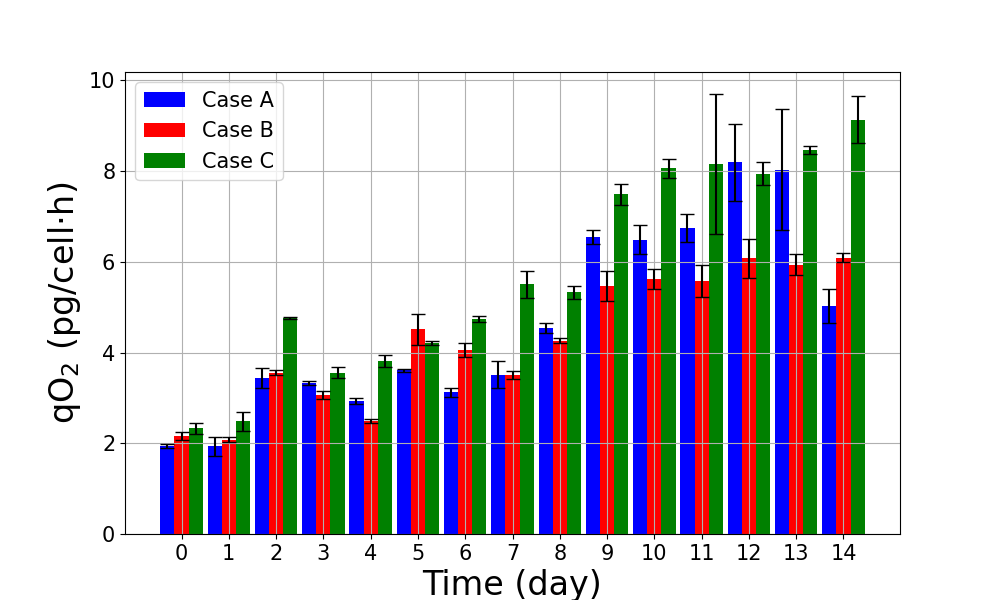
Figure 8: Quantification of qO2 (cell-specific oxygen uptake rates) captures dynamic regulatory adaptation and supports mechanistic attribution of differences across control regimes.
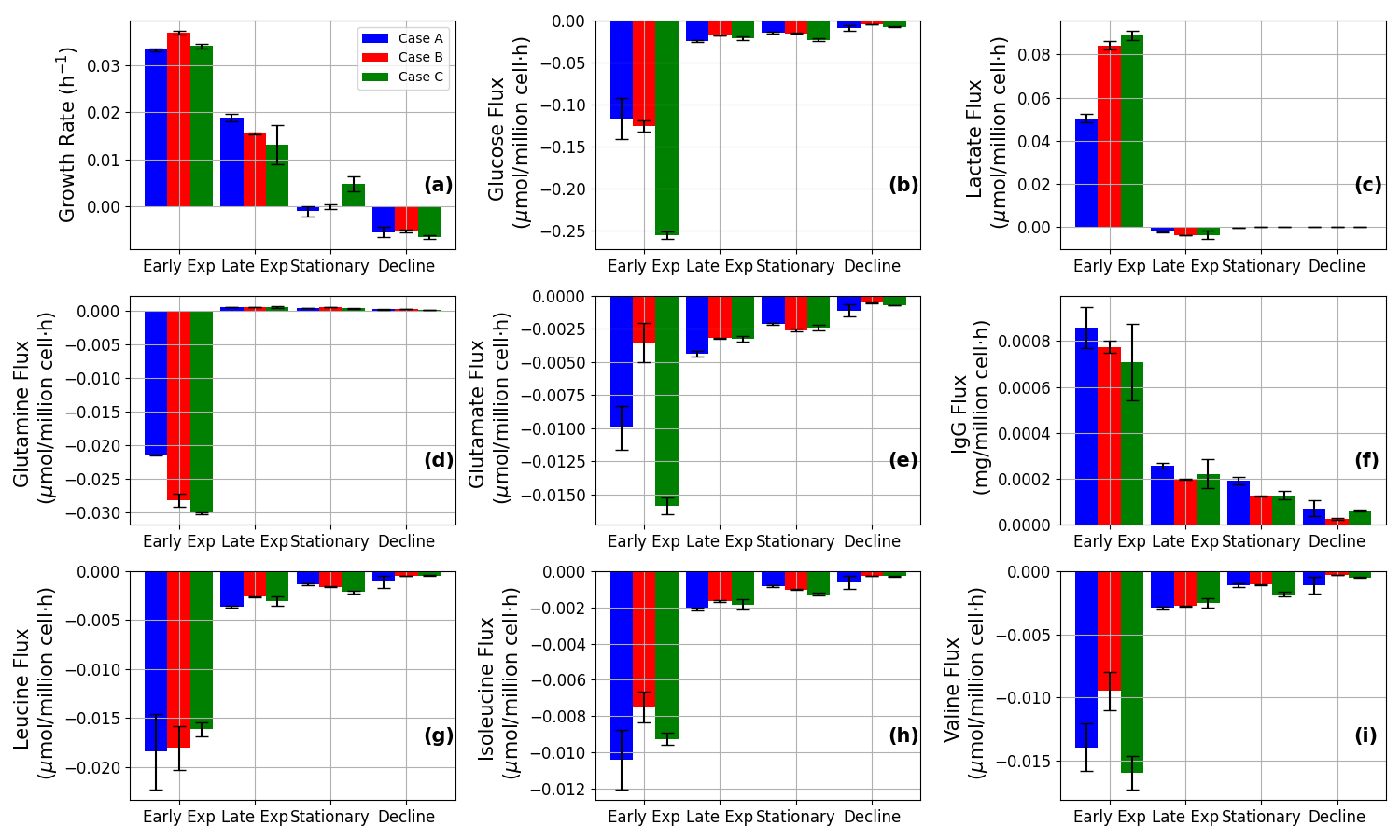
Figure 11: Predicted flux profiles for major metabolites and IgG illustrate phase- and case-specific regulatory phenomena, including shifts in glycolytic and amino acid utilization pathways.
Explicit modeling of asynchronous metabolic phase transitions enables direct inference of phase population distributions and associated cascade effects on metabolite profiles and productivity. Variability between experimental replicates—often a key practical issue in biomanufacturing—is quantitatively explained and bounded by the model’s stochastic propagation.
Implications for Digital Twins and Advanced Process Control
This framework provides a foundation for digital twin platforms capable of efficient, uncertainty-aware prediction and optimization in mammalian cell culture manufacturing:
- Mechanistic Digital Twins: The modularity and physical interpretability of individual modules enable extensibility to new strains, genetic modifications, or industrial process regimes without retraining or loss of explainability.
- Real-time Optimization and Control: Integration of heterogeneous measurement streams (including low-latency online data) enables closed-loop optimization and anomaly detection.
- DoE and Risk Analysis: Monte Carlo simulation over calibrated model parameters provides batch-level risk assessment, improved experimental design, and rational process scale-up strategies.
Conclusions and Outlook
This work establishes that predictive integration of multiscale mechanistic models, with explicit representation of cell-to-cell and population stochasticity and asynchronous metabolic phase transitions, materially advances both quantitative prediction and mechanistic understanding of bioprocess dynamics. The robust out-of-sample predictive performance—supported by reliable uncertainty quantification—provides actionable capabilities for bioprocess digital twins and sets a benchmark for future model-based process control initiatives.
Expansion to additional cell lines, process scales, or product modalities will require appropriate adaptation of network topology and parameterization, but the modular framework directly accommodates these generalizations. Methodological advances in high-throughput single-cell analytics and real-time sensor networks will further enhance the scope, resolution, and feedback capacity of such hybrid models.
There is potential for future cross-fertilization with AI-driven (e.g., reinforcement learning) control methods and advanced multi-omics data assimilation, enabling next-generation robust, sample-efficient optimization and process monitoring across the full spectrum of advanced biomanufacturing environments.