- The paper’s main contribution is introducing miVAE, a novel autoencoder architecture that disentangles latent variables to capture both individual-specific and shared neural features in V1.
- It employs directed generative modeling and variational inference to robustly align multi-modal data, demonstrating enhanced cross-individual performance.
- Experimental results show that miVAE outperforms baseline models, with ablation studies confirming the critical role of latent space design in modeling neural dynamics.
Multi-Modal Latent Variables for Cross-Individual Primary Visual Cortex Modeling and Analysis
Introduction
This paper addresses the significant challenges in modeling the primary visual cortex (V1) across different individuals, particularly due to the inherent variability in neural characteristics and the integration of partial neural recordings with complex visual stimuli. The study introduces a multi-modal identifiable variational autoencoder (miVAE) designed to effectively map neural activity and visual stimuli into a unified latent space, enhancing cross-modal correlations and facilitating robust analysis without the need for subject-specific adjustments.
miVAE Architecture and Methodology
The core innovation of the paper lies in the architecture of the miVAE, which employs two-level latent space disentanglement. Neural activity is modeled with both idiosyncratic latent variables and preserved latent variables, capturing individual-specific details and consistent information across individuals. Visual stimuli are differentiated into activity-related and unrelated variables. This separation refines the latent space, optimizing the correlation between neural and visual inputs.
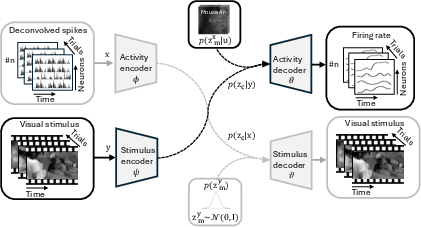
Figure 1: miVAE for modeling V1 neural activity x and corresponding visual stimuli y.
The cross-modal approach uses directed generative modeling to capture stimulus-evoked neural dynamics, reinforcing the correlation between neural activity and visual stimuli within the latent space. Variational inference aids in approximating intractable posteriors, optimizing the variational evidence lower bound (ELBO) across dual domains for effective learning.
Neural Coding and Attribution Analysis
The study validates miVAE using a large-scale mouse V1 dataset, demonstrating superior performance in cross-individual latent representation. Encoding and decoding models show remarkable scalability with increasing dataset sizes, while score-based attribution analysis uncovers distinct neuronal subpopulations and stimulus-sensitive regions.
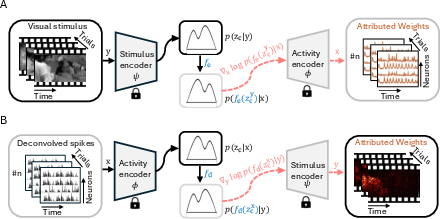
Figure 2: Neural coding in the latent space with following score-based Attribution Analysis.
The attribution strategy utilizes a score function, treating neural encoders as decoders, to relate high-level latent representations back to original data, identifying significant responses in both neural populations and stimulus features.
Experimental Results and Ablation Studies
The miVAE system consistently outperforms baseline models, illustrating its effectiveness in multi-modal and cross-modal latent alignment without subject-specific training. Ablation studies reveal the critical roles of latent variables in capturing cross-individual variations, and dataset scalability further enhances model performance, validating its applicability to broader neuroscience research.
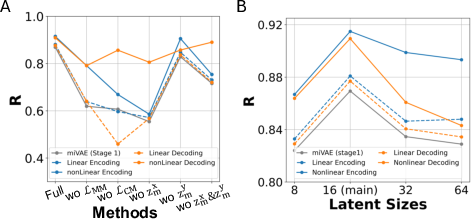
Figure 3: Ablations on modeling methods, showcasing the impact of loss functions and latent variable configurations.
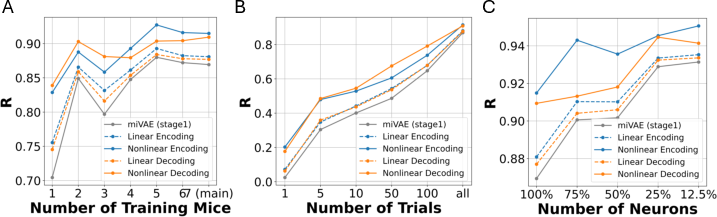
Figure 4: Ablations studies on data scale, showing the relationship between dataset size and model performance.
Conclusion
The miVAE framework presents a robust methodological advancement for cross-individual modeling of V1, facilitating detailed neural and stimulus analysis. Its scalable and interpretable architecture holds promise for broader applications in sensory cortex research, potentially extending beyond visual to auditory systems, enhancing our understanding of complex neural dynamics and cross-sensory integration.



