- The paper presents a framework combining active learning with label-conditional conformal prediction to minimize manual labeling and ensure reliable predictions.
- It utilizes model-agnostic methods like TF-IDF + XGBClassifier to achieve robust performance in resource-constrained environments.
- The active selection of high-uncertainty texts via k-means clustering enhances label diversity and supports real-time deployment in medical text mining.
Conformal Active Learning for Medical Text Mining
The paper "Mining Unstructured Medical Texts With Conformal Active Learning" (2502.04372) introduces a novel framework for extracting relevant information from unstructured medical texts, specifically Electronic Health Records (EHRs), using a combination of active learning and label-conditional conformal prediction. This approach aims to reduce the manual labeling effort required for training classification models while providing reliability through uncertainty quantification. The framework is designed to be model-agnostic, allowing for deployment in resource-constrained environments while addressing privacy concerns.
The core of the proposed methodology is a Conformal Active Learning framework designed to iteratively refine a classification model with minimal manual labeling (Figure 1).
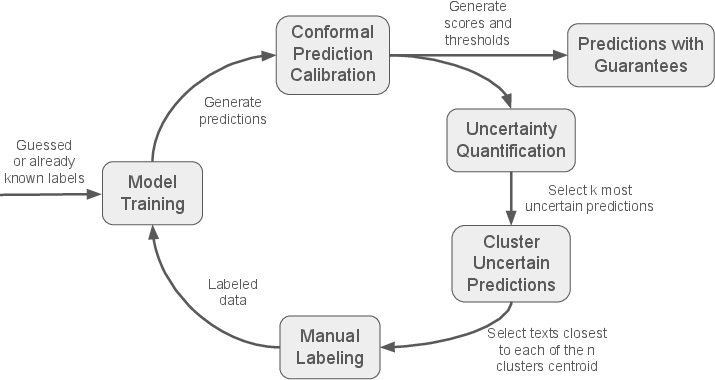
Figure 1: Diagram of the active leaning cycle.
The process starts with an initial set of labels, either guessed or derived from a small subset of manually labeled data. The data is split into training and validation sets. A classification model is trained on the training data, and a label-conditional conformal model is calibrated using the validation set. This conformal model assigns a score to each prediction, reflecting the model's uncertainty about a given label y for a sample text X. The conformity score is calculated as:
s(x,y)=1−p^(y ∣ x)
where p^(y∣x) is the predicted probability of x having label y according to the classification model. Thresholds ty,α for each label y are computed using the validation split to ensure a specified confidence level α. Calibrated set predictions Cα(Xnew) are then made for new data points Xnew, consisting of a subset of labels:
Cα(Xnew)={y:s(Xnew,y)≤ty,α}
This ensures that the predicted labels fall within the specified confidence level 1−α, providing a transparent, uncertainty-aware guarantee of correctness for each classification. The mean conformity score SX over all predicted labels for a sample text X is used to rank data points by uncertainty:
SX=∣Cα(X)∣1y∈Cα(X)∑s(X,y)
The top ktop data points with the highest uncertainty scores are selected and clustered using k-means into kcluster groups (kcluster<ktop). From each cluster, the point closest to the centroid is chosen for manual labeling, ensuring a diverse sample representing a broad range of high-uncertainty predictions. The framework can also incorporate a fraction of low-uncertainty predictions to validate the model's performance on straightforward cases.
Implementation and Deployability
The framework's classification model agnosticism allows training and selection based on available hardware, facilitating on-premise deployment in healthcare institutions. The authors are developing a web interface, OLIM (Open Labeller for Iterative Machine learning), to streamline the iterative labeling process. The code is released as open source under the GPL-v3.0 license, with the entire setup packaged in Docker containers for easy installation.
Experimental Evaluation
The framework was evaluated using a database of Amazon product reviews as a proxy for medical records, due to the sensitive nature of patient information. Four labels were defined for classification: Pet product, Drinkable product, Low quality product, and Damaged product. Experiments were conducted with 100 or 200 manually labeled texts, using ktop=500 and kcluster=6. A 20% subset of labeled data was randomly assigned to the validation dataset in each cycle. The models tested included TF-IDF + XGBClassifier and DeBERTaV3. Additional experiments explored the impact of balancing high and low uncertainty texts and starting with pre-labeled texts.
Results and Discussion
The framework achieved strong performance with the TF-IDF + XGBClassifier model, even with a limited number of manually labeled texts. With 200 manual labels, combining high and low uncertainty in ktop selection and adding a small number of pre-labeled texts, the framework demonstrated effectiveness in optimizing labeling efforts while maintaining robust performance. The deep learning model DeBERTaV3 did not perform as well as expected, suggesting that simpler models can deliver competitive results in resource-constrained environments. Incorporating a mix of high and low uncertainty texts during selection significantly improved performance, with the AUC-ROC score increasing substantially when using a 50/50 or 70/30 split. For less frequent labels, such as Damaged product, providing pre-labeled texts to initialize the framework improved performance substantially. Randomly selecting data for more common labels achieved results comparable to the active learning framework, but performance degraded significantly for rare labels.
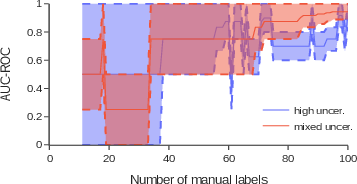
Figure 2: Convergence of AUC-ROC for the Pet product label, using the XGBBoost model, with only high uncertainty and a 70/30 mix of high and low uncertainty.
Conclusion
The Conformal Active Learning framework effectively reduces manual labeling efforts while ensuring reliable predictions. The framework's model agnosticism and open-source implementation facilitate its deployment in healthcare institutions for privacy-preserving, real-time epidemiological surveillance. Future work will focus on testing with real medical data and refining the framework for practical deployment.

