- The paper presents a Progressive Spectrum Diffusion Model (PSDM) that uses compositional prompts to generate synthetic polyp images, improving detection and diagnosis.
- It integrates segmentation masks, bounding boxes, and colonoscopy reports to enhance F1 scores and mean average precision across various datasets.
- The method significantly boosts model performance on standard datasets by addressing out-of-distribution challenges in colorectal cancer screening.
Robust Polyp Detection and Diagnosis through Compositional Prompt-Guided Diffusion Models
Introduction
The paper "Robust Polyp Detection and Diagnosis through Compositional Prompt-Guided Diffusion Models" addresses the challenge of improving polyp detection, classification, and segmentation in colorectal cancer (CRC) screening. Despite advances in deep learning models, they often fall short in diverse clinical environments and with out-of-distribution (OOD) data. Traditional data augmentation methods struggle to capture the complexity of medical images, necessitating innovative approaches for generating diverse and clinically relevant training datasets.
Methodology
The authors introduce a Progressive Spectrum Diffusion Model (PSDM) that exploits compositional prompts created from segmentation masks, bounding boxes, and colonoscopy reports. Compositional prompts enable the integration of coarse and fine-grained clinical annotation, facilitating the generation of synthetic polyp images that reflect real-world variability.
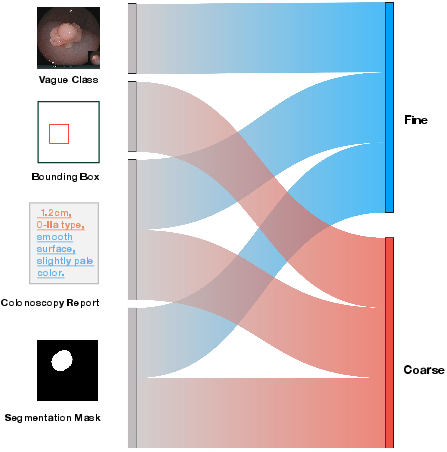
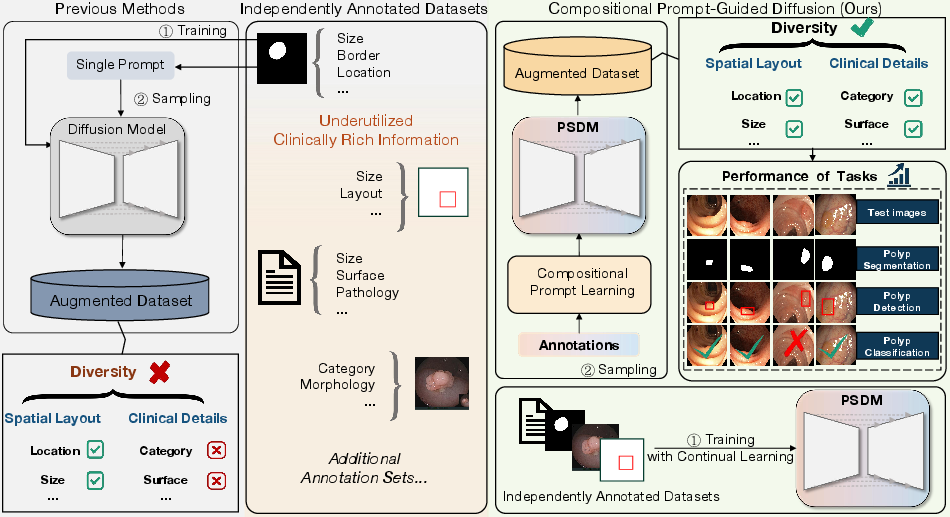
Figure 1: Compositional prompt-guided diffusion framework for generating diverse polyp images. Left: Example prompts with varying levels of granularity. Right: Previous single-prompt methods constrain diversity. In contrast, our PSDM model employs compositional prompts to enhance augmented dataset diversity, resulting in improvements in downstream tasks.
By leveraging advanced data augmentation through PSDM, the paper reports significant improvements across various medical imaging tasks. Specifically, the model enhances F1 scores and mean average precision on the PolypGen dataset by emphasizing the importance of integrating textual and structural annotations.
Results
The augmented dataset generated using PSDM significantly boosted the performance of polyp classification, segmentation, and detection tasks. On standard datasets such as CVC-300 and Clinic-DB, as well as the challenging PolypGen dataset, models trained with PSDM-augmented data consistently outperformed baseline models.
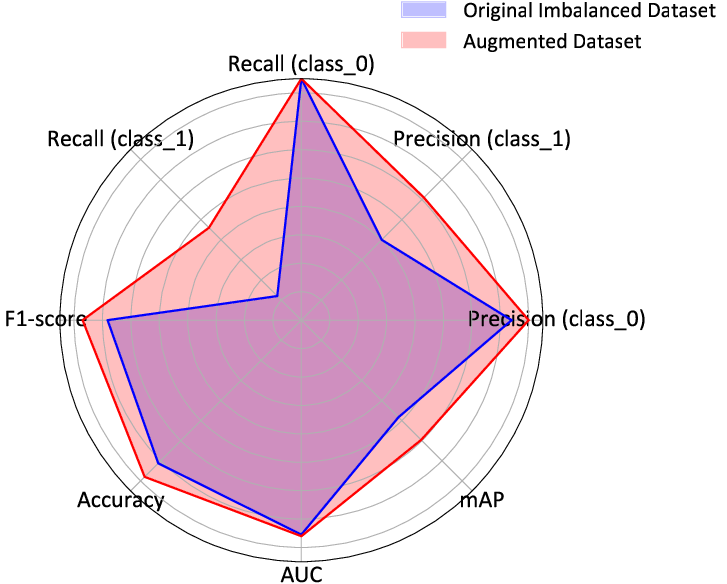
Figure 2: Radar chart illustrating the performance comparison between ResNet models trained on the original imbalanced dataset and the augmented balanced dataset.
The inclusion of text descriptions alongside traditional segmentation led to improved generalization. Notably, ResNet models showcased marked improvements in detecting malignant polyps when trained on a balanced dataset supplemented by PSDM-generated images, as depicted in the radar chart in Figure 2.
Discussion
The introduction of compositional prompts to diffusion models represents a significant advancement in addressing the fidelity and diversity challenges faced by synthetic medical image generation. Through this methodology, the paper highlights the potential of leveraging comprehensive clinical annotations to generate synthetic datasets that enhance model performance in real-world diagnostic tasks.
However, the research acknowledges the lack of standardized evaluation metrics for synthetic medical images, pointing to future work in establishing robust benchmarks for clinical relevance.
Conclusion
By integrating multimodal annotations into a unified diffusion model framework, this research enhances the capacity for generating clinically diverse and accurate polyp images. The approach not only refines data augmentation but also sets a foundation for future applications where synthetic data can bridge gaps in existing clinical datasets, leading to improved diagnostic accuracy in colorectal cancer screening and beyond. The PSDM model drives home the importance of rich, annotated datasets in fortifying model generalization and robustness, paving the way for broader adoption in medical image analysis.


