- The paper demonstrates a novel promptable 3D segmentation model that leverages a large-scale dataset to reduce manual annotation costs by over 85%.
- It introduces a comprehensive network architecture incorporating image and prompt encoders, memory attention, and mask decoders for efficient medical imaging tasks.
- The model achieves superior segmentation performance using the Dice similarity coefficient across complex targets including kidney lesions and pancreas.
MedSAM2: Segment Anything in 3D Medical Images and Videos
MedSAM2 represents an advancement in medical image and video segmentation by leveraging a foundation model tailored for 3D medical contexts. This essay examines its development, architecture, performance, and practical implications for reducing annotation costs in large-scale datasets.
Dataset and Network Architecture
MedSAM2 exploits a substantial dataset comprising 455,000 3D image-mask pairs and 76,000 annotated frames across CT, PET, MRI, ultrasound, and endoscopy (Figure 1). The model architecture extends the Segment Anything Model 2 (SAM2) with a promptable segmentation network, including an image encoder, a prompt encoder, a memory attention module, and a mask decoder. This design facilitates efficient segmentation by capturing spatial continuity in 3D images and videos.
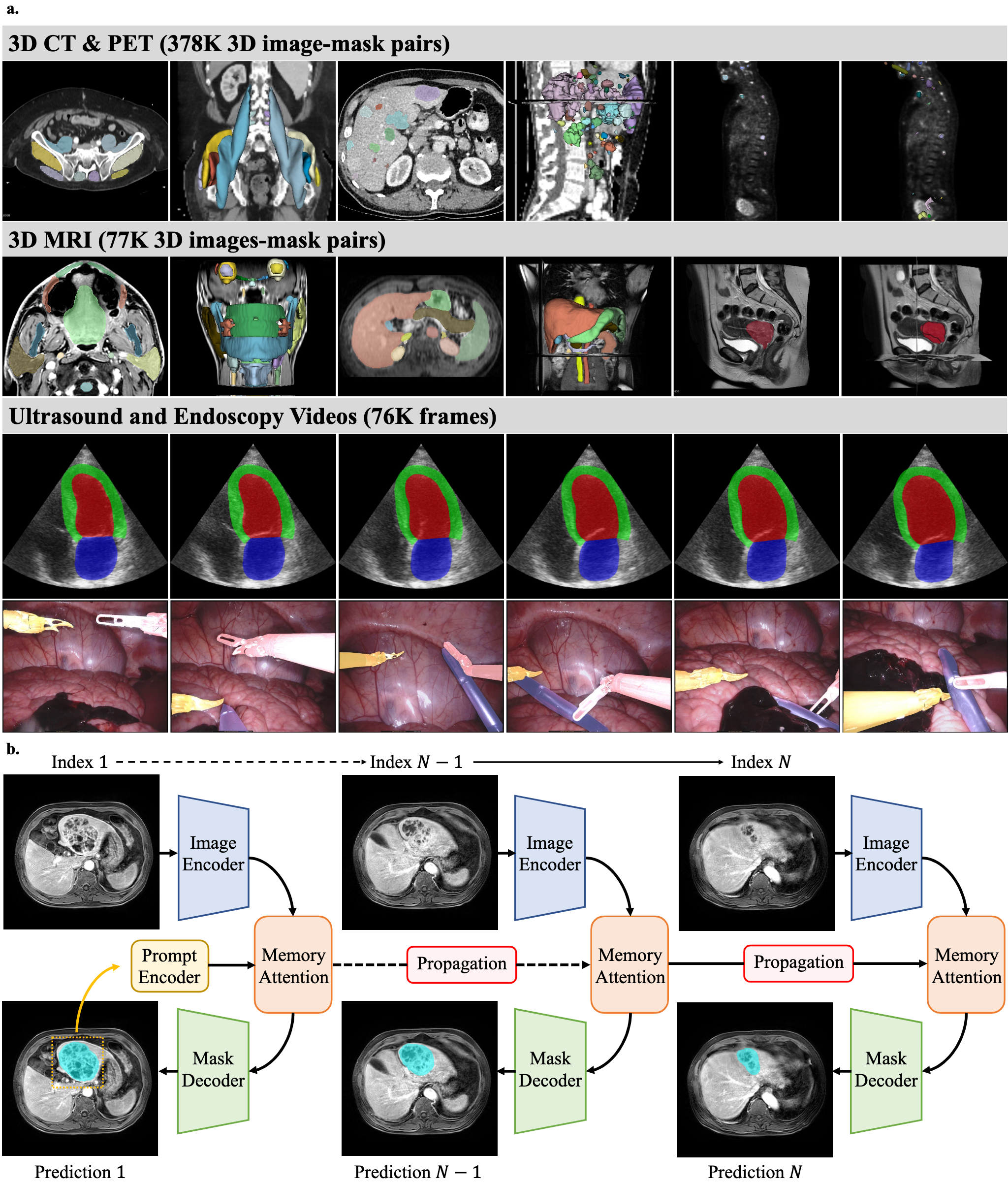
Figure 1: Dataset and network architecture for MedSAM2 development.
MedSAM2's segmentation capabilities were rigorously evaluated against established benchmarks for diverse organs and lesions. Figure 2 illustrates its superior performance across five 3D segmentation tasks using the Dice similarity coefficient. Notably, MedSAM2 surpasses other models such as EfficientMedSAM-Top1, particularly for complex targets like kidney lesions and pancreas, which display significant anatomical variability.
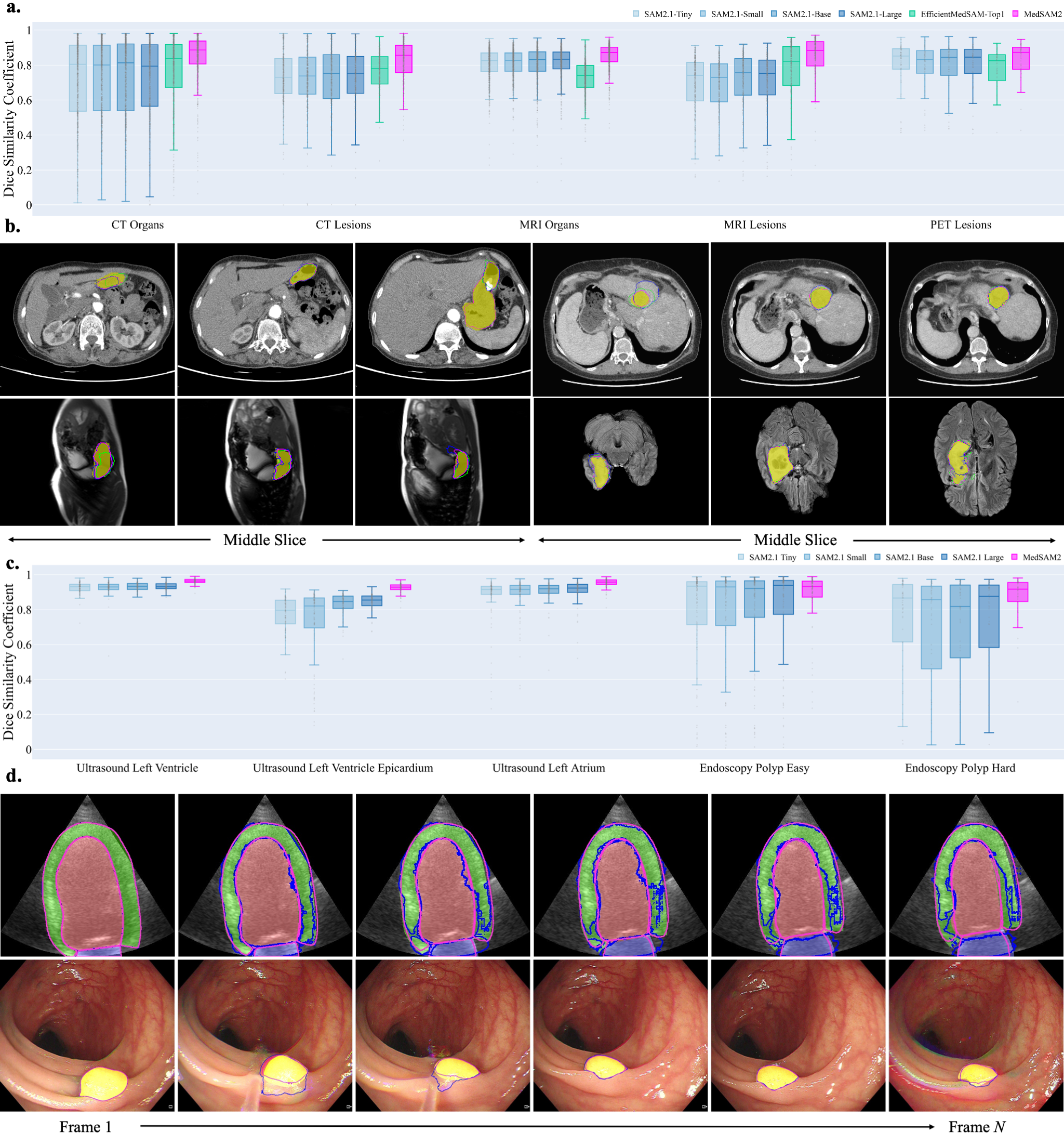
Figure 2: Segmentation performance on hold-out 3D image and video datasets.
Efficient 3D Lesion Annotation
MedSAM2 incorporates a human-in-the-loop pipeline for efficient lesion annotations in CT and MRI scans, demonstrating over 85% reduction in manual segmentation costs. Figure 3 shows the iterative process, achieving significant time savings per annotation round, ultimately advancing from lengthy manual procedures to rapid models like MedSAM2 fine-tuned with domain-specific data.
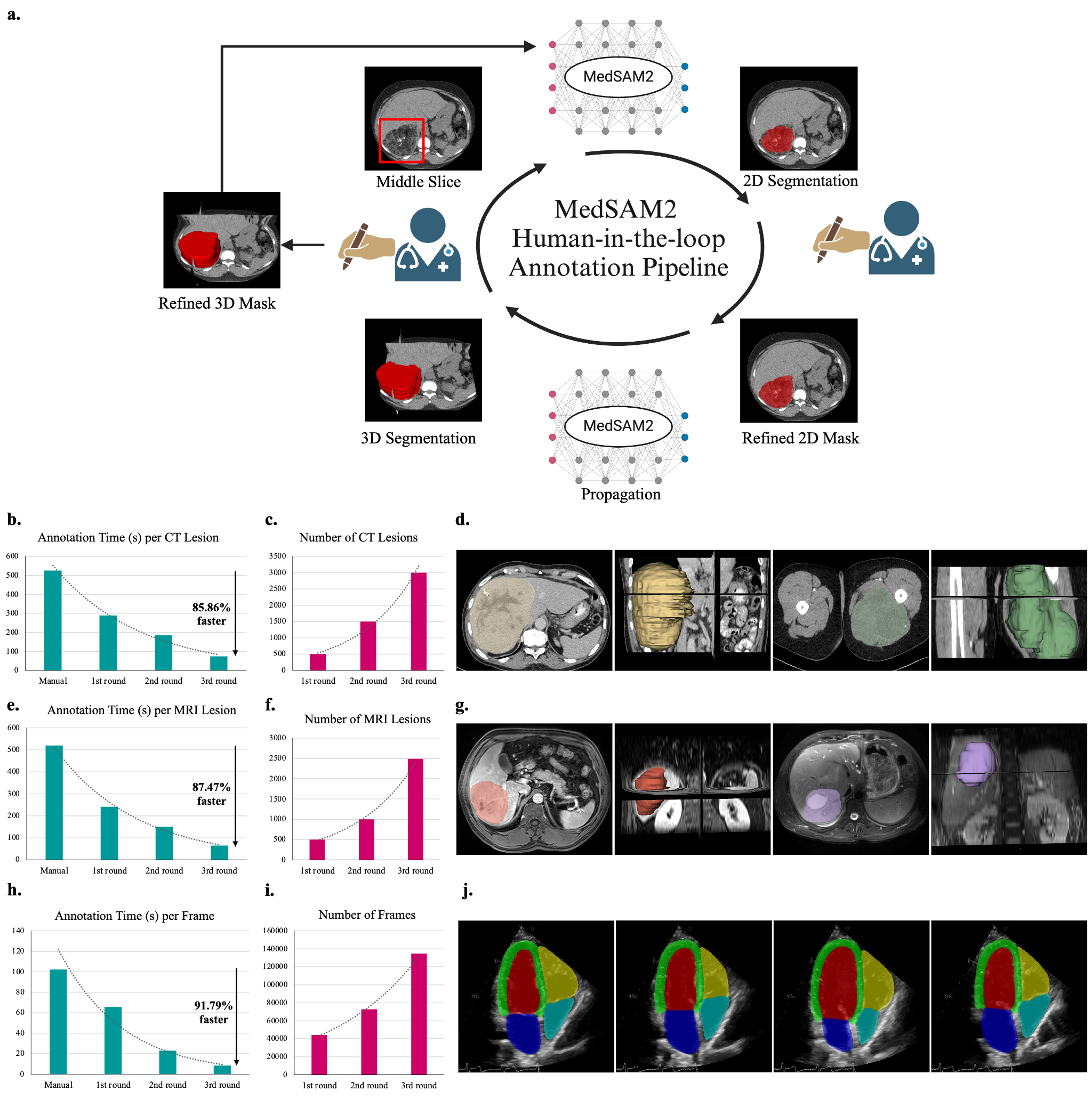
Figure 3: MedSAM2 for efficient lesion annotation in 3D CT and MRI scans.
High-Throughput Video Annotation
MedSAM2 adapts its annotation pipeline for echocardiography videos, effectively handling the dynamic nature of cardiac ultrasound imaging. This method, tailored to tackle motion artifacts and achieve temporal coherence, cuts annotation times significantly, as evidenced in the annotation process of the RVENet dataset (Figure 4).
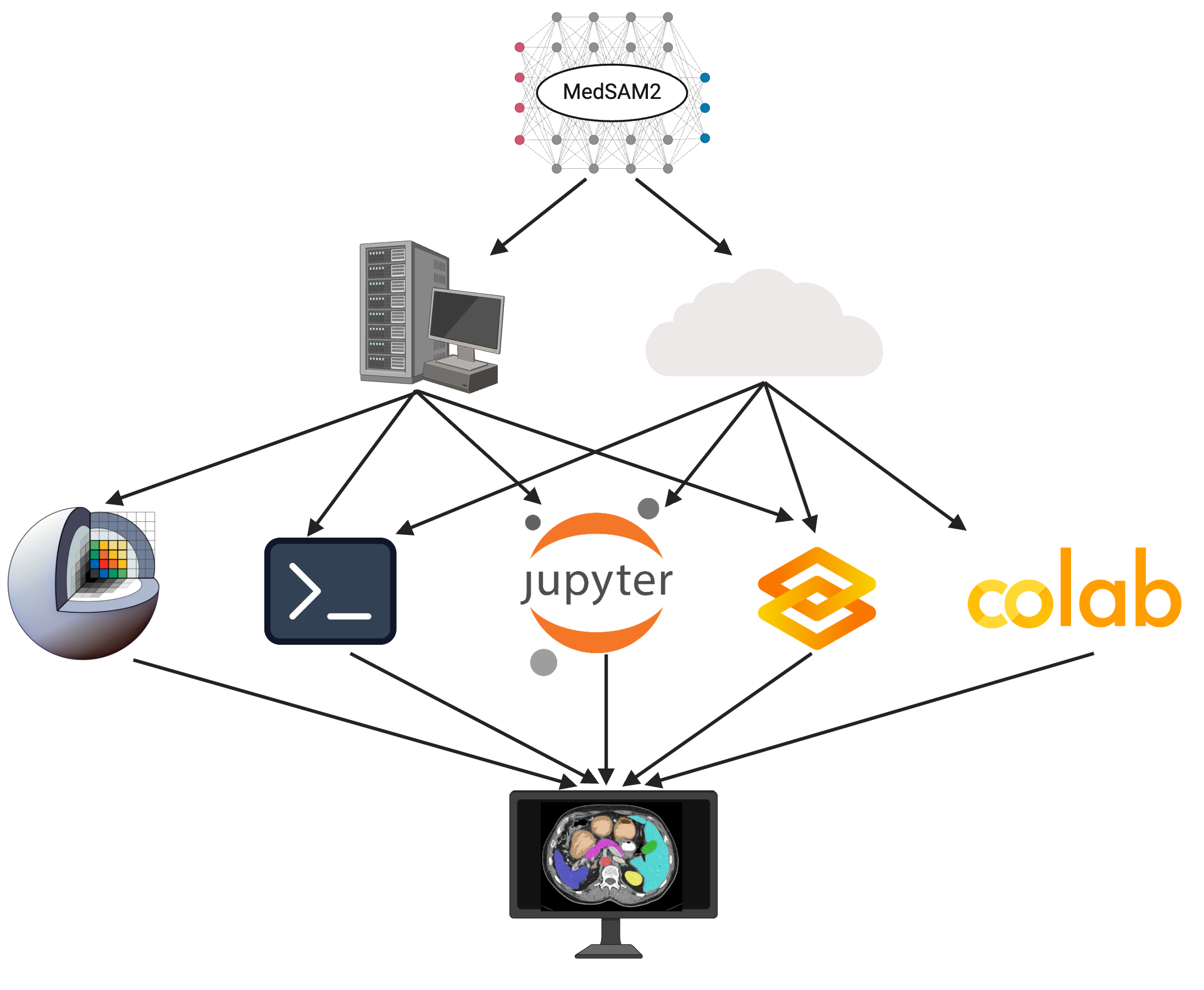
Figure 4: MedSAM2 can be deployed on local desktops and remote clusters with commonly used platforms.
MedSAM2 is implemented across platforms like 3D Slicer, JupyterLab, Gradio, and Google Colab, ensuring community access and easy integration into diverse computational workflows. This flexibility supports varied user needs from clinical researchers to data scientists, further enhancing its utility in both local and cloud settings.
Discussion
MedSAM2 signifies a leap in leveraging foundation models for medical segmentation by addressing domain gaps through transfer learning and efficient interactive designs. The model's scalability and robustness across varied medical imaging modalities underline its potential to streamline clinical workflows, particularly in high-throughput environments like echocardiography and oncology.
While MedSAM2 effectively reduces annotation costs and improves segmentation reliability, its dependency on bounding box prompts limits its application for intricate structures. Future enhancements might focus on expanding prompt types or implementing adaptive memory systems to capture complex motions more adeptly.
Conclusion
MedSAM2's deployment promises a pivotal shift in 3D medical image and video segmentation, enabling more efficient resource utilization, scaling annotated datasets, and enhancing research and clinical applications. Its integration into mainstream platforms paves the way for broader adoption and continued community collaboration in enhancing medical imaging technologies.



