- The paper introduces EquiHGNN, which models complex molecular interactions by leveraging rotational and translational equivariance through hypergraphs.
- The framework integrates symmetry-aware feature initialization and generalized message passing across hyperedges to capture high-order interactions.
- Experimental results show that EquiHGNN outperforms traditional graph models on benchmarks like QM9 and PCQM4Mv2, highlighting its scalability and accuracy.
EquiHGNN: Scalable Rotationally Equivariant Hypergraph Neural Networks
Introduction
The paper presents EquiHGNN, a novel framework designed to model complex high-order interactions in molecular systems using hypergraphs integrated with symmetry-aware features. Unlike traditional graph-based models limited to pairwise connections, EquiHGNN captures multi-way interactions through hypergraphs, enabling a more robust understanding of molecular structures. By enforcing rotational and translational equivariance, this approach ensures that the learned representations are both physically meaningful and consistent with the inherent symmetries of molecular systems.
Equivariant Hypergraph Neural Networks Framework
EquiHGNN leverages hypergraph representations to encode complex molecular interactions such as conjugated π-systems and hydrogen bonding networks. It builds upon the AllSet framework, which formulates hypergraphs as bipartite graphs where nodes represent atoms and hyperedges represent multi-atomic interactions.
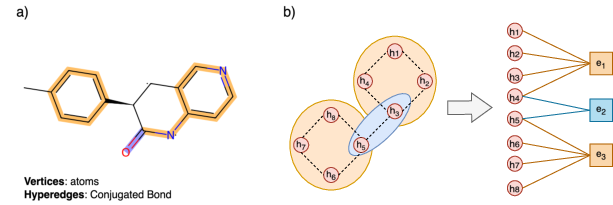
Figure 1: a) Illustration of a hypergraph constructed from a molecule, where vertices represent atoms and hyperedges represent conjugated bonds, highlighted in blue and orange. b) Hypergraph to Bipartite representations.
EquiHGNN initializes node features with both scalar attributes and 3D geometric properties, embedding equivariance naturally. This approach results in a compact and expressive model that captures both topological and geometric information. The framework evaluates three architectural backbones—EGNN for Euclidean space, Equiformer for the Fourier domain, and FAFormer for frame-based symmetry averaging.
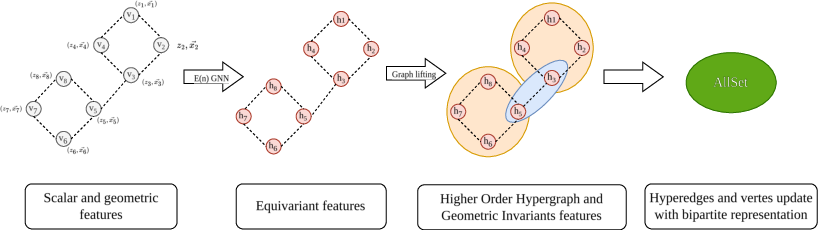
Figure 2: Overview of the Equivariant Hypergraph Neural Network framework.
Methodology
EquiHGNN's methodology emphasizes the integration of geometric deep learning (GDL) techniques to maintain spatial symmetries. It employs equivariant architectures that ensure rotational and translational equivariance, a necessity for accurately modeling molecular properties.
- Feature Initialization: Nodes in the hypergraph are initialized with symmetry-aware geometric representations. Scalar features are derived from molecular properties and coordinates are used to capture spatial relationships.
- Message Passing: The model generalizes message-passing neural networks (MPNNs) to hypergraphs, enabling the capture of high-order interactions. This is achieved through a bipartite graph structure where hyperedges aggregate information from connected nodes and then propagate updated features back to those nodes.
- Architectural Variants: The study compares three equivariant methods:
- EGNN: Utilizes invariant scalar features with coordinate transformations, maintaining the structure through distance-preserving mappings.
- Equiformer: Provides rotational equivariance through spherical harmonics, enabling nuanced geometric learning.
- FAFormer: Uses frame averaging to maintain equivariant properties, integrating attention mechanisms to weigh the importance of spatial interactions.
Experimental Results
EquiHGNN was benchmarked against both traditional 2D graph models and other hypergraph approaches across multiple datasets, including QM9, OPV, PCQM4Mv2, and Molecule3D.
- QM9 Dataset: Demonstrated significant performance improvements, particularly in tasks involving complex geometric relationships. The Equiformer-MHNN variant achieved superior accuracy for predicting ϵHOMO.
- OPV Dataset: Showcased scalability, particularly for larger polymer molecules. Equiformer-MHNN outperformed existing models in three polymer-related tasks, underscoring the scalability and expressiveness of the framework.
- Large-scale Datasets (PCQM4Mv2 and Molecule3D): Highlighted the robustness of EquiHGNN in handling large and complex molecular graphs. EGNN-MHNN achieved the best performance on the PCQM4Mv2 dataset, with significant reductions in MAE compared to baseline models.
Conclusion
EquiHGNN provides a scalable and robust framework for molecular modeling by integrating hypergraph representations with symmetry-aware features. This approach effectively captures both high-order interactions and geometric consistency, outperforming existing models, especially for larger molecular structures. The straightforward integration with various equivariant techniques highlights its adaptability and potential for future development in capturing complex molecular interactions.
Future Work
Future work will focus on exploring additional high-order interactions beyond conjugated bonds, such as leveraging ETNNs and SE3Set methodologies to enhance the expressiveness of hypergraph neural networks. Integrating more advanced equivariant architectures could further boost computational efficiency and generalization capabilities, offering promising avenues for both algorithmic improvements and practical applications in molecular sciences.

