- The paper introduces PharmaDiff, a novel diffusion model that integrates atom-level pharmacophore hypotheses for precise ligand-based de novo drug design.
- It employs an E(3) equivariant graph Transformer with inpainting and cross-attention to enforce 3D pharmacophoric constraints during molecular generation.
- Experiments on the GEOM-Drugs dataset demonstrate improved pharmacophore match scores and docking performance compared to SMILES-based and structure-based methods.
Pharmacophore-Conditioned Diffusion Model for Ligand-Based De Novo Drug Design
Introduction
The paper introduces PharmaDiff, a generative model designed to harness the potentials of diffusion models for ligand-based drug design. Focusing on 3D molecular generation, PharmaDiff integrates an atom-based representation of pharmacophore hypotheses into its generative process, thereby enabling precise conditioning of 3D molecular structures to predefined pharmacophoric constraints. This approach is a significant advancement over traditional methods which typically rely on either structure-based methods requiring protein target structures or ligand-based methods limited by 2D QSARs.
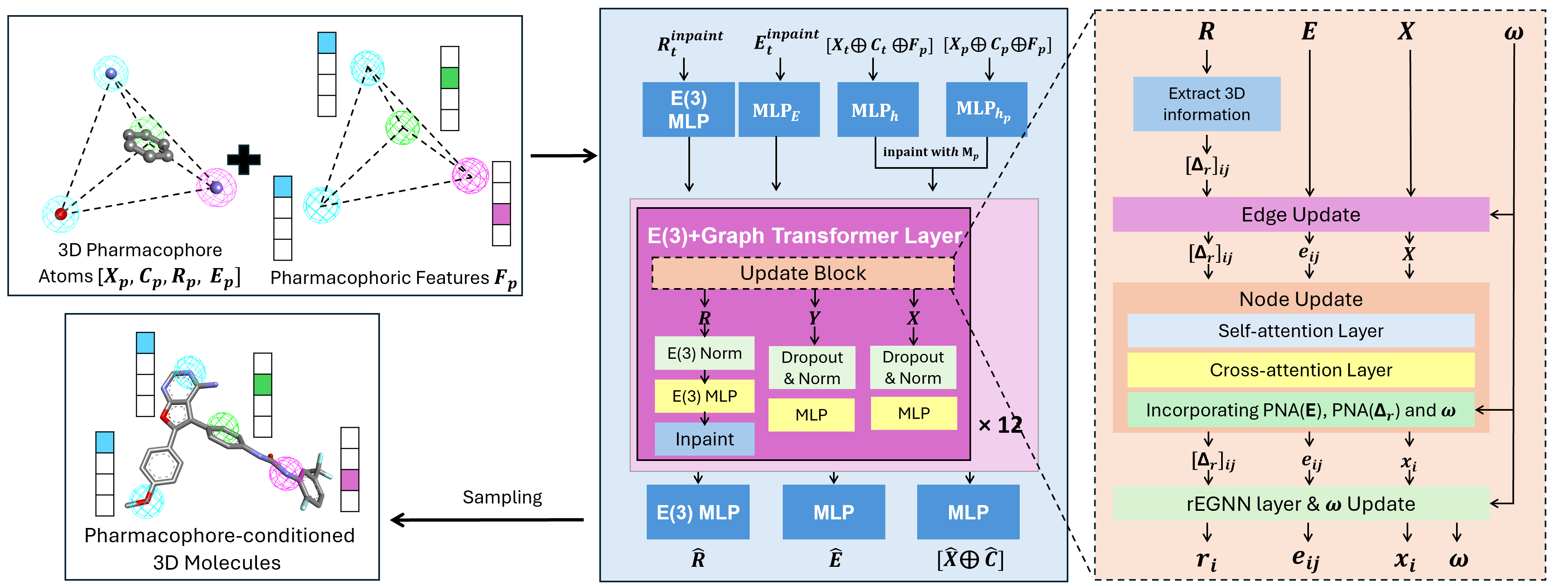
Figure 1: PharmaDiff's architecture overview simultaneously predicting 2D and 3D coordinates conditioned on a pharmacophore hypothesis. The model uses 12 layers of an E(3) graph Transformer architecture, designed to maintain SE(3) equivariance, it used inpainting and a cross-attention layer between molecular and pharmacophore Node Embeddings to enforce pharmacophore constrains.
Model Architecture
PharmaDiff's architecture builds upon the E(3) graph Transformer style, integrating pharmacophore information into the transformation blocks. Specifically, it uses a dedicated multilayer perceptron (MLP) to encode pharmacophoric features, which are then utilized in inpainting, cross-attention, and E(3)-equivariant blocks to ensure that generated molecules conform to defined pharmacophoric constraints.
The use of inpainting allows for the enforcement of pharmacophore-associated atoms within the generative process, albeit with challenges in maintaining atomic connectivity during this process. Cross-attention layers provide dynamic integration between synthesis features and pharmacophore-specific constraints, permitting the generation of molecular graphs that align with intricate 3D pharmacophore patterns.
Noise and Loss Modeling
PharmaDiff uses a noise model that stays true to E(3) symmetries, ensuring the generative process respects rotational and translational invariances crucial for 3D molecular data. The loss function combines MSE for coordinate regression and cross-entropy for categorical features (such as atom types), augmented by pharmacophore-specific penalties to reinforce the generation of designated pharmacophore features.
Experiments and Results
The model was tested on the GEOM-Drugs dataset, aiming to synthesize new drug-like molecules that conform to pharmacophore hypotheses derived from real-world drug-like conformers.
Ligand-Based Drug Design
PharmaDiff outperformed existing methods in achieving higher pharmacophore match scores (MS) and perfect match rates (PMR), indicating superior fidelity to the 3D pharmacophore hypothesis. This advantage is particularly pronounced over models that utilize solely SMILES-based generation techniques, highlighting the effect of directly integrating 3D conditioning.
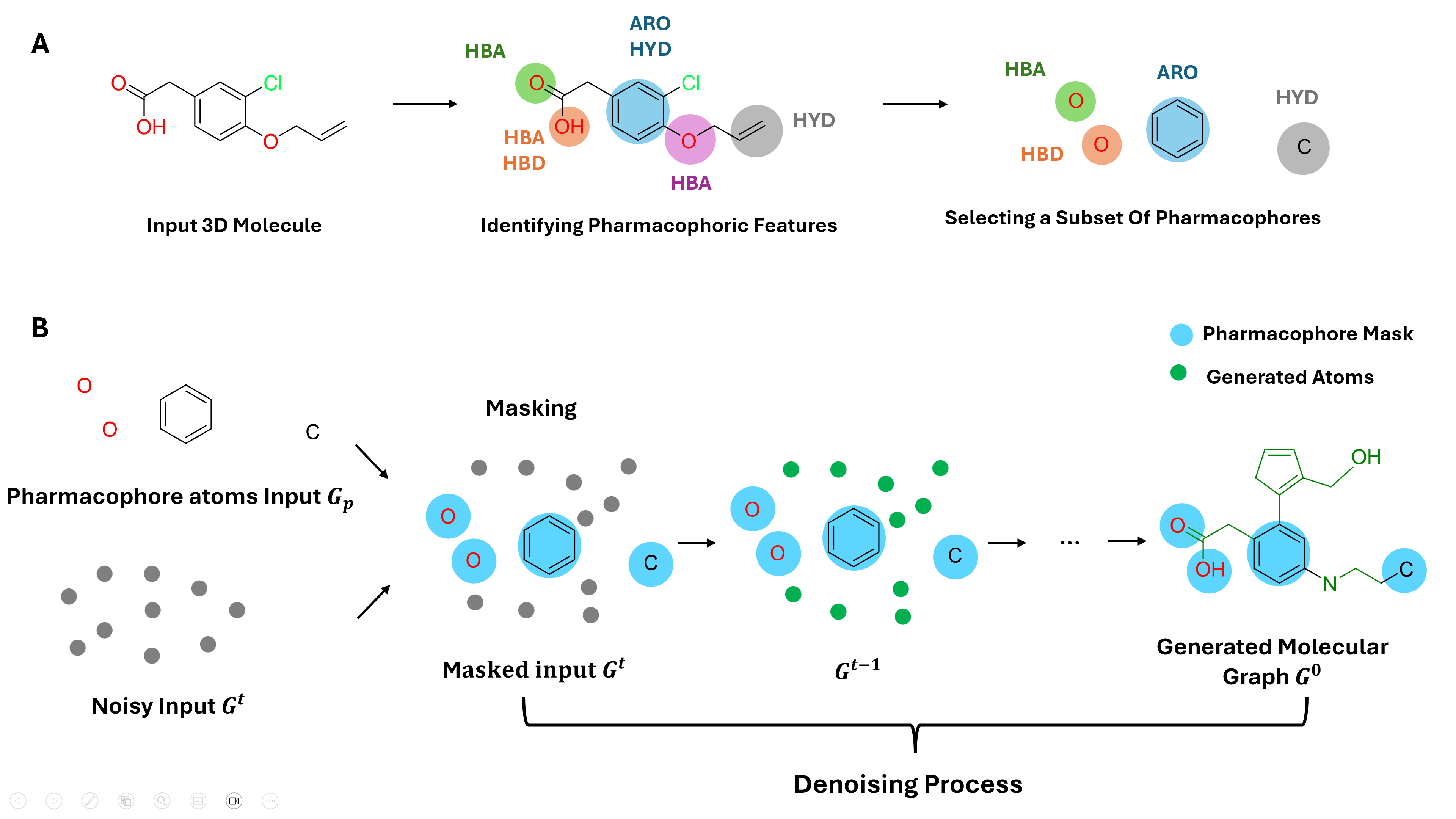
Figure 2: (A) Decomposition of molecular structures into 3D pharmacophore-associated atoms. (B) Conditioning the PharmaDiff model during the denoising process using the pharmacophore graph Gp. The pharmacophoric features shown include hydrogen bond acceptor (HBA), hydrogen bond donor (HBD), hydrophobic (HYD), and aromatic (ARO) groups.
Structure-Based Drug Design
In structure-based applications, PharmaDiff demonstrated superior performance in docking scores against proteins' active sites without reliance on detailed protein structures, outperforming structure-based generative models by producing more synthetically accessible and pharmacophore-accurate molecular structures.
Conclusion
PharmaDiff establishes a new benchmark for pharmacophore-conditioned molecular generation, merging the rigors of 3D pharmacophore hypothesis adherence with the generative prowess of diffusion models. By circumventing the reliance on protein structure data, it broadens the applicability of de novo drug design to novel targets, making it a promising tool for areas like orphan drug development and rapid-response drug design strategies. Future work should focus on enhancing connectivity in generated molecules and explicitly encoding pharmacophoric features to retain key functionalities essential for biological activity.

