- The paper presents a novel biofidelic architecture by converting the Drosophila larval connectome into a fixed recurrent network.
- The BPU achieved 98% accuracy on MNIST and 58% on CIFAR-10, and excelled in chess puzzles versus comparable MLP and Transformer models.
- Scaling via a degree-corrected stochastic block model preserved biological wiring, highlighting potential for advanced, bio-inspired AI systems.
Biological Processing Units: Leveraging an Insect Connectome
Introduction
The paper "Biological Processing Units" explores a novel approach to artificial intelligence by leveraging the connectome of the Drosophila larva. Researchers converted the wiring diagram of this insect brain into a Biological Processing Unit (BPU), a fixed recurrent network derived directly from synaptic connectivity. The unmodified BPU, consisting of approximately 3000 neurons and 65,000 synaptic weights, demonstrated remarkable performance in tasks such as image classification (MNIST and CIFAR-10) and decision-making (ChessBench), surpassing size-matched multilayer perceptrons (MLPs) even without task-specific adaptations.
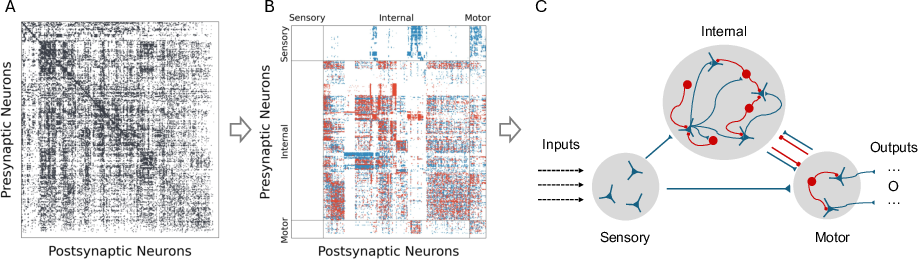
Figure 1: Biological Processing Unit (BPU) architecture based on the larval Drosophila connectome.
Methods
BPU Architecture
The BPU design is based on the Drosophila larval connectome, utilizing an adjacency matrix derived from electron microscopy reconstructions. Synaptic weights are directly sourced from the connectome, remaining unchanged during training. The neurons are partitioned into three pools: sensory, output, and internal. The architecture features a fixed-weight recurrent core that propagates the activity over unrolled steps, with trainable input and output projections optimized via gradient descent. This setup captures the connectome's biological dynamics faithfully without structural modifications.
Connectome Expansion
Scaling the BPU further was achieved using a directed, signed degree-corrected stochastic block model (DCSBM). By preserving the block-level wiring statistics and synaptic polarity, researchers expanded the BPU up to 5×. This approach allowed the investigation of how size influences performance, providing insights into potential advantages of larger biological circuitry.
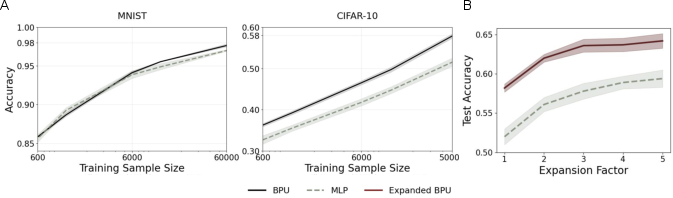
Figure 2: Test accuracy on MNIST and CIFAR-10 for the original connectome-derived BPU and its expanded versions.
Results
Image Classification
The BPU performed competitively across image classification tasks, achieving 98% accuracy on MNIST and 58% on CIFAR-10, outperforming a similarly parameterized 2-layer MLP baseline. When the connectome was expanded using DCSBM, CIFAR-10 accuracy improved further, showcasing the advantages of leveraging detailed biological structures.
(Figure 3)
Figure 3: Test accuracy on MNIST and CIFAR-10 with modality-restricted reservoirs.
Chess Puzzle Solving
The BPU was tested on chess puzzles using both graph neural networks (GNN) and convolutional neural networks (CNN) for encoding. The models exhibited impressive puzzle-solving accuracy, with the GNN--BPU achieving performance levels comparable to smaller Transformer models and the CNN--BPU outperforming a parameter-matched Transformer baseline. Moreover, when enhanced with minimax search and alpha-beta pruning during inference, the CNN--BPU reached 91.7% accuracy, surpassing a 9M-parameter Transformer.
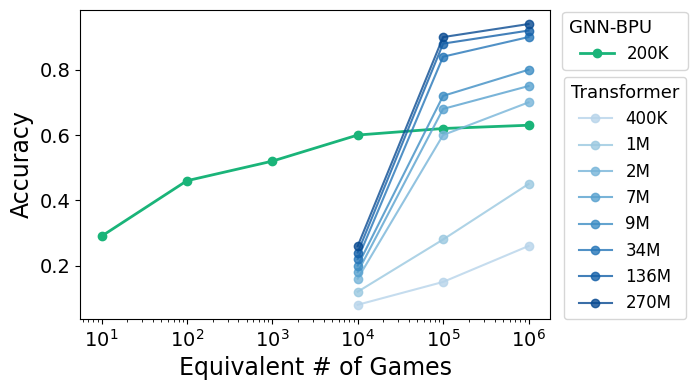
Figure 4: Puzzle-solving accuracy (%) with GNN--BPU model and ChessBench reference models of multiple sizes.
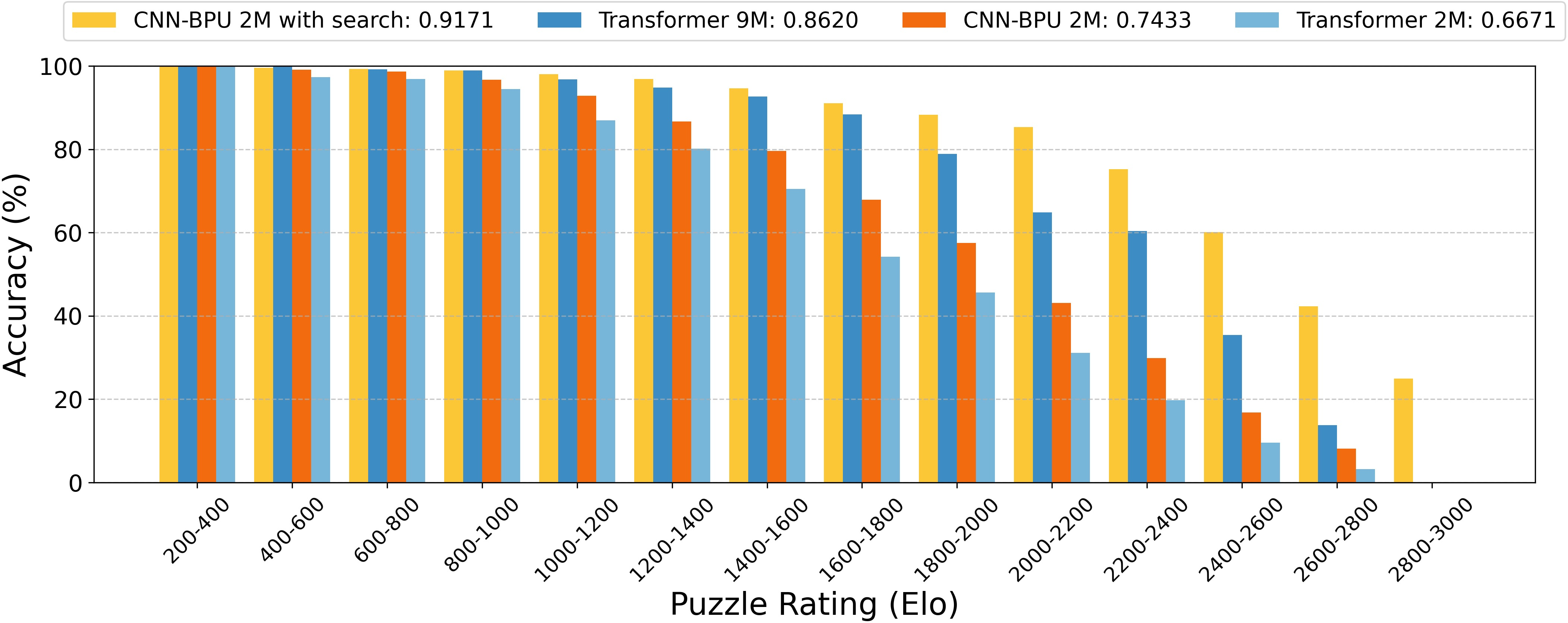
Figure 5: Bars show the percentage of puzzles solved correctly within each Elo bin using CNN-BPU with search.
Discussion
The results presented in the paper suggest that the full Drosophila larval connectome effectively supports complex cognitive tasks, highlighting its latent computational capacity without structural modifications. The findings underscore the potential of biofidelic neural architectures, where biologically evolved circuitry serves as reusable substrates for intelligent computation. Future research may involve exploring task-specific adaptations and investigating the functional specialization of different connectome substructures. As more comprehensive connectomic data becomes available, the principles established here could scale to larger and more cognitively complex brain structures, potentially paving the way for advanced AI systems inspired by biological networks.
Conclusion
This research demonstrates the viability of employing biologically evolved circuits from the Drosophila larval connectome as neural substrates for AI. By adhering closely to the original neural architecture and exploring both image classification and chess puzzle tasks, the BPU showcases robust performance, motivating future scaling to more intricate connectomes.



