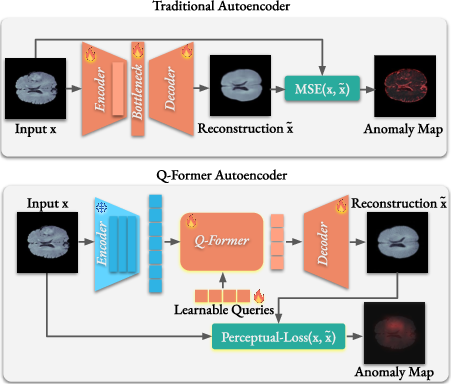
- The paper introduces an unsupervised Q-Former Autoencoder that combines pretrained encoders, a Q-Former bottleneck, and perceptual loss for semantically-guided medical anomaly detection.
- It leverages features from models like DINOv2 and Masked Autoencoder to achieve state-of-the-art AUROC scores of 94.3% on BraTS2021 and 83.8% on RSNA.
- The framework emphasizes scalability via frozen pretrained models while addressing computational demands and potential adaptation challenges for diverse clinical applications.
Introduction
The paper introduces "Q-Former Autoencoder: A Modern Framework for Medical Anomaly Detection," which presents an unsupervised autoencoder-based model integrated with cutting-edge pretrained vision models to enhance anomaly detection in medical imaging. The proposed architecture leverages foundation models like DINO, DINOv2, and Masked Autoencoder to extract features, utilizing a Q-Former as the bottleneck to maintain high-level feature representation and applying perceptual loss for semantically guided anomaly detection.
Methodology
Architecture
The Q-Former Autoencoder (QFAE) architecture comprises three main components: a pretrained encoder, a Q-Former as the bottleneck, and a lightweight Transformer-based decoder. The pretrained models, serving as encoders, capture rich feature representations without domain-specific fine-tuning. The Q-Former aggregates multi-scale features and provides a fixed-length output regardless of input variability. This, coupled with perceptual loss guided by a Masked Autoencoder, focuses on semantic rather than pixel-level detail, refining the network's ability to generalize anomalies beyond what standard MSE-driven autoencoders accomplish.
Training and Loss
The model is trained using perceptual loss derived from features of a pretrained Masked Autoencoder, which aligns reconstruction toward meaningful structures rather than purely minimization of pixel-level error. This approach provides nuanced detection of anomalies, capitalizing on differences in high-level semantics that are characteristic of normal versus abnormal imagery.
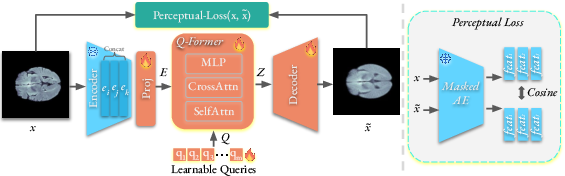
Figure 2: The training of our Q-Former Autoencoder for medical anomaly detection.
Results
The QFAE was evaluated across diverse benchmarks including BraTS2021, RESC, and RSNA datasets, showcasing significant improvements in anomaly detection capability, evidenced by its state-of-the-art AUROC scores: 94.3% for BraTS2021 and 83.8% for RSNA. These results reflect the architecture's effectiveness in capturing and differentiating subtle semantic nuances across various imaging modalities, all without additional domain-specific tuning.
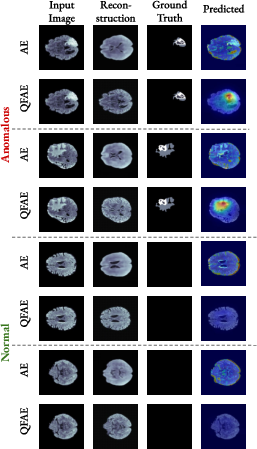
Figure 3: Qualitative examples of anomaly localization on several samples from the BraTS2021.
Implementation Considerations
Scalability and Resource Requirements
The model leverages frozen pretrained encoders, reducing comprehensive training resource constraints. However, the necessity to compute high-dimensional perceptual loss implies significant memory usage, particularly for multi-scale feature extraction. Deployment when translating to real-world systems necessitates consideration of computational power, especially within clinical settings where real-time processing may be imperative.
Limitations
The model's reliance on pretrained vision encoders, while democratizing feature extraction, does not cater to every domain nuance without imaging diversity inherent in foundational model training sets. Processing efficiency and robustness across unexpected anomaly variations require ongoing exploration and possible architectural adaptations to expand its clinical application viability.
Conclusion
The Q-Former Autoencoder represents an advanced leap in unsupervised anomaly detection, emphasizing pretrained models for efficient feature extraction and perceptual loss for semantically driven training. Though demonstrating exceptional performance across multiple datasets, adaptation challenges and computational demands highlight areas for future development. Integration with ongoing advancements in foundational vision models promises even broader applicability across more diverse medical domains.