- The paper introduces GDAIP, a framework that leverages GAT and MME loss for cross-dataset brain parcellation to improve domain adaptation.
- The framework constructs brain graphs from fMRI data and applies semi-supervised training with 5% labeled target vertices to enhance accuracy.
- Evaluations using Dice coefficients and Pearson correlation metrics demonstrate GDAIP’s superior ability to capture individual brain functional organization.
A Graph-Based Domain Adaptive Framework for Individual Brain Parcellation with GDAIP
The paper "GDAIP: A Graph-Based Domain Adaptive Framework for Individual Brain Parcellation" presents a sophisticated methodology for addressing domain shifts in cross-dataset scenarios using fMRI data to generate individual brain parcellations. This research introduces a framework integrating Graph Attention Networks (GAT) and Minimax Entropy (MME)-based domain adaptation for constructing accurate individual brain parcellations by leveraging graph-based representations and semi-supervised training techniques.
Introduction to Cross-Dataset Brain Parcellation
Functional MRI (fMRI) provides a dynamic view of brain activity by capturing blood-oxygen-level-dependent (BOLD) signals, forming a pivotal tool in cognitive neuroscience. Traditional node-based analysis reduces fMRI data dimensionality, leveraging predefined atlases to delineate brain regions and extract functional network features. However, group-level atlases used in these analyses often overlook individual variability, necessitating methods for personalized brain parcellation. This paper leverages deep learning advancements to propose the Graph Domain Adaptation for Individual Parcellation (GDAIP) framework, addressing significant domain shifts seen across different datasets.
Methodology Overview
The GDAIP framework involves constructing brain graphs from fMRI data, representing brain vertices connected by adjacency relationships derived from cortical surfaces (Figure 1). Utilizing a semi-supervised GAT model, GDAIP combines representations from both group-level and individual brain graphs to facilitate cross-dataset transfer learning. During training, labeled vertices from source datasets and 5% from target datasets are deployed for computing cross-entropy losses, while an MME loss governs unlabeled target vertices.
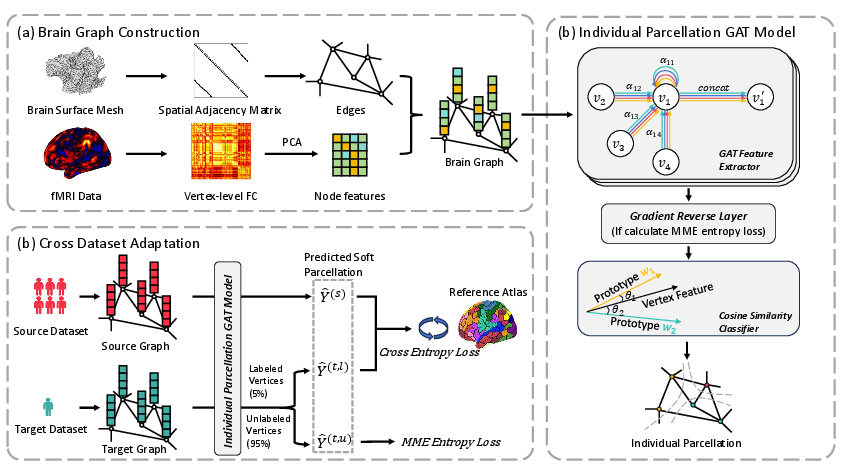
Figure 1: Overview of the GDAIP framework. (a) Brain graphs are constructed using surface-based adjacency and functional fingerprints from fMRI data. (b) The parcellation model includes a GAT-based feature extractor and a cosine similarity classifier, with a Gradient Reversal Layer enabling adversarial MME loss. (c) During training, graphs from source and target datasets are input to the individual parcellation model. Cross-entropy loss is computed on source and 5\% labeled target vertices, while MME loss is applied to the remaining unlabeled target vertices for cross-dataset adaptation.
Graph Construction and GAT Model
In constructing brain graphs, vertices represent cortical regions, and edges manifest through adjacency matrices defining spatial proximity on cortical surfaces. Feature vectors are obtained via PCA from vertex-level functional connectivity matrices, providing dimensionality reduction. The GAT model extracts vertex features through attention mechanisms, enhancing discriminative power and adaptability. Use of a cosine similarity-based classifier further improves generalization, with learned class prototypes organized across region of interests (ROI).
Semi-supervised Training and MME Domain Adaptation
GDAIP employs a semi-supervised learning strategy incorporating MME to improve cross-dataset prediction accuracy. Adversarial optimization through entropy regularization enhances feature separability, a pivotal aspect in the framework’s domain adaptation strategy. Training involves leveraging group-level priors through labeled samples and adapting to individual-specific patterns using semi-supervised settings.
Experimental Analysis
Evaluations were conducted using datasets from the HCP Young Adult and NKI Rockland Samples, assessing adaptation capabilities and functional homogeneity of individual parcellations. Dice coefficient and Pearson correlation metrics were used to quantify intra- and inter-subject parcellation consistency (Figure 2), illustrating the framework's competency in reflecting individual functional organization.
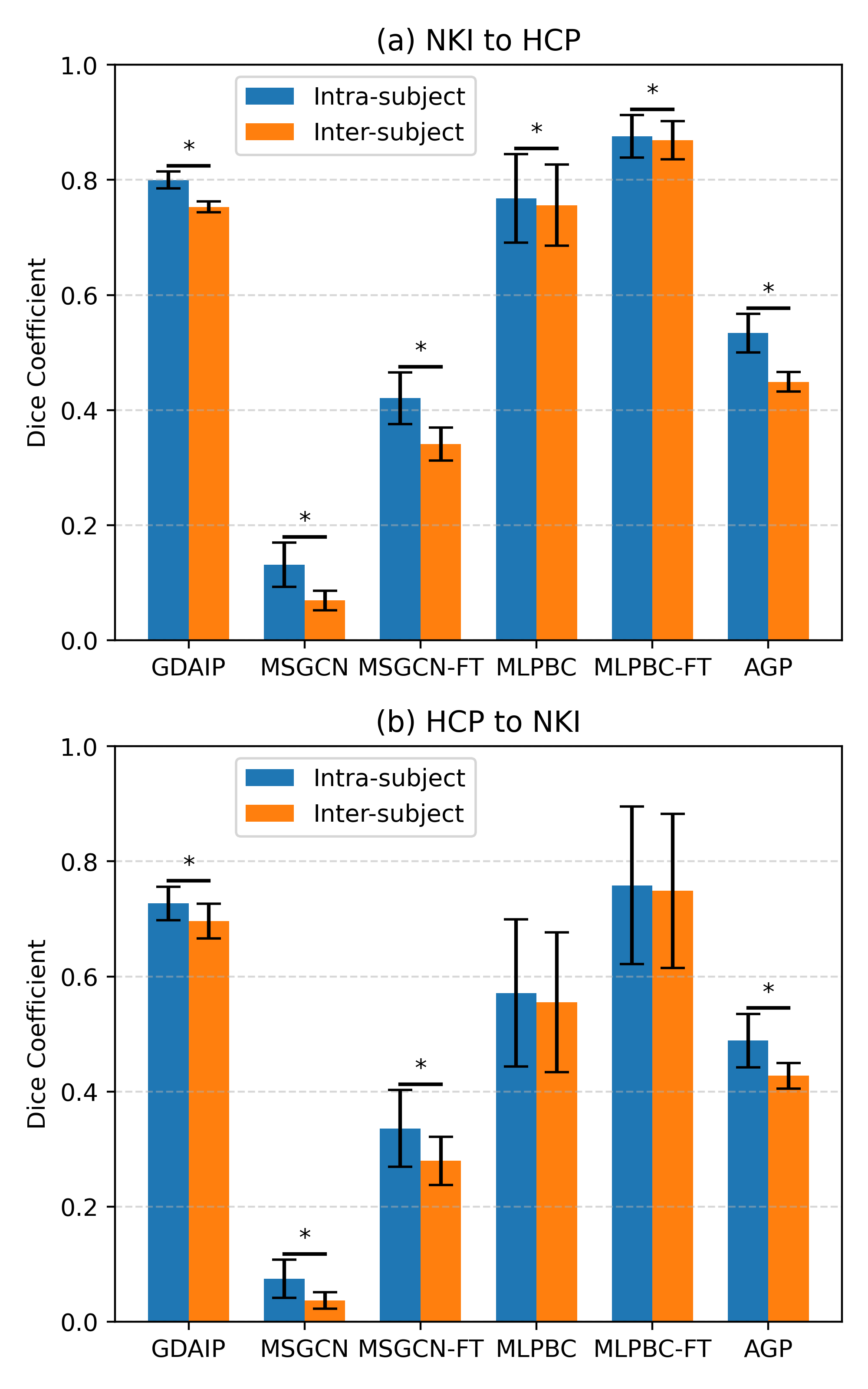
Figure 2: Results of intra- and inter-subject Dice coefficient of individual parcellations across different methods.
Comparative Results and Visualizations
Visualization comparisons between GDAIP-generated parcellations and baseline methods revealed distinct differences, emphasizing the superiority of GDAIP in capturing individual-specific patterns while retaining spatial organization relative to reference atlases (Figure 3).
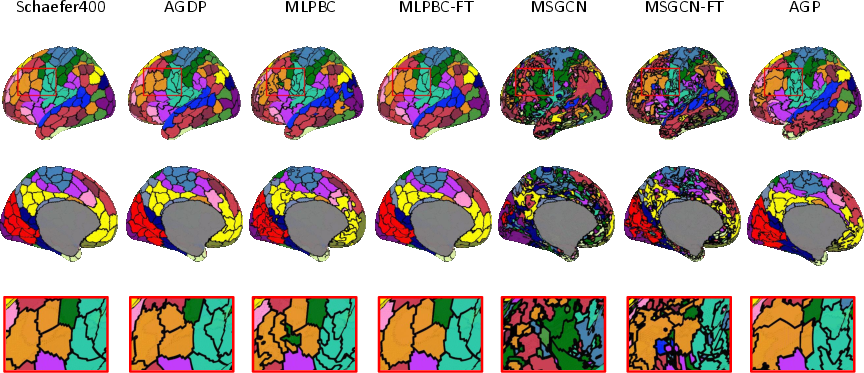
Figure 3: Visualization of the reference Schaefer400 atlas and individual parcellations from different methods. Parcellations are generated for subject 100408 from the HCP dataset, using the NKI dataset as the source domain.
Implications and Future Directions
The study demonstrated that deep learning-based methods must consider domain shifts for practical real-world applicability in individualized brain parcellations. GDAIP’s ability to leverage group-level priors while adapting to subject-specific functional characteristics shows promise for applications in precision medicine. Future research could focus on refining transfer learning methods further to optimize computational efficiency and uncover deeper insights into brain functional topology.
Conclusion
GDAIP successfully implements cross-dataset adaptation in individual brain parcellation, leveraging group-level brain graph priors while generating individualized results reflective of functional organization. Its integration of GAT and MME strategies offers a robust solution to a longstanding challenge in brain dynamics analysis using fMRI, paving the way for impactful applications in neuroscience research and clinical settings.


