- The paper presents a novel method that parallelizes MCMC using a parallel form of Newton’s method for rapid convergence.
- It introduces quasi-Newton and DEER approaches that cut time complexity by decoupling sequential MCMC steps, enhancing efficiency across Gibbs, MALA, and HMC sampling.
- The techniques enable efficient Bayesian computation and real-time updating by leveraging parallelism to handle long sequence dependencies.
Parallelizing MCMC Across the Sequence Length
Introduction
The paper proposes novel methodologies for parallelizing Markov Chain Monte Carlo (MCMC) algorithms across sequence length, aiming to reduce the time complexity traditionally associated with these inherently sequential processes. Utilizing modern hardware capabilities and advances in parallel computing, particularly parallel versions of Newton’s method, this paper presents methods for effectively parallelizing well-established MCMC techniques such as Gibbs sampling, Metropolis-adjusted Langevin algorithm (MALA), and Hamiltonian Monte Carlo (HMC).
Methodology
Parallel Evaluation of Nonlinear Recursions
The cornerstone of this approach is reimagining nonlinear state sequences as solutions to fixed-point equations and applying a parallel form of Newton’s method to resolve these. This allows the decoupling of MCMC iterations across the chain length, promoting parallel computation.
The foundational algorithm, DeepPCR (termed DEER in the context of MCMC), facilitates this by operating with a parallel-in-time formulation of Newton's method, achieving convergence within a logarithmic factor relative to the sequence length. This method efficiently leverages the structure of nonlinear dynamical systems to generate quick, approximate updates that progressively refine the solution until convergence.
Innovations in Parallel MCMC
The paper introduces quasi-Newton methods to combat the computational resource strains induced by traditional Newton’s method, offering a linearly efficient alternative for systems with large dimensionalities. These variants, employing approximations like memory-efficient quasi-DEER, substantially reduce operational memory footprints and runtime by using stochastic estimates to forecast the Jacobian diagonal, rather than computing it fully.
Additionally, the implementation of early-stopping criteria allows for further acceleration by halting iteration sequences once the approximation nears high-fidelity outputs relative to sequential sampling, offering trade-offs between computational overhead and sample accuracy.
Specific Implementations
- Gibbs Sampling: The reparameterization and subsequent application of DEER unify this process, allowing for theoretically driven samples through a deterministic route driven by input randomness.
- MALA: The pseudo code reinterprets the position updates and integrates non-differentiable Metropolis criteria through smooth approximations, utilizing parallel computing for rapid update sequences.
- HMC: The approach extends to leapfrog integrals, enabling either parallelization of integration steps or full-sample-to-sample transformations.
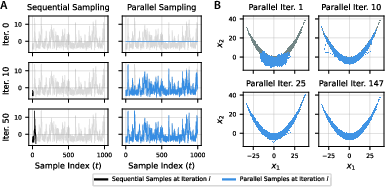
Figure 1: Parallel evaluation of 100K HMC samples targeting the Rosenbrock distribution. Dark lines represent sequential samples, while blue lines indicate parallel samples firming convergence highlights.
Numerical Experiments: Implementations demonstrated enhanced performance across several benchmarks. Particularly, parallel computations reduce time complexities significantly, with quasi-DEER achieving up to 10x speedups compared to sequential approaches, highlighting the efficacy of the method across various dimensionality and complexity paradigms.
Performance tests on both simple models, like Bayesian logistic regression, and complex multimodal distributions, ensured robustness across a spectrum of potential scenarios that MCMC techniques are employed for in practical applications.
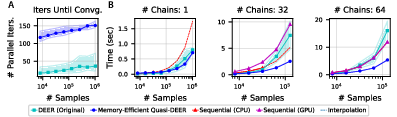
Figure 2: Parallel Gibbs sampling. Shows the scalability and efficiency of the DEER algorithm for large chain lengths.
Conclusion
By unveiling methods to parallelize MCMC over sequence length, the paper contributes a significant advancement towards more efficient Bayesian computation and inference. These methods stand to revolutionize analytics within high-dimensional probabilistic models, significantly reducing the resource barriers that have traditionally constrained MCMC applications. The results indicate that leveraging parallel Newton iterations ensures best-in-class performance for modern MCMC needs, with potential applications in Bayesian learning frameworks and situations demanding rapid sample generation and assessment. The methodologies proposed herein expand the computational boundaries for extensive probabilistic modeling and the practical feasibility of real-time Bayesian updating in dynamic systems.

