- The paper introduces the SHARP pipeline that integrates network proximity with the novel pAGE metric to identify drugs affecting aging hallmarks.
- It leverages a library of 2,358 aging-associated genes mapped onto the human interactome to define hallmark modules and assess drug interactions.
- Validation against clinical candidates and murine lifespan models underscores the potential of network-driven repurposing in precision geroscience.
Network-driven Discovery of Repurposable Drugs Targeting Hallmarks of Aging
Introduction
The study titled "Network-driven discovery of repurposable drugs targeting hallmarks of aging" addresses a significant challenge in aging research: the development of therapeutic interventions targeting the multifactorial nature of aging. By integrating network medicine with genomic data, the study seeks to identify clinically approved or experimental drugs that can modulate the interconnected hallmarks of aging. Employing a network-based approach, the authors utilize a comprehensive library of 2,358 longevity-associated genes mapped onto the human interactome to identify distinct hallmark modules of aging. Through computational analysis, this framework measures the network proximity of over 6,000 compounds to these hallmark modules.
Hallmark Modules and Network Proximity
The research begins by identifying hallmark-associated genes that form significant connected subgraphs, known as hallmark modules, in the human interactome (Figure 1). Network proximity is then utilized to assess the closeness of drug targets to these hallmark modules, with a specific focus on drugs that can perturb aging processes beneficially.
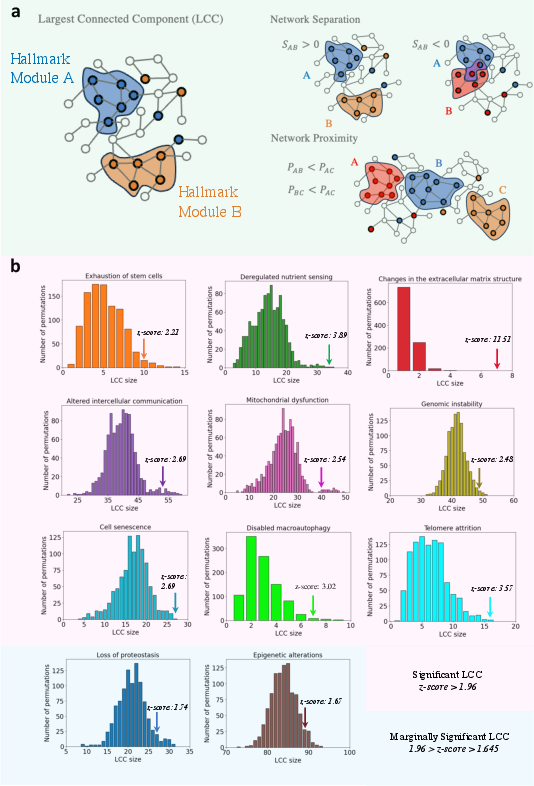
Figure 1: Network characteristics of the hallmarks of aging. Genes associated with similar biological mechanisms often form connected components.
The hallmark modules reveal the underlying network organization of aging genes, providing a structured framework to evaluate interactions between drugs and aging phenotypes. By employing a novel transcription-based metric termed pAGE, the analysis distinguishes whether drug-induced expression changes align or counteract age-related transcriptional shifts.
Pro-Age Metric: Distinguishing Beneficial Interventions
The introduction of the pAGE metric marks a methodological advancement, offering insight into the directionality of drug effects on aging processes (Figure 2). This quantitative measure evaluates whether the expression changes induced by drugs enforce or neutralize known age-related changes, facilitating the identification of pro-longevity compounds.
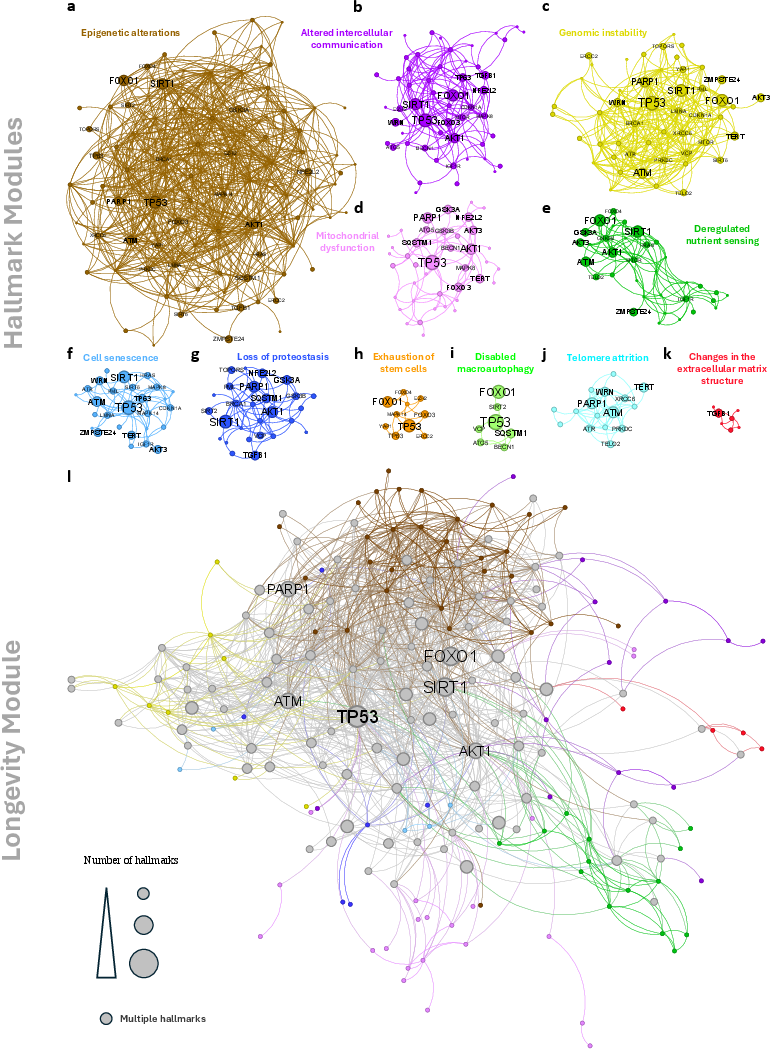
Figure 2: Perturbation pro-Age signature. The network proximity identifies drugs with close targets to aging hallmarks, while pAGE predicts the directionality of impact.
The integration of pAGE with network proximity helps to systematically filter and identify drugs that have the potential to reverse hallmark-associated expression changes, thereby promoting longevity.
Validation and Application of the SHARP Pipeline
The Systematic Hallmark-based Aging Repurposing Pipeline (SHARP) is validated against known drug candidates in clinical trials and those shown to extend lifespan in murine models. The pipeline effectively captures and predicts the therapeutic potential of these drugs (Figure 3).
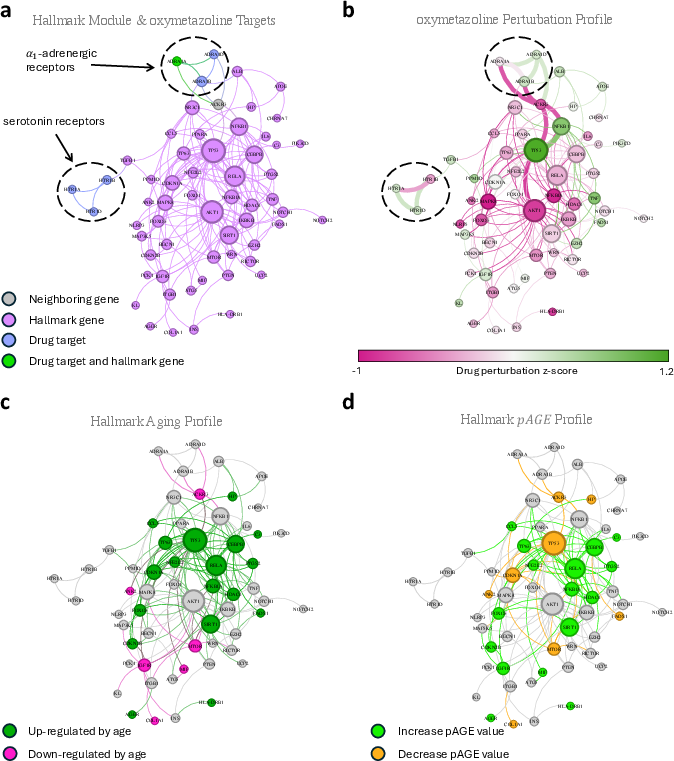
Figure 3: Proximity and pAGE predicts mechanism of action. Example with oxymetazoline illustrating the drug's network-based pathway and predicted impact on longevity.
SHARP identifies 370 drugs with significant proximity to at least one hallmark of aging, with 60 providing pAGE data. This comprehensive approach facilitates the recognition of leading candidates for repurposing, offering both exploratory and predictive capabilities in evaluating drug potential for combating aging.
Discussion and Future Directions
This research advances the field of aging interventions by illustrating the potential of a network-centric approach to drug repurposing. By leveraging comprehensive genetic and network data, the study offers a framework to efficiently filter existing drugs for their ability to modulate aging hallmarks.
While primarily computational, the study lays the groundwork for experimental validation and potential clinical application. The differentiation between pro-longevity and age-accelerating compounds enhances the strategy's robustness in identifying viable therapeutic targets. Future research directions include integrating tissue-specific data, exploring combination therapies, and refining pAGE by accounting for dosage and non-linear effects. This approach exemplifies a crucial step towards Precision Geroscience, emphasizing personalized strategies in aging therapeutics.
Conclusion
The integration of network medicine with genomic data presents a practical method for identifying drug candidates capable of modulating aging's multifactorial mechanisms. This approach not only fosters the understanding of aging processes but also streamlines the path toward developing clinically viable interventions. By advancing the application of network-based repurposing and the pAGE metric, this research underscores the potential for revitalizing anti-aging drug discovery with computational ingenuity.




