How many patients could we save with LLM priors?
Abstract: Imagine a world where clinical trials need far fewer patients to achieve the same statistical power, thanks to the knowledge encoded in LLMs. We present a novel framework for hierarchical Bayesian modeling of adverse events in multi-center clinical trials, leveraging LLM-informed prior distributions. Unlike data augmentation approaches that generate synthetic data points, our methodology directly obtains parametric priors from the model. Our approach systematically elicits informative priors for hyperparameters in hierarchical Bayesian models using a pre-trained LLM, enabling the incorporation of external clinical expertise directly into Bayesian safety modeling. Through comprehensive temperature sensitivity analysis and rigorous cross-validation on real-world clinical trial data, we demonstrate that LLM-derived priors consistently improve predictive performance compared to traditional meta-analytical approaches. This methodology paves the way for more efficient and expert-informed clinical trial design, enabling substantial reductions in the number of patients required to achieve robust safety assessment and with the potential to transform drug safety monitoring and regulatory decision making.
Paper Prompts
Sign up for free to create and run prompts on this paper using GPT-5.
Top Community Prompts
Explain it Like I'm 14
What Is This Paper About?
This paper asks a simple but powerful question: can we use knowledge from large AI models to run safer, faster clinical trials with fewer patients? The authors show a way to plug “expert knowledge” from a LLM into a statistical model of side effects (also called adverse events) in multi-center clinical trials. By doing this, the model makes better predictions, so researchers may not need as many patients to reach solid, trustworthy conclusions.
What Questions Did the Researchers Want to Answer?
They focused on three easy-to-understand goals:
- Can AI help set smarter “starting beliefs” (priors) for a Bayesian model that predicts side effects in clinical trials?
- Do these AI-informed priors make predictions more accurate than the current best standard?
- If yes, how much can this cut down the number of patients needed while keeping accuracy strong?
How Did They Do It? (Methods in Plain Language)
Think of a clinical trial like many schools (sites), each with a different number of students (patients). We want to predict how many “bad events” (side effects) happen per school and overall. Some schools have more events, some fewer. A “hierarchical” model handles this two-level structure: students inside schools.
- Hierarchical Bayesian model: This is a way of starting with a reasonable guess (a prior), then updating it with real data. It lets information “share” across sites so tiny sites aren’t too noisy.
- Poisson–Gamma setup:
- Poisson is a standard way to model counts (like “how many side effects happened”).
- Gamma describes how different sites’ average rates might vary.
- Priors (starting beliefs): Normally, statisticians pick them using past studies or simple rules. Here, the twist is they asked an LLM to suggest the prior settings. That’s like asking a very well-read assistant, trained on lots of medical texts, “What are reasonable values experts might choose to start with?”
What the LLM actually did:
- The team asked two AI models (Llama 3.3 and MedGemma) to provide two numbers that control how similar or different the sites’ side-effect rates should be. These numbers act like knobs that influence how much the model expects:
- Sites to be similar versus varied.
- Side-effect rates to be tightly clustered or spread out.
- They tried two prompting styles:
- Blind: general clinical-trial expertise without naming the disease.
- Disease-informed: adding lung cancer context (NSCLC).
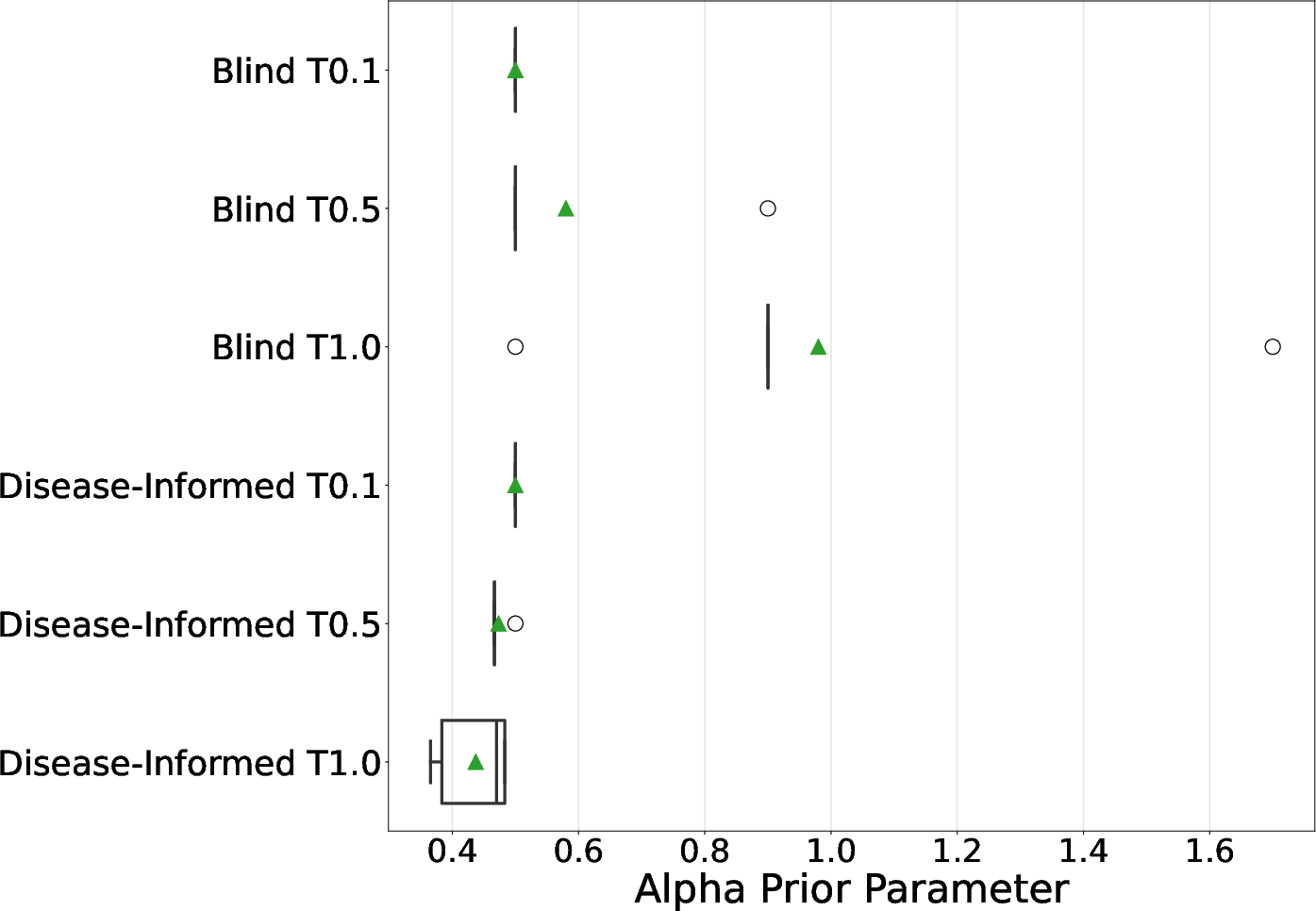
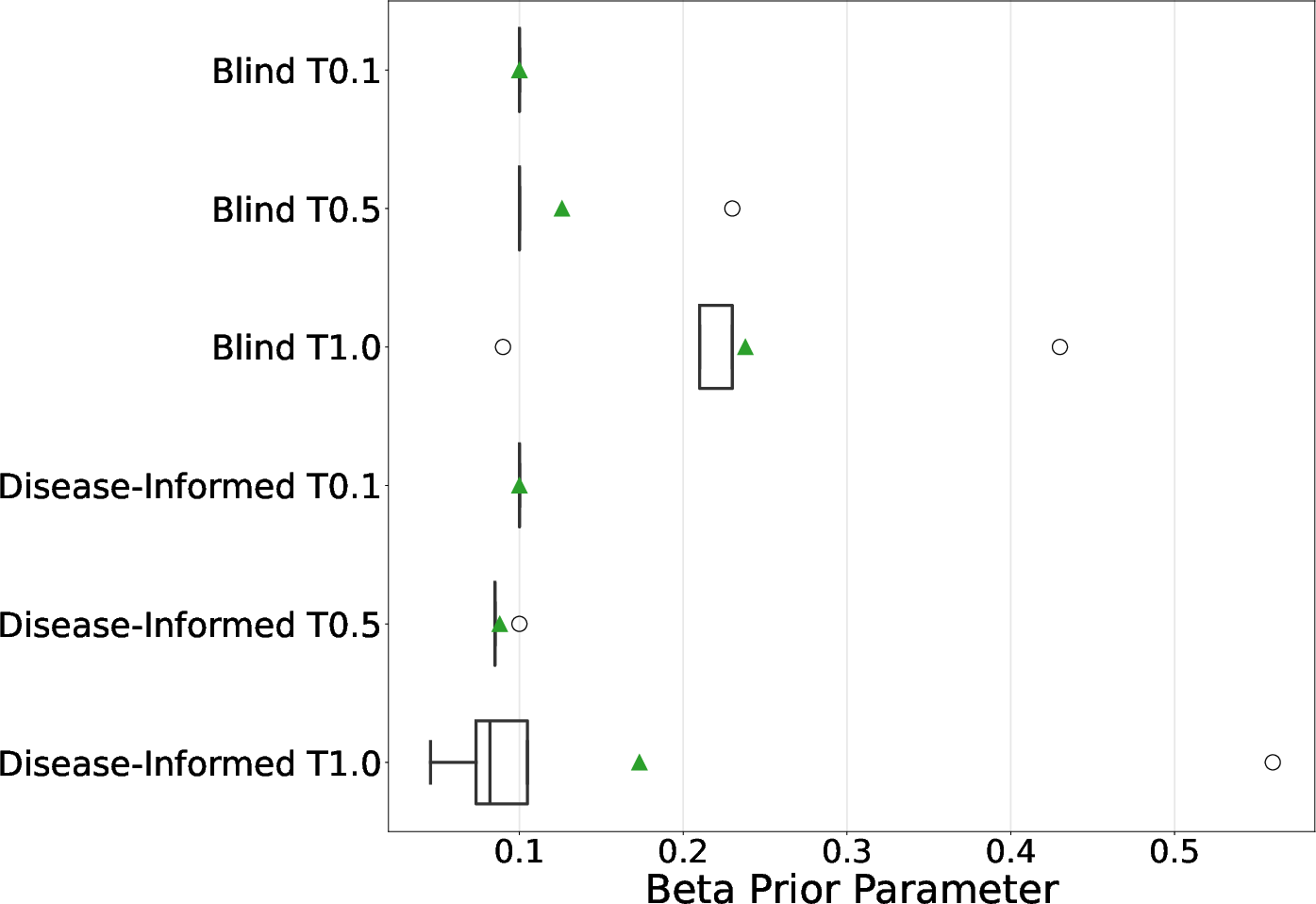
- They also adjusted the AI’s “temperature,” a creativity knob:
- Lower temperature = more consistent answers
- Higher temperature = more diverse answers
How they checked if this worked:
- Real data: 468 patients across 125 sites from a lung cancer trial.
- Cross-validation: They trained on some sites and tested on totally different sites (like practicing on some schools and seeing how well you predict at new schools).
- Score used: Log Predictive Density (LPD). Think of it as a prediction score—higher (less negative) is better.
- They also tested how performance changes if you reduce the amount of training data to see if the AI priors help you “do more with less.”
What Did They Find, and Why Does It Matter?
Here are the main results in simple terms:
- AI-informed priors beat the standard method: Using priors suggested by the LLMs made predictions better than the usual “meta-analytical” priors (a respected baseline from earlier studies).
- Best setup: Llama 3.3 with the blind prompt at higher temperature (T=1.0) performed best in predicting side effects.
- Fewer patients needed: With LLM-based priors, using only about 80% of the training data matched or beat the baseline that used 100%. In this study, that’s roughly 66 fewer patients to get similar accuracy.
- Disease-specific hints not required: Surprisingly, just the LLM’s general medical knowledge was enough. Adding detailed disease context didn’t consistently help.
- Temperature matters: Letting the LLM be a bit more “creative” (higher temperature) sometimes led to better priors and better predictions.
Why this matters:
- Clinical trials are expensive, slow, and involve real people. If you can get the same trustworthy results with fewer participants, you save time, money, and reduce the number of patients exposed to potential side effects—while still protecting safety and accuracy.
What Could This Change in the Real World?
This approach could:
- Help design smaller, faster trials that still meet high scientific standards.
- Improve how we monitor drug safety by combining statistical rigor with expert-like knowledge from AI.
- Support better decisions by regulators and doctors, because the method is transparent (unlike making up fake data). The AI suggests priors; it doesn’t invent new patients.
Limitations and Next Steps
The study used one disease area (lung cancer) and two AI models. Before this becomes standard:
- It should be tested across more diseases and more trials.
- We need to confirm that regulators accept AI-informed priors.
- Researchers should explore combining this with other AI tools (like synthetic data) carefully and transparently.
In short: Using AI to set smart starting points for statistical models can make clinical trials more efficient. This could mean fewer patients, faster answers, and safer, more affordable progress in medicine.
Knowledge Gaps
Knowledge Gaps, Limitations, and Open Questions
The following bullet points identify what remains missing, uncertain, or unexplored in the paper, phrased concretely to guide future research:
- External validity: Results are demonstrated on a single NSCLC control-arm dataset; it is unknown whether LLM-elicited priors generalize across diseases, therapeutic areas, trial phases, geographies, or sponsor/site ecosystems.
- Model family limitations: The Poisson–Gamma framework assumes equidispersion and no zero inflation; no comparisons are made to alternative likelihoods (e.g., Negative Binomial, Poisson–Lognormal, zero-inflated/recurrent-event models) that may better fit AE counts with heavy tails and excess zeros.
- Missing exposure adjustment: AE counts are modeled without offsets for patient exposure time/follow-up duration; the impact of including exposure offsets remains unexplored.
- No covariates: Patient-level (e.g., age, ECOG, comorbidities) and site-level covariates (e.g., region, site volume) are absent; how LLM priors interact with hierarchical models including covariates is unknown.
- Single-endpoint scope: Only total AE counts are modeled; extensions to multiple AE categories, severity grades, relatedness, SAEs, or time-to-first/recurrent AE processes are not examined.
- Control-arm only: The work does not model treatment arms or treatment effects; it is unclear whether LLM priors improve safety signal detection or treatment effect estimation in comparative settings.
- Prior family rigidity: Hyperpriors are restricted to independent Exponential distributions for alpha and beta; sensitivity to hyperprior families (e.g., Gamma, Lognormal, half-t, hierarchical shrinkage, or heavy-tailed priors) and to dependence between alpha and beta is not assessed.
- Prior-data conflict: No formal checks (e.g., prior predictive checks, conflict diagnostics, KLD-based measures) are reported to detect and mitigate prior–data discordance.
- Effective sample size (ESS): Claims of increased “effective sample size” are not quantified using established ESS definitions (e.g., Morita/Neuenschwander methods); a formal ESS calculation is missing.
- Power/type I error: The link between improved LPD and operating characteristics (power, type I error, false safety signal rates) is not established via simulation or analytical derivations.
- “Patients saved” quantification: The paper infers fewer patients are needed from small LPD gains; a formal prospective sample size/power analysis under informative priors is not provided.
- Calibration and coverage: No evaluation of posterior calibration (e.g., coverage of predictive/credible intervals), Brier scores, or decision-relevant calibration metrics is reported.
- Prior robustness safeguards: No guardrails are proposed for mis-specified or overconfident priors (e.g., robustification, mixture/hedging priors, prior tempering, or prior truncation).
- Temperature tuning methodology: Temperature is varied heuristically; criteria or procedures to choose temperature that optimize predictive performance while controlling prior informativeness are not developed.
- LLM uncertainty representation: The approach averages 5 elicited values; it does not elicit distributions or uncertainty intervals from the LLM nor model uncertainty via mixture priors over hyperparameters.
- Prompt design scope: Only “blind” vs. “disease-informed” prompts are tested; richer strategies (e.g., retrieval-augmented prompting with literature grounding, citation requirements, chain-of-thought verification) and their effect on prior quality are unexplored.
- Potential contamination: The possibility that LLMs’ training data include the evaluated dataset or closely related sources is not investigated; safeguards against data leakage/memorization are absent.
- Human expert baseline: There is no comparison to priors elicited from domain experts using standard structured elicitation protocols to establish whether LLM priors match or surpass human-informed priors.
- Reproducibility/versioning: LLM drift (model updates) and its impact on prior reproducibility are not addressed; procedures for version control, provenance, and audit trails of elicited priors are missing.
- Evaluation metric narrowness: Only LPD is reported; alternative metrics (e.g., WAIC/LOO, RMSE of site-level rates, calibration, decision-centric utilities) and statistical significance of observed differences are not analyzed.
- Practical clinical utility: The translation from improved LPD to concrete regulatory or pharmacovigilance decisions (e.g., AE signal thresholds, stopping rules) is not demonstrated.
- MCMC diagnostics: Convergence and mixing diagnostics (e.g., R-hat, ESS of chains, divergence rates) and sensitivity to sampler settings are not reported; potential dependence of inference stability on prior informativeness is unknown.
- Prior influence quantification: The magnitude of prior influence on the posterior (e.g., via prior-to-posterior KLD or fraction of information) is not quantified across conditions.
- Site heterogeneity structure: The assumption that site heterogeneity is fully captured by a single Gamma layer is untested; alternative multi-level or nonparametric structures (e.g., Dirichlet process mixtures) are not explored.
- Mathematical clarity of evaluation: The LPD computation integrates over a “new site” distribution; whether this aligns with the intended generalization target and whether alternative site-level predictive setups change conclusions is not examined.
- Cost–benefit analysis: No assessment of the operational cost of LLM elicitation (API use, governance overhead) versus projected trial size reduction is provided.
- Regulatory alignment: The paper does not outline how LLM-derived priors fit within GxP/ICH E9(R1)/21 CFR Part 11 expectations, nor how to document, validate, and justify such priors to regulators.
- Ethical risk assessment: The ethical implications of deploying LLM-driven priors in high-stakes trials (e.g., risk of biased priors disadvantaging subpopulations) are not assessed.
- Robust aggregation: Arithmetic averaging of elicited parameters may be inappropriate on the parameter scale; mixture-of-priors or median-of-means robustness to outliers in LLM outputs is not investigated.
- Sensitivity to outliers: AE counts include extreme values (up to 140); robustness to influential observations and heavy tails (e.g., via robust likelihoods or priors) is not studied.
- Prospective deployment: The framework is not tested in a prospective or adaptive design (e.g., interim monitoring with prior updates), leaving questions about stability under real-time use.
- Transparency of LLM rationale: The LLM is not required to provide citations or quantitative justifications for the elicited values; methods to audit and validate the evidence base are missing.
- Breadth of LLMs tested: Only two LLMs are evaluated; comparative performance across diverse general-purpose and biomedical models (and open vs. closed models) remains unknown.
Practical Applications
Immediate Applications
Below are actionable uses that can be deployed with today’s tools and practices, drawing directly from the paper’s findings (LLM-elicited priors outperforming meta-analytic baselines and enabling ~20% training data reductions for adverse-event modeling in multi-center trials).
- LLM-informed prior elicitation for Bayesian safety models in ongoing and near-term trials
- Sector: Healthcare/Pharma (biostatistics, clinical development), CROs
- What to do: Replace default/meta-analytic hyperpriors with LLM-elicited priors for hierarchical Poisson–Gamma AE models; operationalize the paper’s blind prompting + temperature sweep (e.g., T ∈ {0.1, 0.5, 1.0}, 5 queries/condition, mean aggregation) and select priors via prior predictive checks and cross-validation.
- Tools/Workflow: Stan/PyMC models + a small “LLM-to-prior” Python module that:
- Calls
Llama 3.3or domain LLM (e.g.,MedGemma) with the provided JSON schema prompt. - Automates temperature sensitivity runs and aggregates results.
- Logs prompts, seeds, model/version, and outputs for auditability.
- Assumptions/Dependencies: Hierarchical count model is appropriate for the endpoint; access to a capable LLM; governance for model-version locking; prior predictive checks included; reproducibility (prompt, temperature, seeds) is pre-specified in the SAP.
- Sample-size and power calculators that incorporate LLM-elicited priors at design time
- Sector: Healthcare/Pharma; CROs; Biostatistics consultancies
- What to do: Use LLM-derived priors to run Bayesian trial simulations for AE endpoints to estimate power and expected effective sample size; evaluate reductions similar to the paper’s 20% training data savings (context-dependent).
- Tools/Workflow: Simulation notebooks + CLI that:
- Retrieves LLM priors via the JSON API.
- Runs prior predictive checks and operating characteristic simulations (e.g., LPD, false signal rates).
- Exports design scenarios for protocol/SAP inclusion.
- Assumptions/Dependencies: Regulatory acceptance of prior-informed designs; simulation assumptions match clinical context; transparent documentation of elicitation and validation.
- Pre-specified, auditable prior-elicitation documentation for regulatory submissions
- Sector: Healthcare/Pharma; Regulatory Affairs; Quality; Policy
- What to do: Add a “LLM prior elicitation” appendix to the SAP and CSR:
- Include full prompts, model versions, temperature grid, seeds, and outputs.
- Show cross-validated performance and prior predictive checks vs meta-analytic baseline.
- Tools/Workflow: Version-controlled prompt registry; automated report generation (knitr/Quarto/Jupyter) embedding code, outputs, and diagnostics.
- Assumptions/Dependencies: Agency comfort with explicit, parametric priors vs synthetic data augmentation; internal SOPs for LLM use and validation; privacy/security review.
- Data Monitoring Committee dashboards using LLM priors for ongoing safety assessment
- Sector: Healthcare/Pharma; CROs; Clinical Operations
- What to do: Use the hierarchical model with LLM priors for interim AE monitoring and site-level information borrowing; provide posterior predictive distributions for new sites.
- Tools/Workflow: Shiny/Streamlit dashboard wired to posterior updates; alert thresholds for abnormal site AE rates; reproducible prior artifacts frozen before trial start.
- Assumptions/Dependencies: Pre-specification in protocol; controlled access to trial data; careful governance to avoid “drift” in priors mid-trial.
- Multi-center hospital quality and safety analytics (non-trial)
- Sector: Healthcare providers; Health systems; Public health
- What to do: Apply the Poisson–Gamma framework with LLM-elicited hyperpriors to model count outcomes (e.g., falls, infections) across hospitals/wards, improving stability for small units.
- Tools/Workflow: Monthly pipeline that re-fits models and flags outliers; simple JSON-prior interface for analysts to regenerate priors annually or per program.
- Assumptions/Dependencies: Availability of reliable event counts; alignment between clinical context and LLM general clinical knowledge (blind prompts sufficed in the paper); change management with quality teams.
- Teaching and replication kits for Bayesian workflows with LLM priors
- Sector: Academia; Biostatistics education; Methodology labs
- What to do: Course modules and lab assignments reproducing the paper’s cross-validation and sample-efficiency experiments; extend to other datasets.
- Tools/Workflow: Open-source notebooks; a minimal “llm_priors” package; datasets from Project Data Sphere or similar.
- Assumptions/Dependencies: Access to permissible LLM endpoints or local models; institutional policies for LLM use.
- Lightweight productization inside CROs/biostats teams (“LLM-to-Prior” microservice)
- Sector: CROs; Pharma statistics groups; Software vendors
- What to do: Wrap the elicitation protocol in a service offering that delivers:
- Prior candidates + diagnostics + documentation pack.
- SAP-ready language and reproducibility bundles.
- Tools/Workflow: Containerized API; CI/CD to pin LLM versions; unit tests for JSON parsing, prior predictive checks.
- Assumptions/Dependencies: Vendor risk management; contract language on acceptable use of generative AI; cost control for LLM calls.
- Ethics and IRB templates for disclosing LLM use in statistical priors
- Sector: Policy; IRBs; Research governance
- What to do: Template language describing how LLMs inform hyperpriors (not data), how uncertainty is handled, and how the approach reduces participant exposure.
- Tools/Workflow: Template repository and checklists; standardized sensitivity-analysis sections.
- Assumptions/Dependencies: Institutional willingness to adopt; clarity that priors are transparent and stress-tested (not hallucinated facts).
Long-Term Applications
Below are applications that likely need further validation across endpoints, diseases, models, or require regulatory standardization and product engineering.
- Extension beyond count data to other endpoints and models
- Sector: Healthcare/Pharma; Methodology
- What: LLM-elicited priors for binary (logistic), ordinal, time-to-event (survival), and continuous outcomes; mixture priors or non-conjugate priors with prior predictive calibration.
- Dependencies: New prompts eliciting appropriate hyperparameters; robust prior informativeness controls; extensive simulation and external validation.
- Regulatory standards and guidance for LLM-informed priors
- Sector: Policy; Regulators; Standards bodies
- What: Methodological guidance for:
- Pre-registration of prompts/temperatures/models.
- Documentation/auditing requirements.
- Acceptable sensitivity analyses and prior predictive checks.
- Dependencies: Multi-stakeholder pilots; consensus on governance and acceptable use; clarity on change control and version locking.
- Indication-specific prior registries and “prior marketplaces”
- Sector: Healthcare/Pharma; Knowledge platforms; Software
- What: Curated, peer-reviewed libraries of elicited priors by indication, phase, and endpoint (with provenance and performance metrics).
- Dependencies: Community curation; legal/IP concerns; updating cadence as literature evolves.
- Combination with synthetic data augmentation for small-sample trials
- Sector: Healthcare/Pharma; Methodology; Software
- What: Joint pipelines that use LLM priors plus validated synthetic data to further improve predictive performance and robustness.
- Dependencies: Risk controls for leakage/bias; formal criteria for when augmentation helps vs harms; regulatory acceptance.
- Adaptive and platform trials with improved inter-arm/site borrowing
- Sector: Healthcare/Pharma; Trial design
- What: Use LLM priors to stabilize hierarchical borrowing across sites/arms/baskets in platform or basket trials, potentially accelerating decisions with fewer participants.
- Dependencies: Complex design simulation; operating characteristics across many scenarios; prespecified borrowing rules and safeguards.
- Post-marketing pharmacovigilance and signal detection
- Sector: Pharma; Regulators; Public health
- What: Hierarchical models (sites=regions/hospitals) with LLM priors for AE signal detection in observational settings; faster escalation on true signals.
- Dependencies: Case definition harmonization; observational confounding; calibration against known signals.
- Cross-industry transfer of the elicitation workflow to other count processes
- Sector mapping and examples:
- Insurance/Finance: Claims frequency modeling across regions/branches.
- Manufacturing/Robotics: Defect or failure counts across lines/robots.
- Software/IT Ops/Cybersecurity: Incident/bug rates across services/regions.
- Energy/Utilities: Outage events across substations/crews.
- What: Adopt LLM-informed hierarchical priors to stabilize sparse units and improve forecasts with fewer observations.
- Dependencies: Domain-appropriate prompts; validation against historical baselines; governance for AI use in risk-sensitive sectors.
- Privacy-preserving, on-prem LLM deployments for prior elicitation
- Sector: Large pharma; Hospitals; Highly regulated industries
- What: Run vetted biomedical LLMs behind the firewall to satisfy data governance while retaining elicitation benefits.
- Dependencies: Model hosting, cost and MLOps; internal evaluation of model fitness and drift.
- Governance and bias-auditing frameworks for LLM priors
- Sector: Policy; QA; Risk management
- What: Tooling to quantify prior informativeness, detect instability across temperatures/prompts, and surface potential publication-bias artifacts encoded in LLMs.
- Dependencies: Benchmarks spanning diseases/endpoints; standardized diagnostics and acceptance thresholds.
- Semi-autonomous “AI trial statistician” assistants
- Sector: Software; CROs; Pharma stats
- What: Agents that propose candidate models, elicit priors, run simulations, compile SAP-ready documentation, and surface sensitivity/risk analyses.
- Dependencies: Guardrails; human-in-the-loop review; integration with probabilistic programming and eTMF/CTMS systems.
Notes on Feasibility and Key Assumptions (cross-cutting)
- External validity: The paper demonstrates gains on one NSCLC control-arm dataset; generalization to other diseases/endpoints requires replication.
- Model fit: Benefits hinge on appropriate hierarchical structure (e.g., Poisson–Gamma for counts). Other outcome types need tailored elicitation.
- LLM quality and versioning: Elicited priors depend on the model used, prompt, temperature, and seeds; all must be locked and logged for reproducibility and audit.
- Bias and provenance: LLMs encode patterns from literature (including publication bias). Prior predictive checks, cross-validation, and sensitivity analyses are essential.
- Regulatory acceptance: Transparent, parametric priors are more interpretable than synthetic data augmentation, but require clear documentation and pre-specification.
- Operational readiness: Teams need lightweight tooling (JSON I/O, diagnostics, report generation) and SOPs to ensure consistent, auditable use.
- Security and privacy: Even though prompts need not include PHI, enterprises may require on-prem deployment and strict access controls for LLM use.
Collections
Sign up for free to add this paper to one or more collections.