An AI system to help scientists write expert-level empirical software
Abstract: The cycle of scientific discovery is frequently bottlenecked by the slow, manual creation of software to support computational experiments. To address this, we present an AI system that creates expert-level scientific software whose goal is to maximize a quality metric. The system uses a LLM and Tree Search (TS) to systematically improve the quality metric and intelligently navigate the large space of possible solutions. The system achieves expert-level results when it explores and integrates complex research ideas from external sources. The effectiveness of tree search is demonstrated across a wide range of benchmarks. In bioinformatics, it discovered 40 novel methods for single-cell data analysis that outperformed the top human-developed methods on a public leaderboard. In epidemiology, it generated 14 models that outperformed the CDC ensemble and all other individual models for forecasting COVID-19 hospitalizations. Our method also produced state-of-the-art software for geospatial analysis, neural activity prediction in zebrafish, time series forecasting and numerical solution of integrals. By devising and implementing novel solutions to diverse tasks, the system represents a significant step towards accelerating scientific progress.
Paper Prompts
Sign up for free to create and run prompts on this paper using GPT-5.
Top Community Prompts
Explain it Like I'm 14
Overview
This paper introduces an AI system that helps scientists write high‑quality computer programs for research. These programs are designed to get the best possible “score” on a task, like predicting future COVID‑19 hospitalizations or combining biology datasets so they’re easier to compare. The AI uses two main tools: a powerful LLM (a kind of AI that writes and edits code) and a smart search strategy that tries many ideas and keeps improving them. The goal is to speed up scientific discovery by letting AI do a lot of the slow, careful coding work.
What problems is the paper trying to solve?
Scientists often need “empirical software”—programs that are judged by a clear score (like accuracy or error). But:
- Writing this software by hand can take months or years.
- There’s no easy way to try all the different ideas that might work.
- Progress can stall because people don’t have time to explore enough options.
The paper asks: Can an AI system automatically write, test, and improve scientific code to reach expert‑level results across many fields?
How does the AI system work?
Think of the system like a team coach running drills to beat a high score in a game:
- Scorable tasks: The task has a clear way to measure success, like a leaderboard rank or an error number. That’s the “score.”
- Big AI writer (LLM): The AI reads the task and writes code to try solving it. It can also read research ideas from papers or search results and mix them into the code.
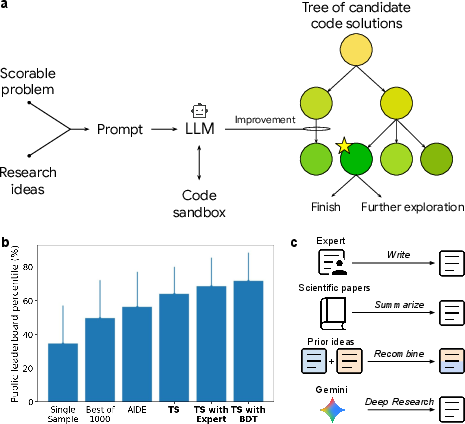
- Tree search: Imagine a branching tree of possible ideas. Each branch is a different version of the code. The AI: 1) writes a version, 2) runs it in a safe environment (“sandbox”), 3) gets the score, 4) keeps the best versions, 5) explores promising branches further (like finding the best path in a maze).
- Idea recombination: The AI can combine parts of different methods—like taking the “best defense” from one team and the “best offense” from another—to make a stronger hybrid.
In simple terms: the AI repeatedly writes code, tests it, learns what works, and makes smarter versions—guided by the score.
What did they test it on, and what did they find?
The researchers tried the system on a variety of real scientific problems. Here are the highlights, and why they matter:
- Kaggle mini‑competitions (practice ground)
- What: 16 simple prediction contests used to compare performance against thousands of humans.
- Result: The AI with tree search clearly beat single attempts and even many repeated attempts without search.
- Why it matters: Shows the method can reliably improve code by exploring and refining ideas.
- Single‑cell biology (combining datasets from different labs)
- What: Remove “batch effects” (technical differences) while keeping real biology signals.
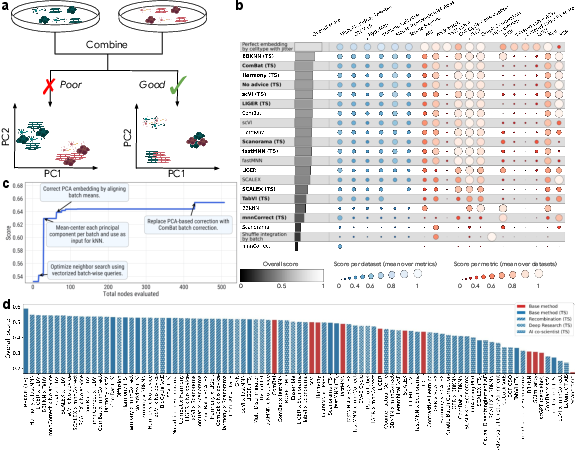
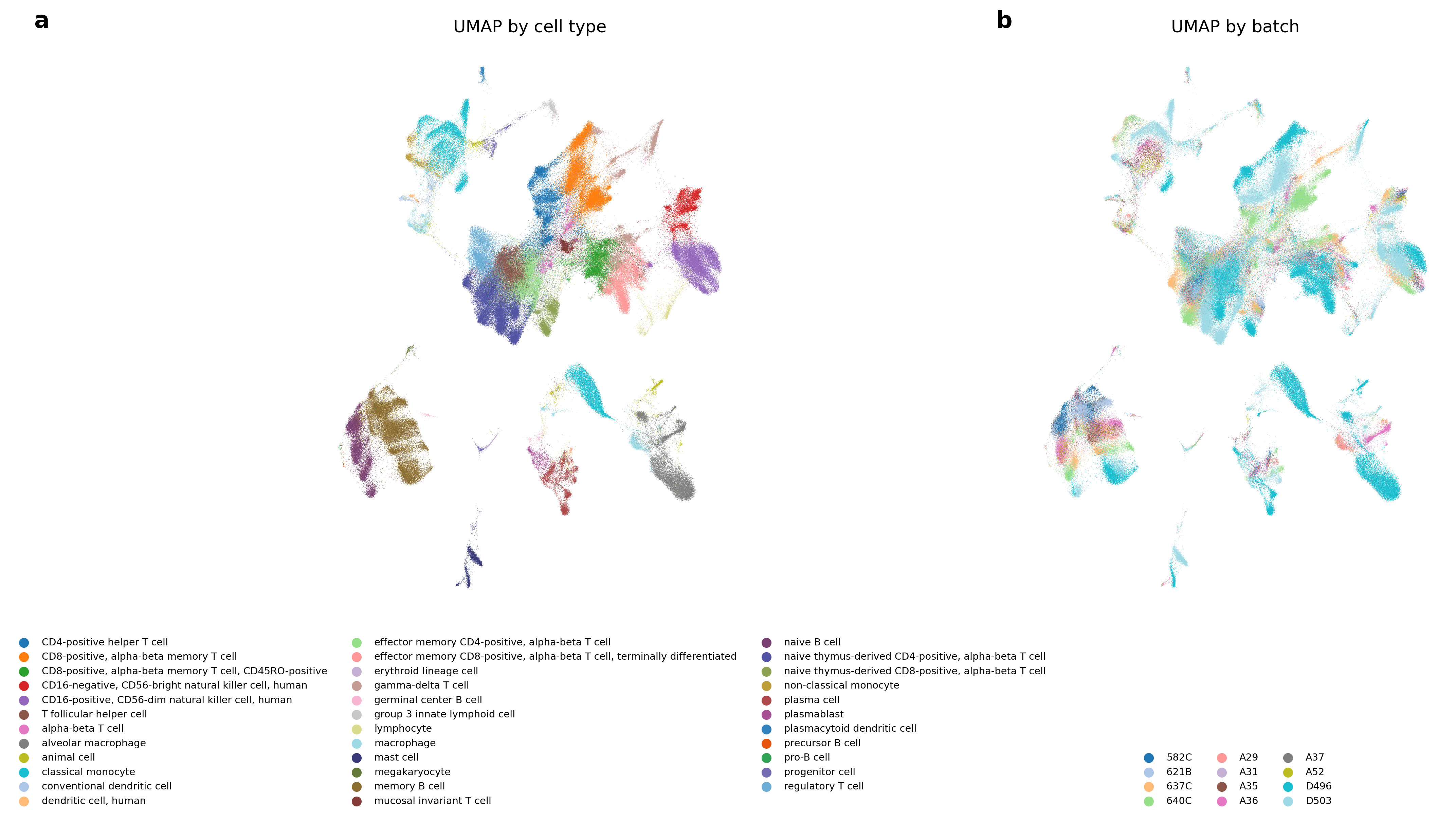
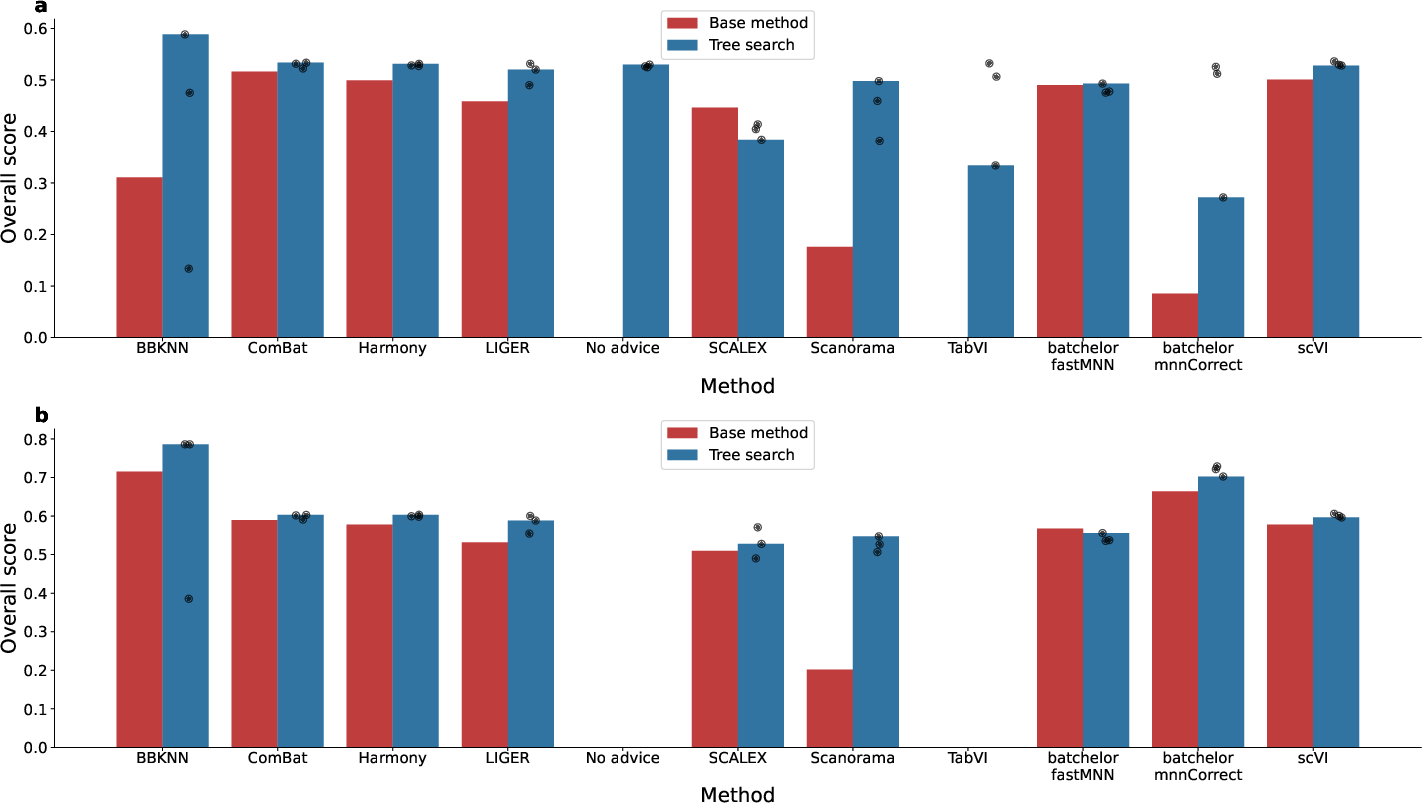
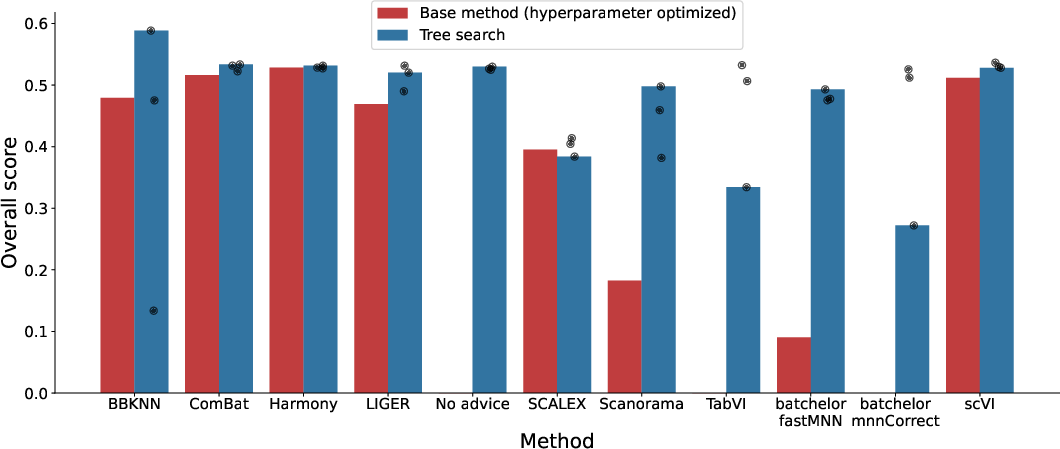
- Result: The AI replicated and improved on many published methods. It often beat the leaderboard methods, and in several cases created better hybrids by combining techniques. One top result improved a leading method by about 14% on the overall score.
- Why it matters: Better data integration helps build massive cell atlases and speeds up discoveries in health and disease.
- COVID‑19 hospitalization forecasts (CDC Forecast Hub setting)
- What: Predict new hospitalizations across U.S. states weeks ahead, with honest uncertainty.
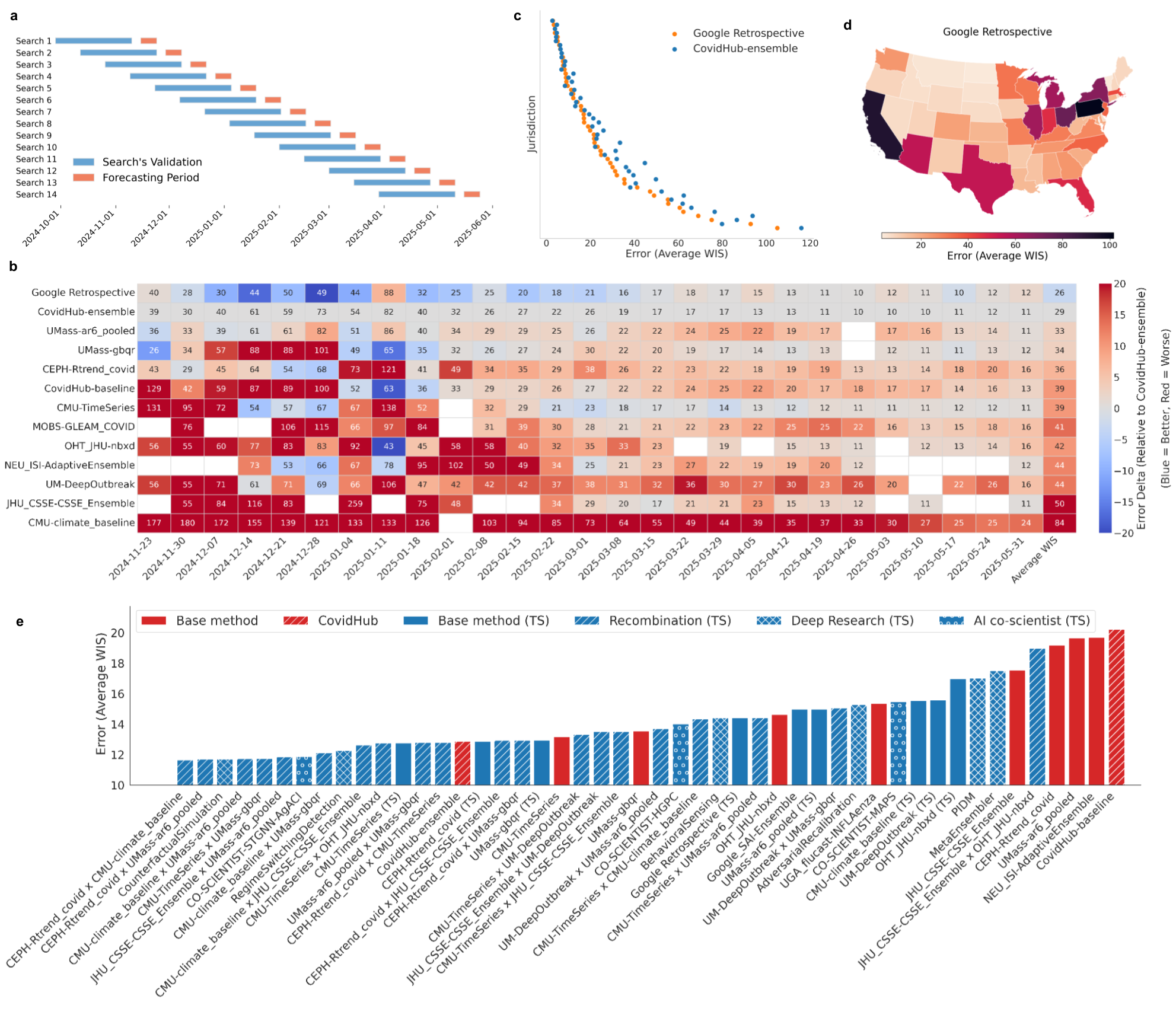
- Result: The AI produced 14 different models that outperformed the CDC’s ensemble during the test period. Many winners were hybrids combining stable trend models with flexible machine learning or epidemiological ideas.
- Why it matters: Stronger forecasts can help hospitals and policymakers prepare resources and respond to outbreaks.
- Satellite image segmentation (mapping land cover pixel by pixel)
- What: Label each pixel (e.g., building, road, water) in challenging images.
- Result: The top AI‑generated models reached mean IoU around 0.80–0.82, beating recent academic methods reported for the same dataset.
- Why it matters: More accurate maps can support environmental monitoring and disaster response.
- Whole‑brain neural activity in zebrafish (predicting future brain signals)
- What: Forecast the activity of 70,000+ neurons from short past windows.
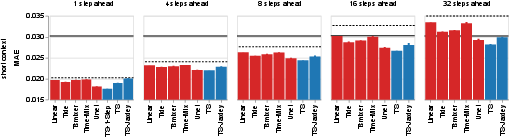
- Result: The AI built fast time‑series models that outperformed most baselines and even beat a heavy video model in most settings, training in hours instead of days. A special 1‑step model led at very short horizons.
- Why it matters: Fast, accurate brain forecasting can help scientists study how brains process information and behave.
- General time‑series forecasting (GIFT‑Eval benchmark)
- What: Predict many kinds of time series (from seconds to years) across many domains.
- Result: The system found strong per‑dataset solutions and also built a single general‑purpose forecasting library that adapts to different datasets and competes with top entries, including foundation models.
- Why it matters: Useful for finance, climate, energy, healthcare—anywhere you need good forecasts.
- Numerical integration (solving tough integrals)
- What: Improve on standard methods that sometimes fail or lose precision.
- Result: The paper introduces the setup and motivation; this section is incomplete in the provided text, but the goal is to create a more reliable, flexible integrals solver.
- Why it matters: Better integrators help in physics and engineering simulations.
Why is this important?
- Speed: The AI can try hundreds or thousands of ideas much faster than a person can code them, quickly climbing to expert‑level solutions.
- Breadth: It works across very different fields—biology, epidemiology, mapping, neuroscience, forecasting, and math.
- Creativity: By reading research and recombining ideas, it often invents new hybrids that perform better than any single method.
- Reliability: It doesn’t just tune parameters; it can implement, verify, and improve whole algorithms, even when there’s no public code.
What could this change in the future?
- Faster scientific progress: Scientists spend less time building and debugging complex code and more time on questions and insights.
- Better tools for everyone: High‑quality, domain‑specific software could become more accessible, even for smaller teams.
- New discoveries: Systematic idea exploration can uncover “needle‑in‑a‑haystack” solutions that people might miss.
- Teamwork between humans and AI: Scientists can guide the AI with goals and ideas, while the AI explores, tests, and refines—like having a tireless research assistant that can also read the literature.
In short, this AI system shows that automated, score‑driven code design—powered by smart search and research‑aware code writing—can match or beat expert methods across tough scientific tasks, helping science move faster and farther.
Knowledge Gaps
Below is a consolidated list of concrete knowledge gaps, limitations, and open questions left unresolved by the paper. Each item is phrased to guide future research toward actionable follow-ups.
- General/system-level: Lack of formal ablations
- No comprehensive ablation quantifying the marginal contributions of tree search vs. single-shot LLM, sampling depth, idea injection, external search, and recombination across the scientific benchmarks (only shown on Kaggle).
- No analysis of alternative search strategies (e.g., beam search, evolutionary strategies, MCTS variants, temperature/entropy schedules) and their sample-efficiency vs. performance trade-offs.
- General/system-level: Compute, efficiency, and carbon cost
- Missing reporting of total compute budgets (LLM calls, GPU/CPU time), cost per successful solution, and carbon footprint across domains, preventing fair efficiency comparison to expert pipelines.
- No scaling law characterization linking node budget, prompt length, and performance gains or diminishing returns.
- General/system-level: Reproducibility and stability
- Incomplete reporting of reproducibility across random seeds, runs, and environments for all domains (replicates shown sporadically).
- No evaluation of version drift (library updates) on the generated code’s correctness and performance, nor pinning/lockfile practices to ensure long-term reproducibility.
- General/system-level: Safety and sandboxing guarantees
- Insufficient details on sandbox/security model (allowed syscalls, network isolation, dependency whitelisting), and defenses against metric hacking, data leakage, or exfiltration during code execution.
- No assessment of dependency supply-chain risks (e.g., unvetted pip packages) or safeguards against license-incompatible code synthesis.
- General/system-level: External knowledge integration and provenance
- Unclear procedures to verify copyright compliance, provenance of incorporated ideas, and potential code verbatim copying from external sources (plagiarism and licensing risks).
- No mechanism to track, attribute, and audit which external papers/sections materially influenced a given solution (provenance trails).
- General/system-level: Novelty and scientific validity
- Claims of “novel methods” lack systematic novelty assessment (e.g., code similarity, idea lineage mapping) to distinguish recombination, reimplementation, and genuine innovation.
- Optimizing a single score risks Goodharting; the paper does not systematically check domain-specific scientific validity beyond the target metric.
- General/system-level: Multi-objective trade-offs
- No frameworks to control or analyze trade-offs among competing objectives (performance vs. interpretability, fairness, compute, stability, or domain constraints).
- General/system-level: Generalization and task selection bias
- Tasks were selected to align with authors’ agendas; unclear how representative these are of the broader landscape of “scorable” scientific tasks.
- No stress-tests on out-of-domain generalization, dataset shifts, or adversarial perturbations.
- General/system-level: Human-in-the-loop and governance
- No procedural guidance on when and how human experts should audit, constrain, or veto generated code, especially in safety-critical domains.
- No policy for credit/attribution when recombining previously published methods, nor governance for authorship and recognition.
- Kaggle benchmark
- Reliance on public leaderboards risks overfitting; no evaluation on private leaderboards or unseen holdouts to verify generalization.
- The effect of “expert advice” prompts on fairness is not analyzed; unclear whether this injects privileged knowledge that biases comparisons.
- scRNA-seq batch integration
- Biological validity beyond benchmark metrics is not assessed (e.g., preservation of gene-gene relationships, downstream differential expression, pathway enrichment, and rare-cell recovery).
- Generalization across tissue types, species, and novel labs not explicitly tested; robustness to extreme batch confounders is unknown.
- Scalability limits (cells × genes) and memory/runtime vs. state-of-the-art implementations are not reported.
- Metric computation failures are scored as zero; the bias this introduces and its impact on rankings are not analyzed.
- Limited statistical significance reporting across runs and datasets; uncertainty in performance deltas is not quantified.
- No assessment of potential metric gaming (e.g., transformations that inflate specific integration metrics while harming downstream biology).
- COVID-19 forecasting (CovidHub)
- Real-time constraints and data-revision/backfill handling are not addressed; unclear if only data available at forecast issuance time were used (risk of lookahead bias).
- Evaluation period (3 reference dates × 4 horizons × 52 jurisdictions) is limited; statistical significance and seasonality robustness are not established.
- Uncertainty calibration (beyond WIS) and coverage properties (e.g., empirical coverage of quantiles) are not reported.
- Replications that omitted external data underperform originals; comparisons are not normalized for data access parity.
- No live, prospective submissions to the Hub; operational feasibility and robustness under real-time data drift remain untested.
- Geospatial segmentation (DLRSD)
- Comparability to prior work is unclear because a custom 80/20 split was used; prior papers may have different splits or protocols.
- Dataset is small and homogeneous; no cross-dataset generalization (e.g., to larger, real-world remote sensing corpora) or domain-shift tests.
- Computational costs of heavy TTA and large encoders vs. marginal gains are not analyzed; no ablation on augmentation vs. architecture.
- Uncertainty quantification and calibration for pixel-wise predictions (important for downstream decisions) are not evaluated.
- Neuroscience (ZAPBench)
- Evaluation focuses on MAE; additional metrics (e.g., correlation, variance explained, region/stimulus-specific performance, causal response fidelity) are not reported.
- Interpretability and biological plausibility of the learned cross-neuron interactions are not analyzed; no alignment with forthcoming structural connectome.
- Long-horizon degradation behaviors and robustness across stimulus regimes are under-explored.
- Biophysical Jaxley-based model ignores explicit synaptic dynamics; open question: can hybrid models incorporate structural priors to outperform data-only models efficiently?
- Time-series forecasting (GIFT-Eval)
- Fairness of compute budgets vs. foundation models is not established; per-dataset bespoke search may confer an advantage.
- The unified solution’s adaptive presets (including holiday features for specific countries) may encode implicit domain knowledge; generality and portability are untested.
- Risk of validation-overfitting from per-dataset configuration selection is not quantified; no nested CV or external holdout validation across datasets.
- No error analysis on failure modes by domain/frequency (e.g., highly intermittent series, regime-switching, irregular sampling).
- Numerical integration (Gaussian quadratures)
- The section is incomplete; missing empirical results, integral classes addressed (oscillatory, singular, highly peaked), error bounds, and comparisons to QUADPACK/scipy integrate under adversarial cases.
- No theoretical guarantees or adaptive control strategies for precision, cancellation handling, and convergence detection.
- Search-space design and idea recombination
- No systematic methodology to bound or prioritize the combinatorial idea space; risk of myopic recombination without global novelty or theory-driven guidance.
- Automated recombination is evaluated empirically, but its mechanisms for avoiding incompatible or degenerate hybrids are not characterized.
- Metric hacking and evaluation harness integrity
- Limited discussion of anti-cheating mechanisms (e.g., guarding against code that reads holdout labels, tampers with metrics, or leaks private data) beyond general “sandbox” references.
- Licensing and IP constraints
- Open question: how to ensure generated code complies with licenses of recombined methods and avoids contaminating IP when using LLMs trained on mixed-license corpora.
- Deployment and maintenance
- Maintainability, readability, and test coverage of generated code are not assessed; no guidelines for transitioning solutions into production or for ongoing model/code monitoring.
- Societal/ethical implications
- No discussion of broader impacts, including displacement of expert labor, shifts in scientific credit, or guardrails for misuse in sensitive domains.
- Theoretical understanding
- No formal analysis of when tree search over LLM proposals is guaranteed (or likely) to find high-quality solutions, or conditions under which it fails (e.g., deceptive reward landscapes, sparse-reward settings).
Practical Applications
Immediate Applications
The paper introduces an LLM + Tree Search system that automatically generates and iteratively improves expert-level empirical software for “scorable tasks.” The following are practical, deployable use cases across sectors, along with expected tools/workflows and feasibility considerations.
- Bioinformatics: single-cell RNA-seq batch integration at atlas scale
- Use case: Integrate multi-lab scRNA-seq datasets (e.g., Human Cell Atlas) with improved batch correction (e.g., BBKNN(TS)+ComBat) that outperforms state-of-the-art on OpenProblems.
- Sector: Healthcare, biotech, academic genomics.
- Tools/products/workflows: Drop-in “AutoBatchIntegration” module for Scanpy/Seurat pipelines; containerized service that ingests AnnData and outputs integrated embeddings/graphs; CI for method recombination and leaderboard tracking.
- Assumptions/dependencies: Access to evaluation metrics and held-out datasets to avoid overfit; adherence to data governance; compute budget; expert sign-off for downstream biological interpretation; robustness across species/tissues/platforms.
- Public health: jurisdiction-level COVID-19 hospitalization forecasting
- Use case: Weekly forecasts with uncertainty outperforming CDC’s CovidHub ensemble; local resource planning (staffing, bed capacity, PPE); scenario analysis (e.g., variant emergence).
- Sector: Government/public health, hospital systems, health insurers.
- Tools/products/workflows: “EpiCast” forecasting service with rolling validation, WIS-based model selection, Monte Carlo scenario simulation, regime-switching detection; data connectors to CDC/CovidHub feeds; dashboard for jurisdictions.
- Assumptions/dependencies: Timely and stable reporting; data quality shifts; legal agreements for external data sources (mobility, wastewater); calibration monitoring; governance for policy use.
- Geospatial analysis: remote sensing semantic segmentation
- Use case: Land-use/land-cover mapping, deforestation monitoring, disaster damage assessment with mIoU > 0.80 on DLRSD using UNet++/SegFormer + TTA.
- Sector: Environmental monitoring, agriculture, insurance, logistics, public safety.
- Tools/products/workflows: “GeoSeg Studio” for fine-tuning pretrained encoders with strong augmentations and test-time augmentation; integration with GIS stacks (QGIS, ArcGIS); batch inference on satellite imagery.
- Assumptions/dependencies: Labeled data quality, domain shift across sensors/regions/seasons; compute for TTA at inference time; licensing of pretrained backbones; cloud inference costs.
- Neuroscience: fast whole-brain neural activity forecasting
- Use case: Predict multi-step neuronal activity from short context windows; assist in analyzing calcium/voltage imaging; enable closed-loop experiments.
- Sector: Academic neuroscience, neurotech, pharma preclinical.
- Tools/products/workflows: “NeuroForecast” module with cross-neuron conditioning and global context; specialized 1-step and multi-step models; JAX-based biophysical variants via Jaxley; rapid iteration in lab pipelines.
- Assumptions/dependencies: Standardized preprocessing; alignment between imaging modalities; validation on new animals/conditions; interpretability for biological insights; resource constraints in lab GPUs.
- General time-series forecasting (per-dataset and unified models)
- Use case: Demand/load/sales/claims/traffic forecasting; unified library competitive with foundation models on GIFT-Eval; adaptive presets with date/holiday/trend features.
- Sector: Retail, energy, finance, logistics, healthcare operations.
- Tools/products/workflows: “AutoTime” Python package and service; dataset adapters; automatic feature engineering (trend/seasonality/holidays), gradient boosting + ensembles; config presets with validation-based selection.
- Assumptions/dependencies: Availability of exogenous features (calendars, holidays, promotions); distribution shift handling; series with non-calendar dynamics; governance for automated model deployment.
- Numerical analysis: robust evaluation of difficult integrals
- Use case: More reliable numerical quadrature for integrals with singularities/cancellations where standard routines fail; simulation, control, pricing.
- Sector: Engineering, physics, quantitative finance, robotics.
- Tools/products/workflows: “RobustQuad” as a SciPy-compatible drop-in integrator; auto-selection of rules/transformations; test suite of difficult integrands; error diagnostics.
- Assumptions/dependencies: Thorough benchmark coverage; floating-point precision control; performance vs rigor trade-offs; reproducibility in safety-critical contexts.
- Literature-grounded replication and recombination of methods
- Use case: Re-implement published algorithms from PDFs/summaries; systematically recombine methods to surpass parents; track adherence and performance.
- Sector: Academia, R&D across domains, ML Ops.
- Tools/products/workflows: “Empirical Software Studio” with idea ingesters (papers/textbooks/search), adherence checks, tree search orchestrator, artifact registry, reproducibility reports, and model cards.
- Assumptions/dependencies: Access to papers and code licenses; accurate summarization of methods; strong evaluation harnesses to prevent “metric gaming”; compute for large searches.
- Research engineering workflow automation
- Use case: Metric-driven code evolution integrated into CI/CD for scientific software; automated hyperparameter and algorithmic exploration beyond manual intuition.
- Sector: Software/ML engineering, enterprise R&D, cloud platforms.
- Tools/products/workflows: GitHub/GitLab actions that trigger tree searches on benchmarks; sandboxed execution; automated reports; rollback on regressions; budget-aware search scheduling.
- Assumptions/dependencies: Secure sandboxing; quota and cost management; dataset versioning; deterministic evaluation; IP and code governance.
- Education and training for scientific computing
- Use case: Teaching empirical modeling through observable “breakthrough plots,” trees of ideas, and code adherence audits; rapid prototyping for student projects.
- Sector: Higher education, bootcamps, corporate upskilling.
- Tools/products/workflows: Classroom-ready datasets and rubrics as scorable tasks; instructor dashboards for method diversity/recombination; student-safe sandboxing.
- Assumptions/dependencies: Curated tasks with clear metrics; compute allocations; academic integrity policies for AI assistance.
- Small business and practitioner forecasting
- Use case: No-code forecasting for inventory, staffing, and cashflow; automated date-feature presets and validation-based selection.
- Sector: SMBs, operations, accounting.
- Tools/products/workflows: Lightweight SaaS with connectors (POS/ERP), holiday-aware presets, anomaly alerts, and simple uncertainty bands.
- Assumptions/dependencies: Data cleanliness and granularity; guardrails for overfitting; responsible defaults and explanations for non-experts.
Long-Term Applications
These opportunities build on the system’s core innovations (idea ingestion, tree search over code, recombination) but require more research, scaling, or integration with other systems.
- AI co-scientist for end-to-end hypothesis generation and testing
- Use case: Continuous literature mining, idea synthesis, code generation, evaluation, and iteration; autonomously exploring high-dimensional method spaces.
- Sector: Cross-disciplinary academia and industrial R&D (materials, drug discovery, climate).
- Tools/products/workflows: Multi-agent orchestration; literature QA; hypothesis trackers; experiment planners; knowledge graphs; cross-benchmark generalization reports.
- Assumptions/dependencies: Reliable retrieval; attribution and IP; scientific credit frameworks; safety and oversight; preventing evaluation leakage.
- Closed-loop lab automation and autonomous experimentation
- Use case: Connect algorithm discovery to robotic labs for wet/dry experiments; optimize protocols and models jointly.
- Sector: Biotech, chemistry, advanced manufacturing.
- Tools/products/workflows: Orchestration between lab robots (LIMS) and TS agent; safety constraints; active learning for experiment selection; digital twins.
- Assumptions/dependencies: Robust simulators; calibration between simulated and real outcomes; bio-safety and regulatory compliance; high reliability of evaluation metrics.
- Precision medicine analytics pipelines
- Use case: Automated batch integration and multimodal fusion (omics, imaging, EHR) for subtyping and biomarker discovery; adaptive models per hospital/cohort.
- Sector: Healthcare providers, pharma, diagnostics.
- Tools/products/workflows: Privacy-preserving/federated search; validation across institutions; explainability for clinical deployment; regulatory-grade documentation.
- Assumptions/dependencies: Data sharing constraints; FDA/EMA pathways; fairness/bias evaluation; clinical endpoint alignment.
- National/municipal early-warning and policy simulation systems
- Use case: Real-time epidemiological forecasting across pathogens; integration with wastewater, mobility, school/workplace signals; scenario planning and intervention simulation.
- Sector: Public policy, emergency management.
- Tools/products/workflows: Data fusion pipelines; counterfactual Monte Carlo and regime-switching detection at scale; interpretable dashboards; response playbooks.
- Assumptions/dependencies: Data access agreements; robust nowcasting for missingness; calibration under rapid shifts; ethical use of mobility and sensitive data.
- Global environmental monitoring and rapid mapping
- Use case: Continuous land-use updates, deforestation alerts, flood/fire damage assessment; fusion of satellite, drone, and IoT data.
- Sector: Climate tech, insurers, governments, ESG reporting.
- Tools/products/workflows: Foundation segmentation models with active recombination; uncertainty-aware mapping; on-demand edge inference; change detection.
- Assumptions/dependencies: Persistent labeling programs; sensor harmonization; latency and bandwidth constraints; standardized ESG/regulatory frameworks.
- Neuro-AI with structural-functional integration
- Use case: Combine functional forecasts with structural connectomes; hybrid interpretable models guiding stimulation or BCI strategies.
- Sector: Neurotech, medical devices, academia.
- Tools/products/workflows: Differentiable biophysical simulators + learned functional layers; safety testing for closed-loop; regulatory-ready telemetry and auditing.
- Assumptions/dependencies: Access to high-quality connectomes; on-device inference constraints; patient safety and IRB approvals; generalization across individuals.
- Time-series foundation models and privacy-preserving deployment
- Use case: Unified forecasting models that adapt to diverse domains with minimal supervision; federated/edge TS to respect data locality.
- Sector: Finance, healthcare ops, telecom, smart grids.
- Tools/products/workflows: Federated tree search; domain adapters; concept-drift monitors; confidential computing.
- Assumptions/dependencies: Transferability across domains; data minimization; explainability for high-stakes uses; cost-effective continual learning.
- Automated numerical method discovery
- Use case: Discover new quadrature/ODE/PDE solvers via recombination and search; hybrid symbolic–numeric procedures with provable error bounds.
- Sector: Engineering simulation, aerospace, energy, quantitative research.
- Tools/products/workflows: Formal verification layers; interval arithmetic; conformance test suites; compiler-level optimizations.
- Assumptions/dependencies: Theoretical validation; performance-portability to accelerators; acceptance in safety-critical pipelines.
- Governance, reproducibility, and audit tooling for AI-assisted science
- Use case: Standardized pipelines for reproducibility, adherence auditing, data provenance, and credit assignment for AI-generated methods.
- Sector: Academia, publishers, funders, industry R&D.
- Tools/products/workflows: Reproducibility badges; traceable idea/code lineage; benchmark registries; model cards and datasheets; automated IRB/compliance checks.
- Assumptions/dependencies: Community standards; interoperable metadata; incentives for sharing; alignment with regulatory frameworks.
- Education at scale via scorable-task curricula
- Use case: Transform curricula into scorable tasks with automated feedback, “breakthrough” visualizations, and safe idea recombination sandboxes.
- Sector: Universities, online learning platforms, workforce development.
- Tools/products/workflows: Instructor consoles; plagiarism-resistant task design; compute grants; student model portfolios.
- Assumptions/dependencies: Resource provisioning; equitable access; assessment redesign; clear AI-use policies.
Collections
Sign up for free to add this paper to one or more collections.