- The paper introduces CancerGUIDE, a hybrid framework that estimates LLM accuracy in NSCLC guideline adherence using proxy benchmarks and internal consistency signals.
- It employs synthetic supervision and consistency-based pseudo-labeling, achieving a Spearman correlation of 0.88 and an AUROC of 0.800 for treatment prediction accuracy.
- The framework supports unsupervised error detection and calibrated confidence scoring, reducing reliance on costly expert annotations for high-stakes clinical decisions.
CancerGUIDE: Estimating LLM Accuracy for Cancer Guideline Adherence via Internal Disagreement
Introduction
The "CancerGUIDE" paper addresses the challenge of evaluating LLMs for clinical guideline adherence, specifically in the context of non-small cell lung cancer (NSCLC) treatment recommendations. The work is motivated by the high cost and scarcity of expert-annotated datasets required for rigorous evaluation of LLMs in high-stakes medical applications. The authors propose a hybrid framework that leverages both proxy benchmarks and internal model consistency signals to estimate model accuracy and provide calibrated confidence scores for treatment recommendations, thereby supporting regulatory compliance and scalable deployment.
A key contribution is the creation of a longitudinal, expert-annotated dataset comprising 121 NSCLC patient cases, each annotated by board-certified oncologists with corresponding NCCN guideline trajectories. The annotation process involved tracing patient notes through the NCCN decision tree, recording node sequences, and assessing guideline adherence. Inter-annotator reliability was moderate (treatment match: 0.636, path overlap: 0.692), reflecting the inherent complexity and subjectivity of clinical reasoning.
The guideline-compliance prediction task is formalized as a structured prediction problem: given patient notes x, the model predicts a guideline-compliant path y through the decision tree. Two evaluation metrics are defined:
- Path Overlap: Jaccard similarity of nodes across predicted and reference paths.
- Treatment Match: Binary indicator of whether the final treatment matches the ground truth.
Proxy Benchmarking in Zero-Label Settings
To address the annotation bottleneck, the authors introduce six proxy benchmarking methods, divided into two main categories:
- Synthetic Supervision: Generation of synthetic patient notes and corresponding guideline paths, with structured and unstructured variants. The synthetic data pipeline includes consistency checks and LLM-based verification to filter out low-fidelity samples.
- Consistency-Based Pseudo-Labeling: Assignment of pseudo-labels based on self-consistency (agreement across multiple model rollouts) and cross-model consistency (agreement across different models), using both path overlap and treatment match as axes of agreement.
The effectiveness of these proxy benchmarks is evaluated by their correlation with expert-annotated ground truth. Notably, self-consistency pseudo-labels provide a robust proxy for benchmarking, achieving high Spearman correlation (r=0.88) and low RMSE (0.08) with expert annotations.
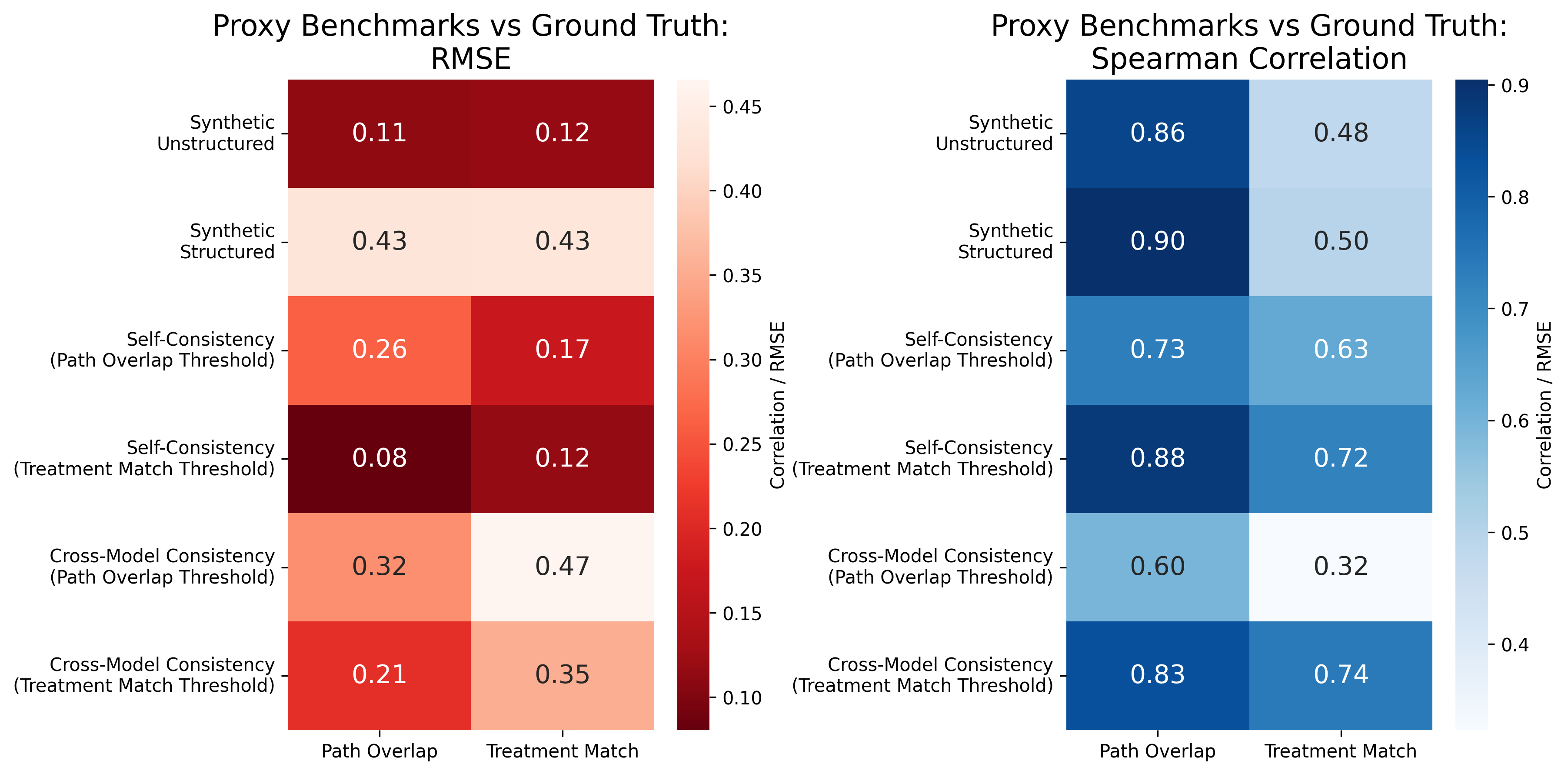
Figure 1: Self-consistency pseudo-labels provide a robust proxy for benchmarking, with strong correlation to expert-annotated benchmarks.
Consistency as a Predictor of Model Accuracy
A central finding is that model self-consistency—measured as the fraction of repeated predictions across multiple runs—correlates strongly with actual accuracy. This relationship holds for both path overlap and treatment match metrics, though the strength of the correlation varies by model family.
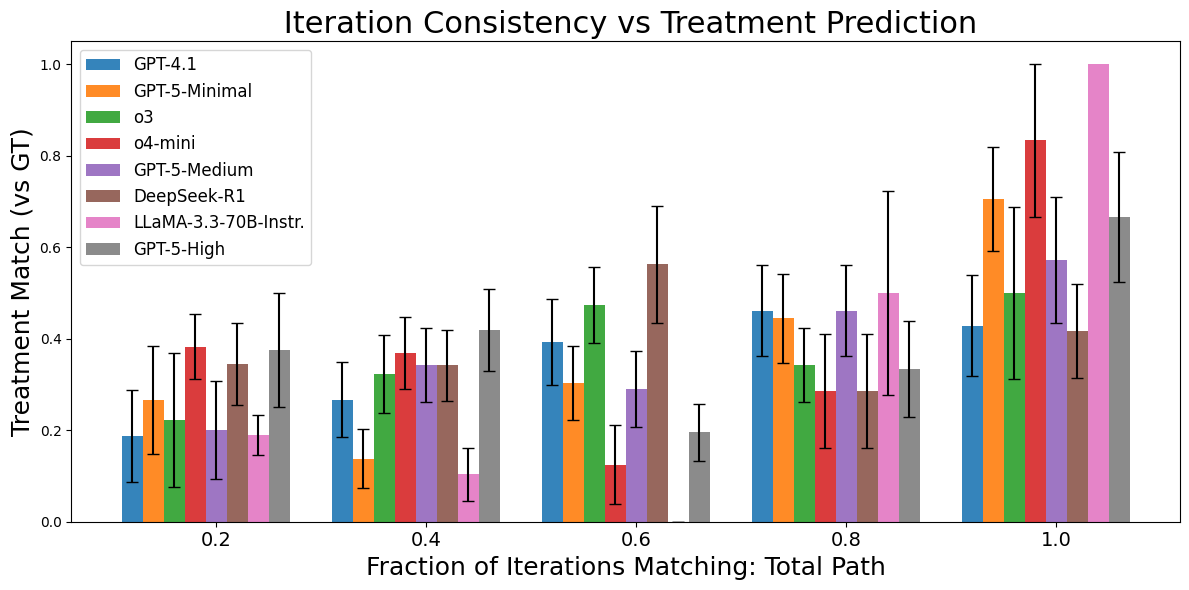
Figure 2: Model accuracy increases with self-consistency across prediction runs, supporting the use of consistency as a proxy for accuracy.
The authors develop a meta-classifier that uses features derived from self-consistency, cross-model consistency, and proxy benchmark scores to predict the correctness of individual treatment recommendations. The classifier achieves an average AUROC of 0.800 across models, enabling ROC-based performance reporting in line with regulatory requirements.
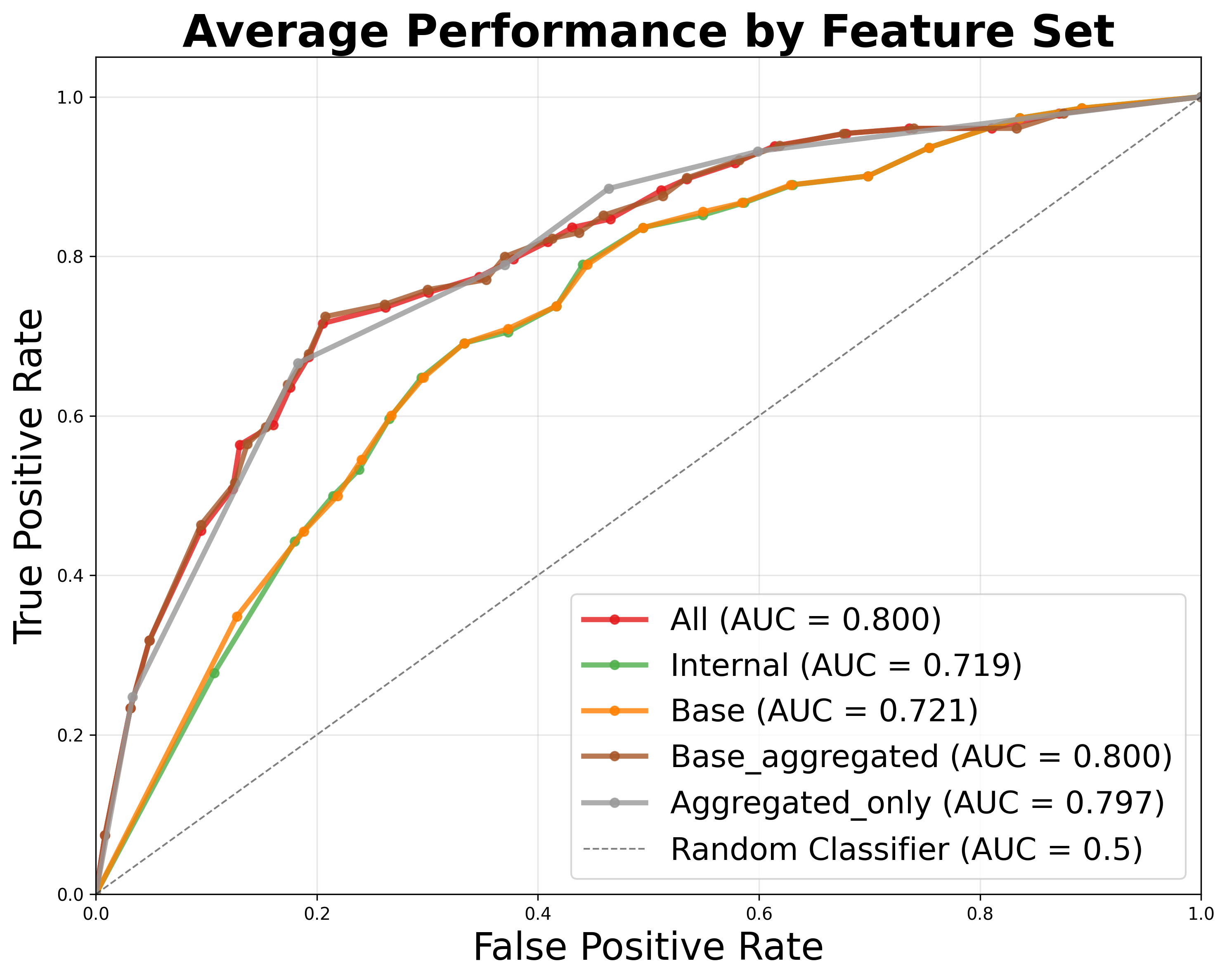
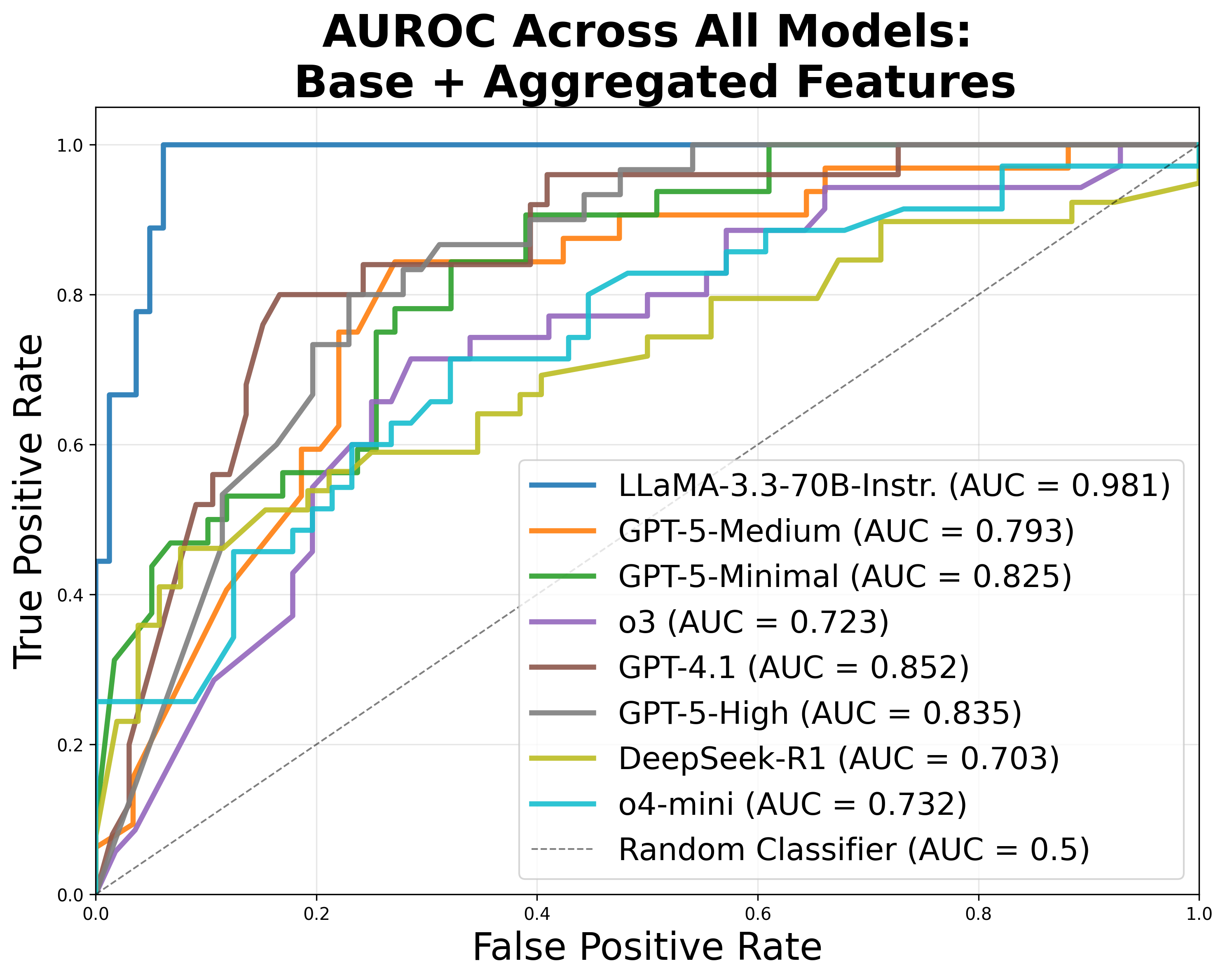
Figure 3: Signals from self- and cross-model consistency provide high AUC for accuracy classification, outperforming proxy benchmark results.
Error Analysis and Unsupervised Evaluation
The framework also supports unsupervised error detection. By clustering on consistency-derived features, the system can distinguish between correct and incorrect predictions with an F1 score of 0.666, even in the absence of human labels. Error analysis reveals that 40.42% of model errors on human-annotated data are detectable via internal disagreement, particularly around guideline compliance and tumor staging.
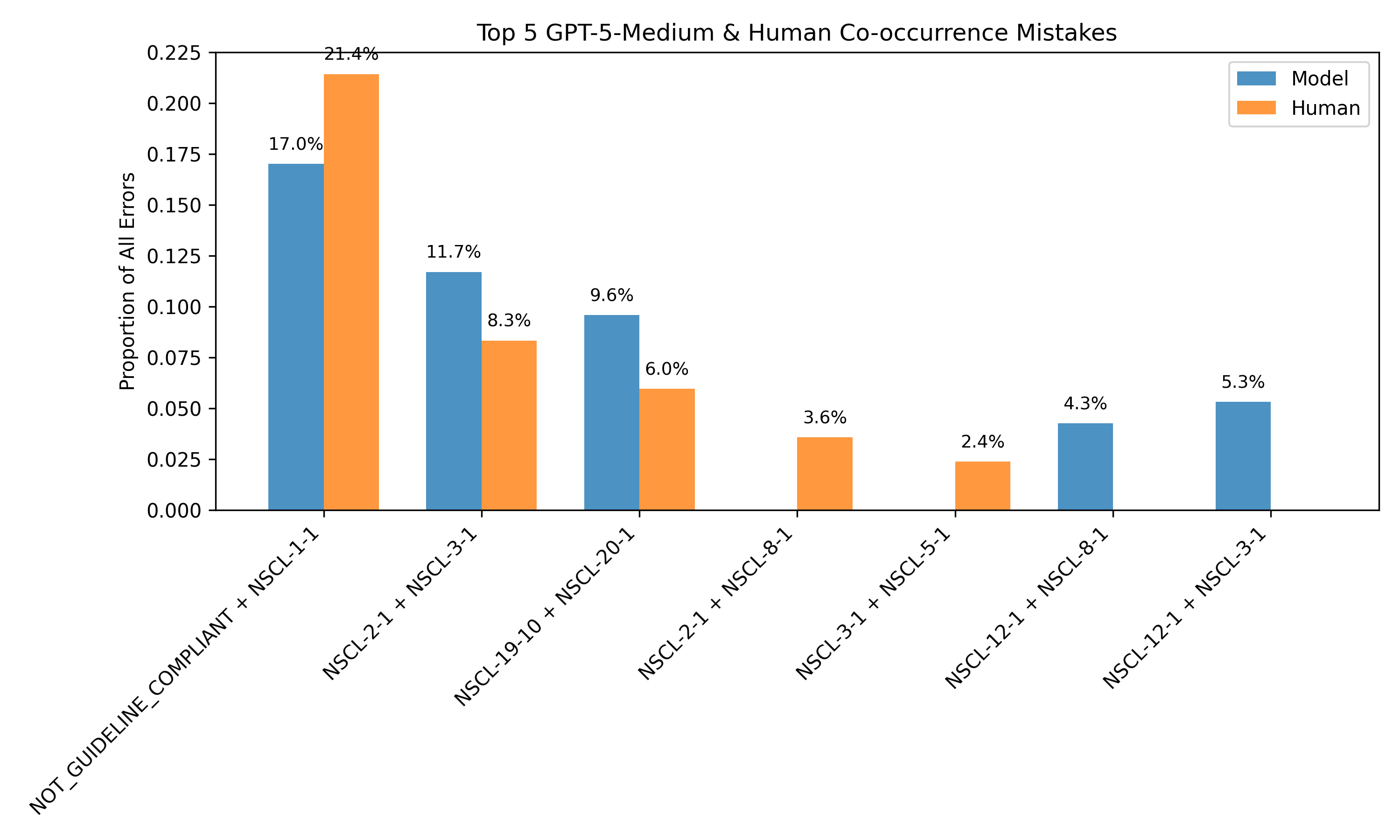
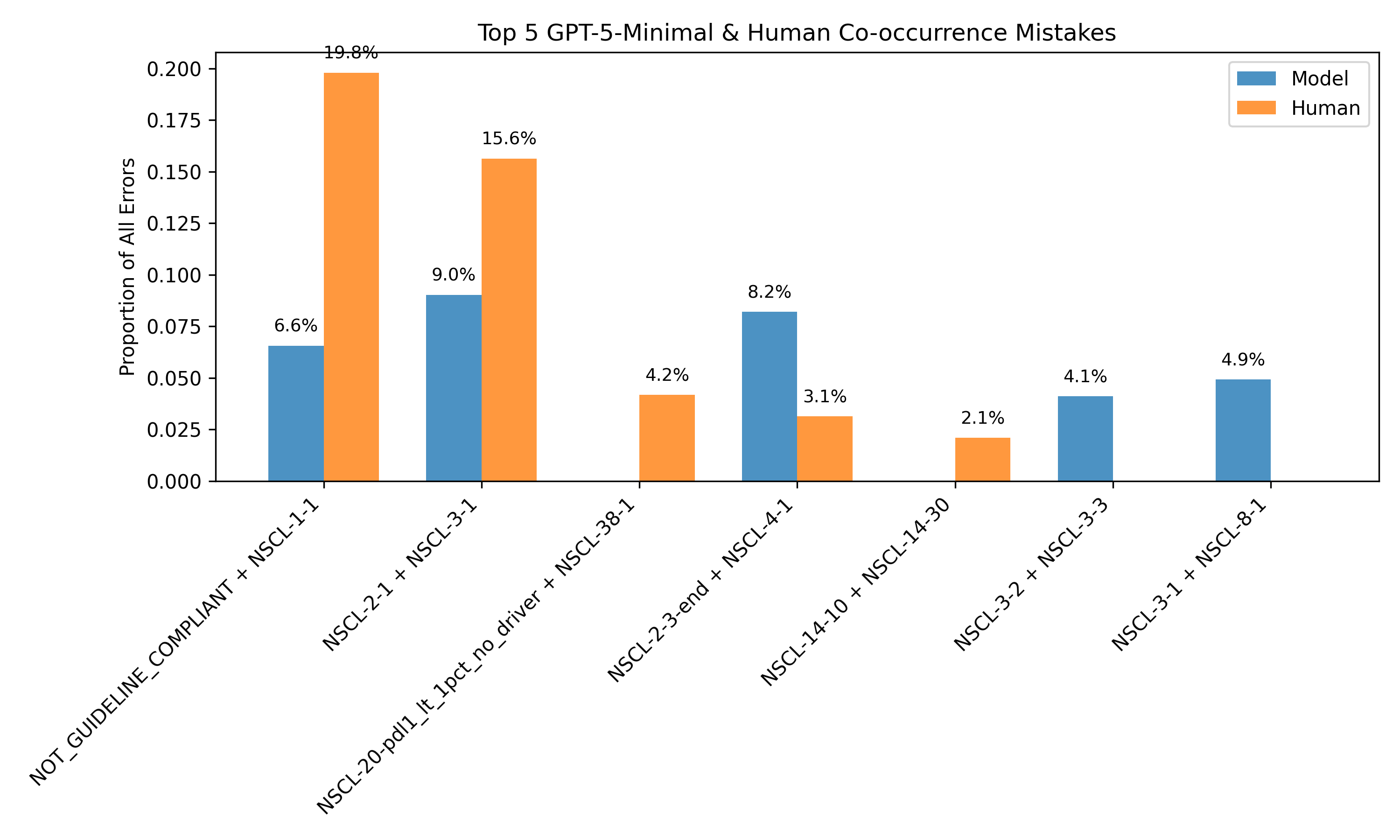
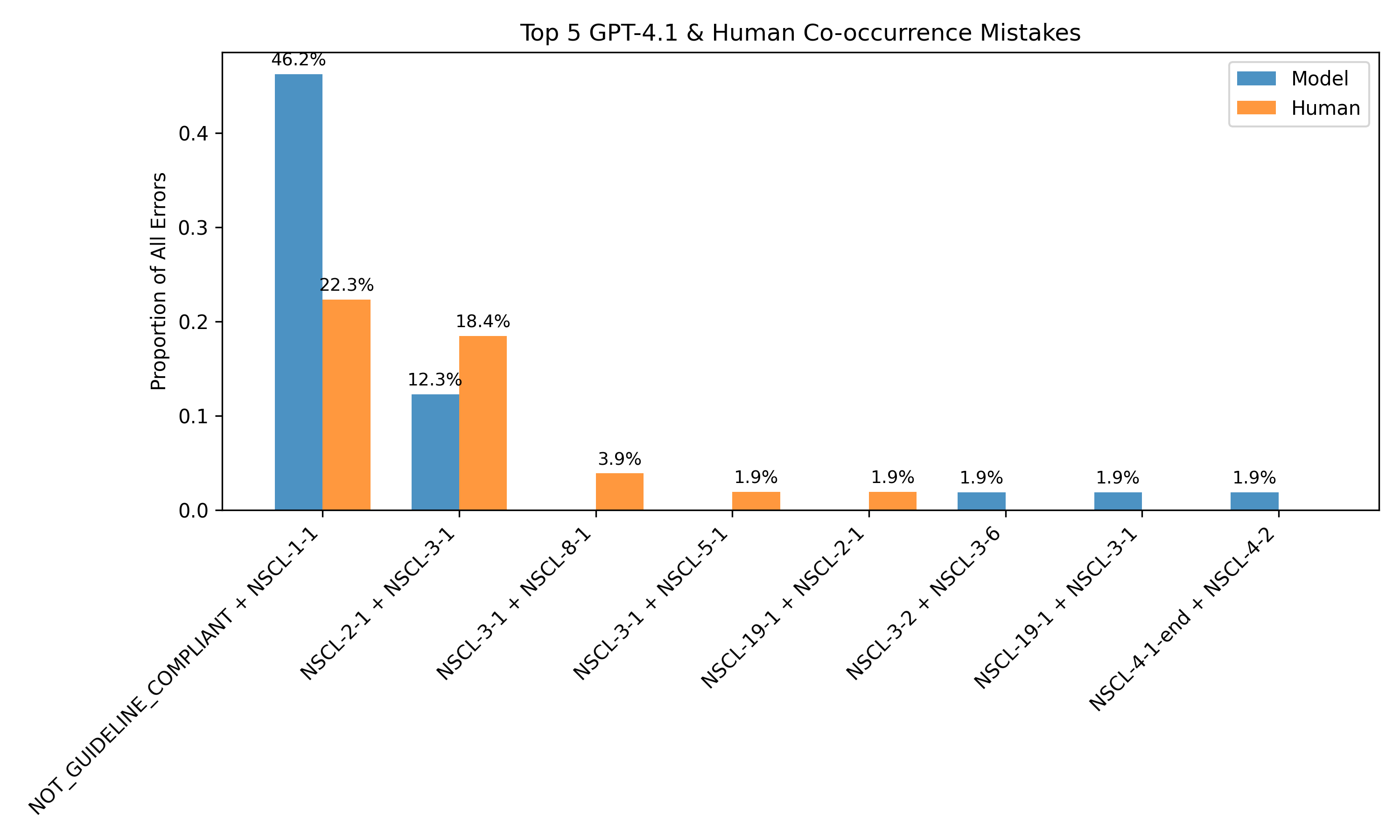
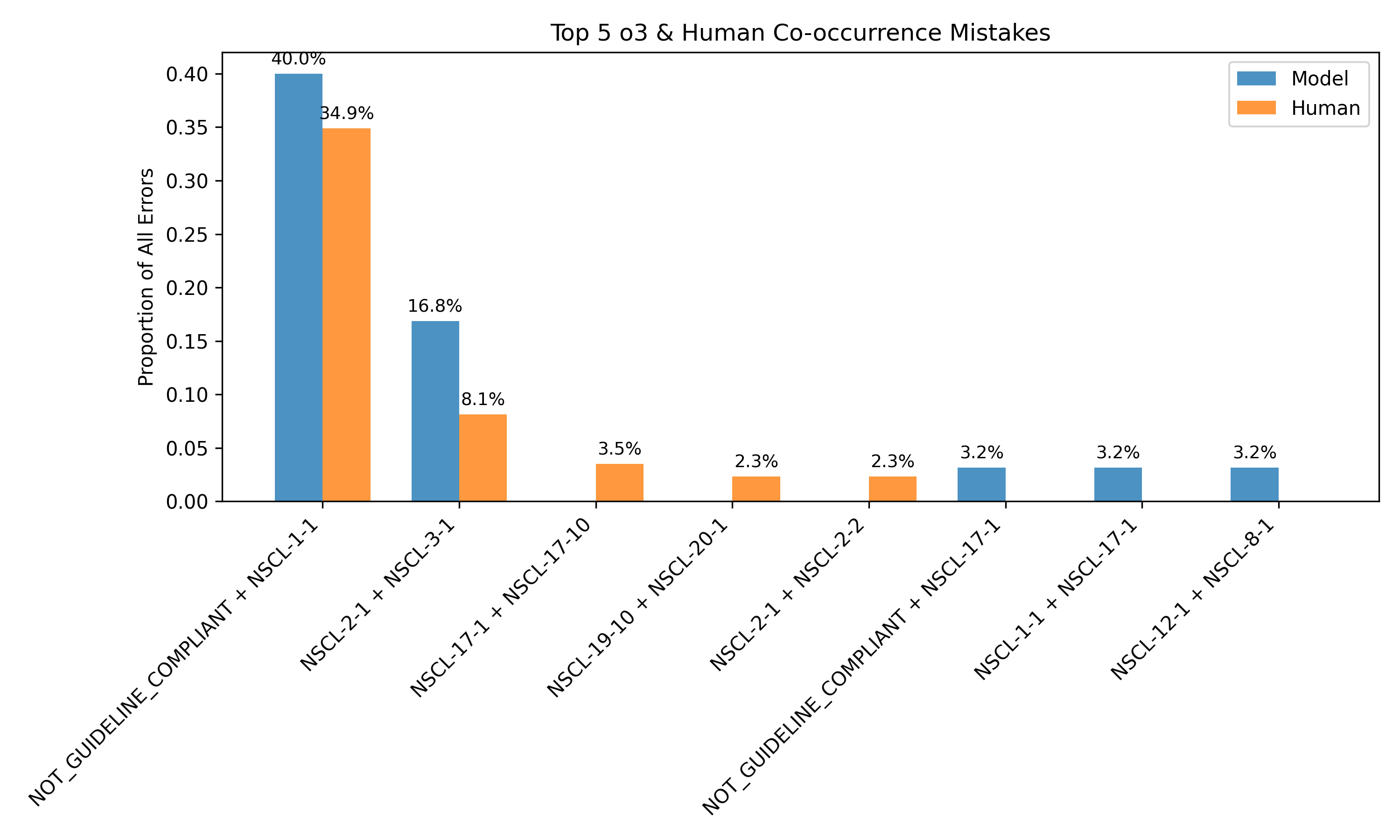
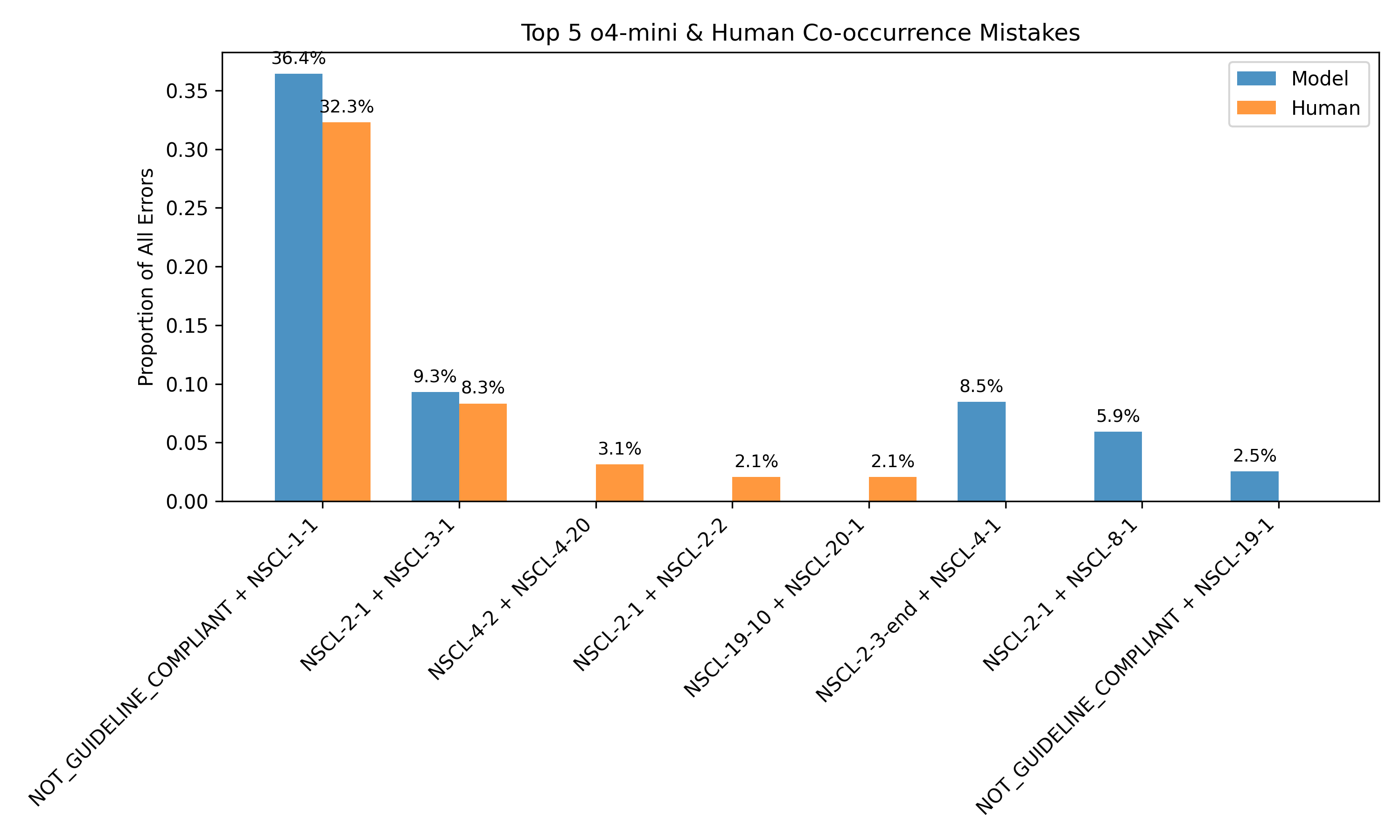
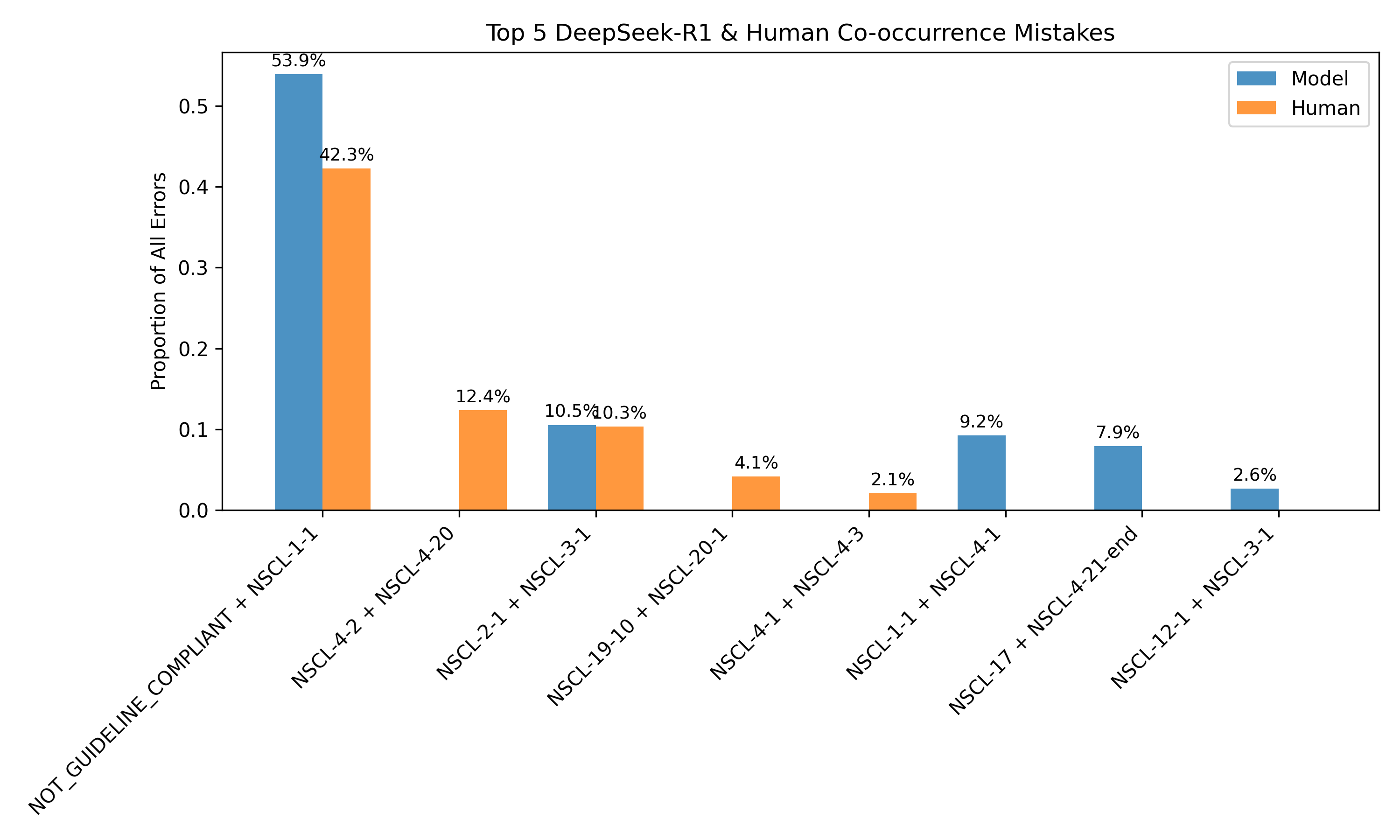
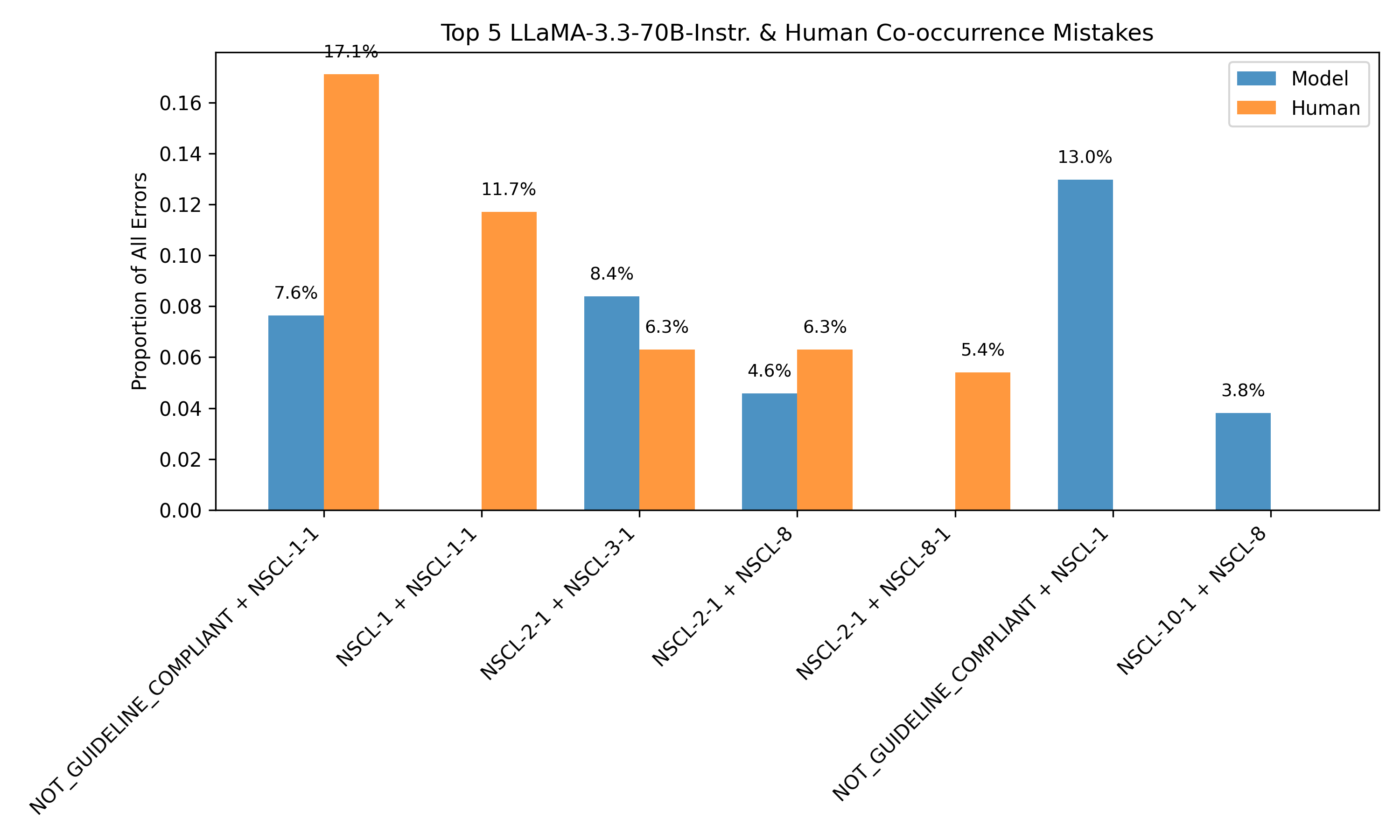
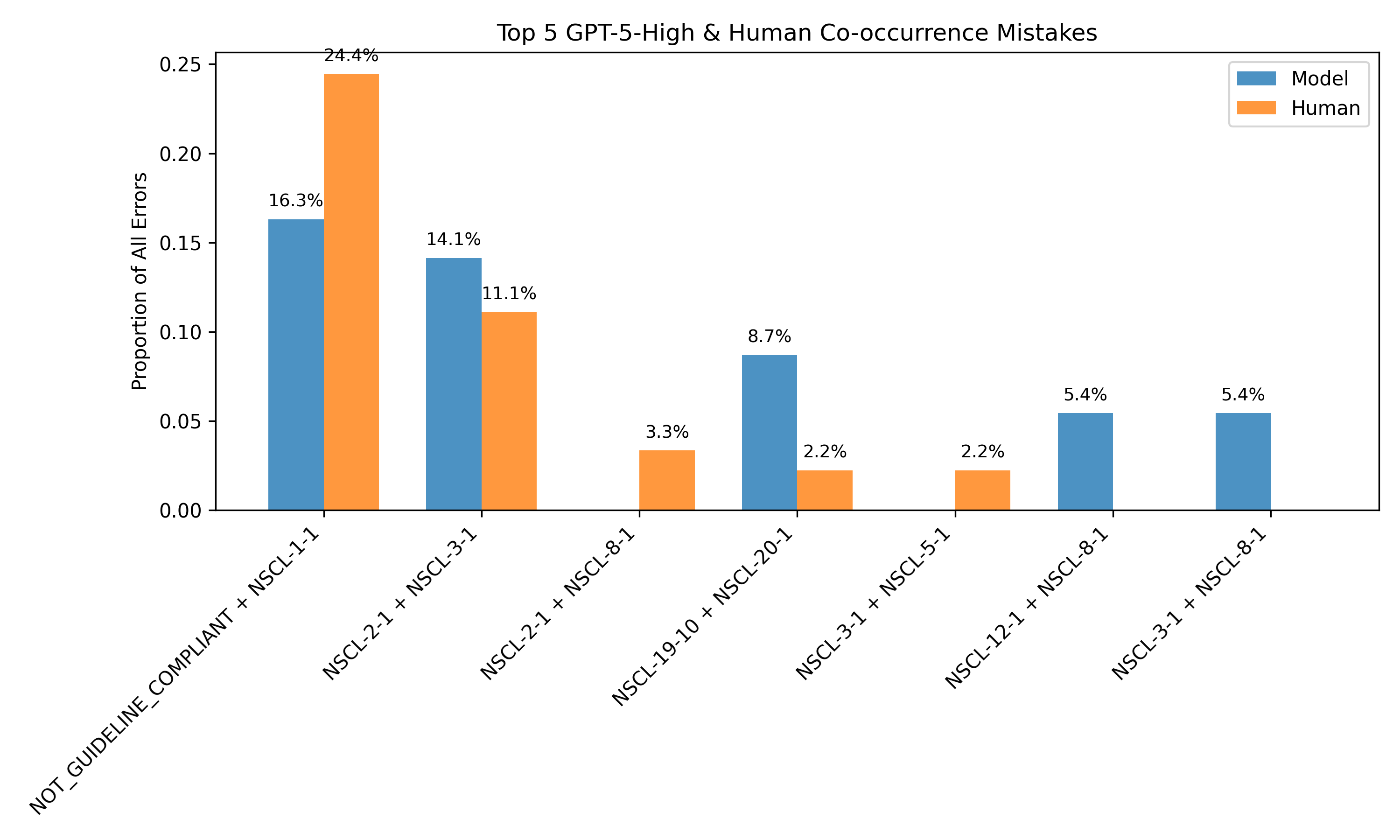
Figure 4: Most common discrepancies between human and model annotations align with discrepancies observed in model rollouts, enabling error identification without human labels.
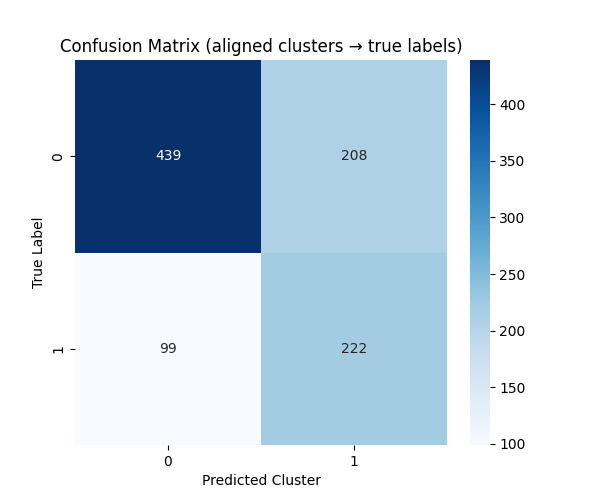
Figure 5: Confusion matrix for K-means clustering over consistency-derived features, demonstrating unsupervised separation of correct and incorrect predictions.
Evaluation on the expert-annotated dataset shows that the best-performing models (e.g., GPT-5, o3) achieve path overlap scores around 0.45 and treatment match rates around 0.36, indicating substantial room for improvement in clinical accuracy. Feature ablation studies demonstrate that cross-model consistency is the strongest predictor of accuracy, but self-consistency alone remains informative, especially when cross-model information is unavailable.
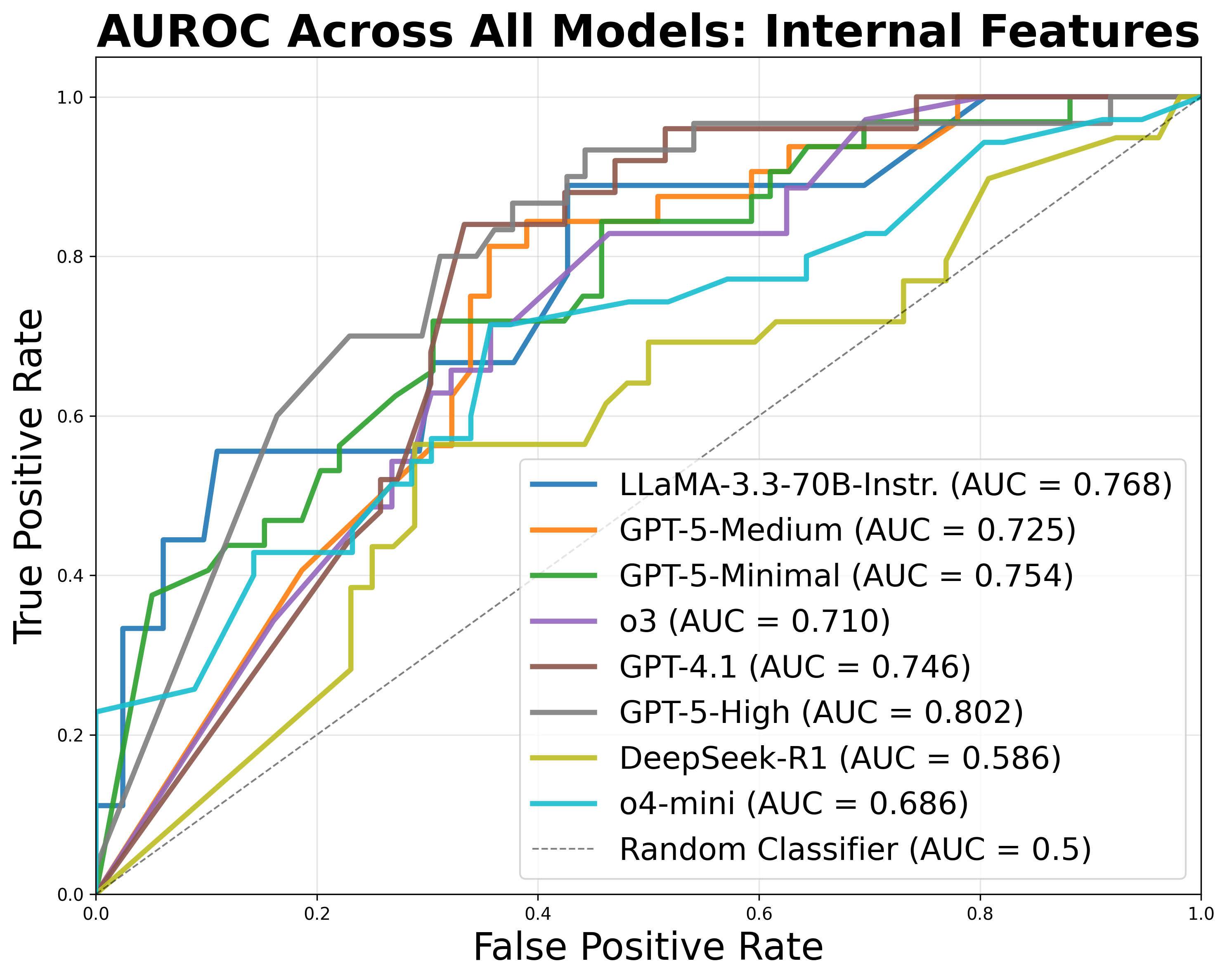
Figure 6: ROC curves for classification of treatment prediction accuracy using exclusively internal features, showing robust performance even without cross-model signals.
Implications and Future Directions
The CancerGUIDE framework provides a scalable, interpretable, and regulation-aligned approach for evaluating LLMs in clinical guideline adherence tasks. By leveraging internal disagreement and proxy benchmarks, the system reduces reliance on costly expert annotation and enables real-time confidence estimation for model outputs. This is particularly relevant for deployment in clinical decision support, where safety, transparency, and regulatory compliance are paramount.
The methodology is extensible to other cancer types and clinical domains, contingent on the availability of structured guidelines and annotated datasets. Future work should focus on expanding the dataset, increasing the diversity of annotated cases, and exploring adaptive learning strategies that explicitly model human uncertainty. Additionally, integrating proxy benchmark data into model alignment pipelines may further mitigate data bottlenecks and improve downstream performance.
Conclusion
CancerGUIDE demonstrates that internal disagreement estimation—via self- and cross-model consistency—enables robust, scalable evaluation of LLMs for cancer guideline adherence, even in zero-label settings. The framework achieves strong correlation with expert benchmarks, supports unsupervised error detection, and provides calibrated confidence scores for regulatory reporting. These advances represent a significant step toward practical, trustworthy deployment of LLMs in high-stakes clinical decision support.













