- The paper introduces a Bayesian pliable lasso that integrates a horseshoe prior to enforce heredity and quantify uncertainty in high-dimensional GLMs with missing responses.
- It employs a Gibbs sampler with data augmentation to efficiently impute missing outcomes and achieve rapid posterior convergence.
- Simulation studies and a neuroimaging application demonstrate superior estimation, prediction, and variable selection compared to existing methods.
Bayesian Pliable Lasso with Horseshoe Prior for Interaction Effects in GLMs with Missing Responses
Introduction and Motivation
The paper introduces a Bayesian extension of the pliable lasso for high-dimensional regression and generalized linear models (GLMs), focusing on the estimation of both main effects and structured interactions between predictors and modifying covariates. The frequentist pliable lasso enforces strong heredity constraints and sparsity, but lacks a mechanism for uncertainty quantification and prior incorporation. The Bayesian formulation leverages the horseshoe prior to induce global-local shrinkage on both main and interaction effects, naturally enforcing heredity and enabling principled posterior inference. The framework is further extended to handle missing responses via data augmentation, making it applicable to real-world datasets with incomplete outcomes.
The pliable lasso model is generalized to GLMs by parameterizing the canonical link as:
ηi=β0+Zi⊤θ0+j=1∑pxij(βj+Zi⊤θj)
where xij are predictors, Zi are modifying covariates, βj are main effects, and θj are vectors of interaction effects. The strong heredity constraint is enforced by the hierarchical prior: θj can be nonzero only if βj is nonzero.
The horseshoe prior is placed jointly on (βj,θj) for each j, with shared local and global scale parameters:
βj∼N(0,λj2τ2),θj∼N(0,λj2τ2Iq),λj∼Cau+(0,1),τ∼Cau+(0,1)
This structure ensures adaptive shrinkage, aggressively shrinking noise while preserving large signals, and couples the main and interaction effects in accordance with the heredity constraint.
Posterior Inference via Gibbs Sampling
Efficient posterior computation is achieved through a Gibbs sampler exploiting the inverse-gamma representation of the half-Cauchy prior [makalic_simple_2016]. The sampler cycles through updates for:
- Main effects βj and interaction effects θj (Gaussian conditionals)
- Local and global shrinkage parameters (λj2, τ2; inverse-gamma conditionals)
- Intercept terms (β0, θ0; Gaussian conditionals)
- Noise variance σ2 (inverse-gamma conditional)
- Imputation of missing responses (Gaussian conditional for each missing yi)
The imputation step treats missing responses as latent variables, drawing from their full conditional given current parameter values. This approach assumes ignorable missingness and does not require modification of other conditional distributions.
Simulation Studies
Extensive simulations are conducted across a range of settings, varying the type and correlation structure of predictors and modifiers, sample size, and proportion of missing data. The pliable horseshoe (pHS) is compared against the frequentist pliable lasso (pLasso), standard lasso, and standard horseshoe (HS).
Key findings:
- Estimation and Prediction: pHS consistently achieves the lowest estimation and prediction errors across all settings, especially when modifiers are binary or in high-dimensional regimes.
- Variable Selection: Both HS and pHS excel in variable selection accuracy, with pHS outperforming in small sample sizes and in the presence of interactions.
- Robustness to Missing Data: pHS maintains strong performance up to moderate levels of missingness; degradation is observed only at extreme missingness (e.g., 70%).
- High-Dimensional Scalability: pHS outperforms competitors in scenarios with p≫n, maintaining low estimation and prediction errors and high selection accuracy.
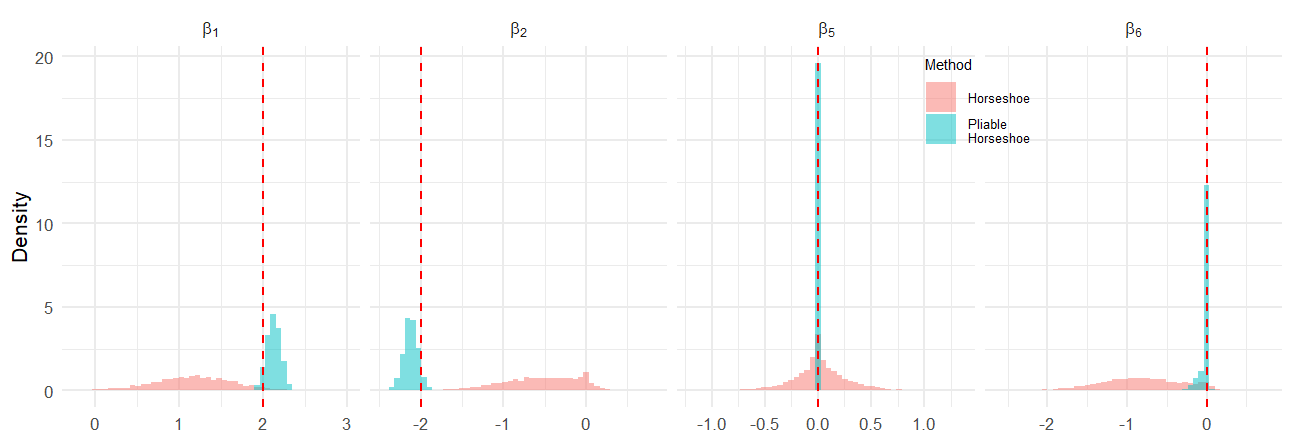
Figure 1: Histogram plots comparing posterior distributions for selected βj under HS and pHS in Setting I (n=200,p=10,q=4), illustrating improved recovery of true values by pHS.
Trace and autocorrelation plots (see Appendix) confirm rapid mixing and convergence of the Gibbs sampler for both main and interaction effects.
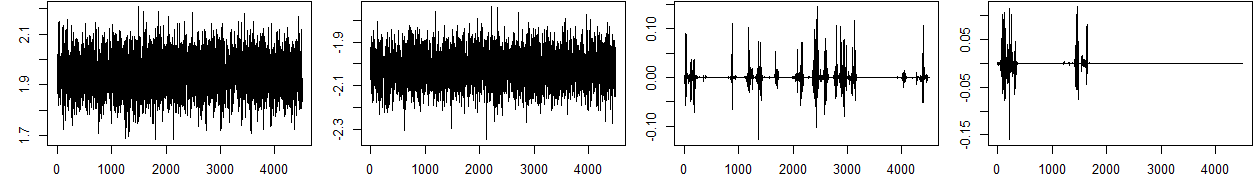
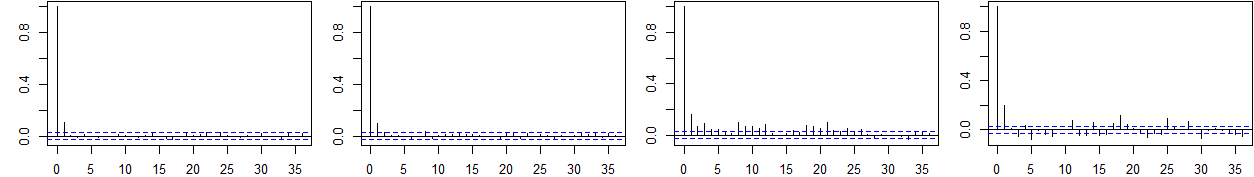
Figure 2: Trace and ACF plots for selected βj in Setting I, demonstrating efficient sampling and convergence properties of the Gibbs sampler.
Application to Neuroimaging Data
The method is applied to the OASIS Brain Data, predicting right hippocampal volume from demographic and neuroimaging covariates, with dementia status as a modifier. pHS and pLasso both identify normalized whole brain volume (nWBV) as a key predictor, with pHS providing wider credible intervals reflecting posterior uncertainty. Interaction effects are estimated but not found to be significant. In repeated train/test splits, pHS and pLasso yield the lowest prediction errors, outperforming HS and lasso.
Implementation and Practical Considerations
The method is implemented in the R package hspliable, leveraging Rcpp for computational efficiency. The Gibbs sampler is tractable for moderate p and q; for very large-scale problems, further acceleration (e.g., via variational inference) is warranted. The approach is directly extensible to other exponential family models (e.g., logistic, Poisson) via appropriate data augmentation (e.g., Polya-Gamma for logistic regression).
Theoretical and Practical Implications
The Bayesian pliable lasso with horseshoe prior provides a principled framework for sparse interaction modeling in high-dimensional GLMs, with automatic uncertainty quantification and heredity enforcement. The joint shrinkage mechanism is theoretically optimal for sparse regimes and empirically robust to challenging data structures, including missingness and high dimensionality. The approach is particularly suited to scientific domains requiring interpretable effect-modifier relationships, such as genomics, personalized medicine, and neuroimaging.
Future Directions
Potential extensions include scalable inference for ultra-high-dimensional data, adaptation to survival and nonparametric models, and explicit modeling of non-ignorable missingness mechanisms. The framework is also amenable to hierarchical and multi-level interaction structures, broadening its applicability.
Conclusion
The Bayesian pliable lasso with horseshoe prior advances the state-of-the-art in high-dimensional interaction modeling for GLMs, offering superior estimation, prediction, and variable selection performance, robust handling of missing data, and interpretable uncertainty quantification. The method is practically implemented and validated on both synthetic and real-world biomedical data, with clear advantages over existing frequentist and Bayesian approaches.


