- The paper introduces a novel explicit 3D Gaussian representation that models protein density directly in real-space.
- The methodology employs closed-form 2D projection and gradient pruning to markedly reduce memory usage and accelerate training.
- Experimental results demonstrate that GEM achieves state-of-the-art resolution and robust performance, nearing physical imaging limits.
GEM: 3D Gaussian Splatting for Efficient and Accurate Cryo-EM Reconstruction
Introduction
The GEM framework introduces a novel approach to cryo-electron microscopy (cryo-EM) reconstruction by leveraging 3D Gaussian Splatting (3DGS) for explicit, real-space modeling of protein density. Traditional Fourier-space methods, while computationally efficient, suffer from information loss due to repeated transforms and interpolation. Recent neural field-based real-space methods (e.g., NeRFs, Transformers) improve accuracy but incur prohibitive cubic memory and computation costs, especially for large datasets and high-resolution reconstructions. GEM circumvents these limitations by representing protein density as a compact set of 3D Gaussians, each parameterized by 11 values, and introducing efficient gradient computation restricted to the subset of Gaussians contributing to each voxel. This design yields substantial improvements in both computational efficiency and reconstruction fidelity.
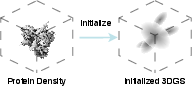
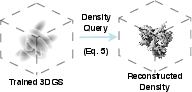
Figure 1: Protein density is represented by 3D Gaussians. Before training, the parameters of 3D Gaussians are randomly initialized.
Methodology
Explicit 3D Gaussian Representation
GEM models the protein density as a sum of M 3D Gaussians, each defined by a center pj, covariance Σj (decomposed into rotation Rj and scale Sj), and density coefficient ρj. The overall density at position x is:
V^(x)=j=1∑Mρjexp(−21(x−pj)⊤Σj−1(x−pj))
This explicit representation is both sparse and local, as Gaussians are instantiated only in nonzero-density regions, and each contributes to multiple pixels during projection.
Efficient 2D Projection and Differentiable Rendering
The projection of the 3D Gaussian mixture onto 2D is performed via closed-form integration along the optical axis, yielding a 2D Gaussian mixture for each view. This avoids the need to query the full 3D grid, substantially reducing memory and compute requirements. The projected image I^i is then convolved with the microscope's contrast transfer function (CTF) in Fourier space, and the loss is computed as the ℓ2 distance to the observed noisy image.
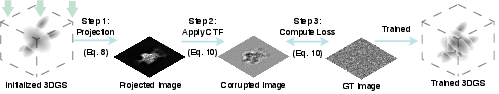

Figure 2: The overview of GEM training. The training begins with randomly initialized Gaussians. The 3D Gaussians are projected following Equation \ref{eq:render}.
Gradient Pruning and Parallel Rasterization
To further optimize efficiency, GEM introduces a thresholding strategy: only Gaussians with non-negligible contributions to a given pixel are considered during gradient computation. This pruning, combined with parallel rasterization, eliminates the cubic scaling of memory and computation inherent to neural field methods. The remaining Gaussians are sorted along the z-axis for efficient accumulation.
Experimental Results
Efficiency and Scalability
GEM demonstrates up to 48× faster training and 12× lower memory usage compared to state-of-the-art baselines (CryoDRGN, CryoNeRF), as measured on four standard cryo-EM datasets. The memory footprint is stable and primarily determined by the number of Gaussians, enabling practical deployment on commodity GPUs.
Reconstruction Quality
Gold-Standard Fourier Shell Correlation (GSFSC)
GEM consistently achieves superior GSFSC resolution, with intersection points at the $0.143$ threshold further to the right (lower Å values) than baselines, indicating higher reconstruction quality. On datasets EMPIAR-10049 and EMPIAR-10028, GEM approaches the Nyquist-Shannon limit, demonstrating near-optimal recovery of fine structural details.
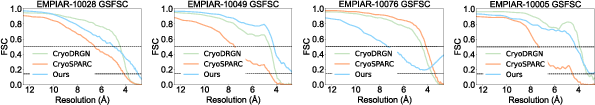
Figure 3: GSFSC of GEM and baselines. Intersection points further to the right correspond to better reconstruction quality.
Local Resolution Estimation
GEM yields higher local resolution across all datasets, with larger regions in blue-green on the resolution maps, indicating more accurate and detailed reconstructions. On challenging datasets, GEM attains local resolution close to the physical limit imposed by the microscope.
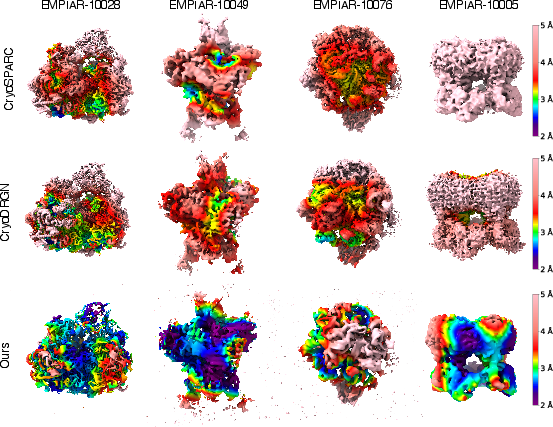
Figure 4: Local resolution maps of reconstructions from GEM, CryoSPARC, and CryoDRGN. Larger blue-green regions indicate better local resolution.
Fourier Slice Correlation (FSLC)
GEM achieves higher and more isotropic FSLC values, with more uniform yellow regions on the correlation maps, reflecting robust directional consistency and stability across sampled orientations.
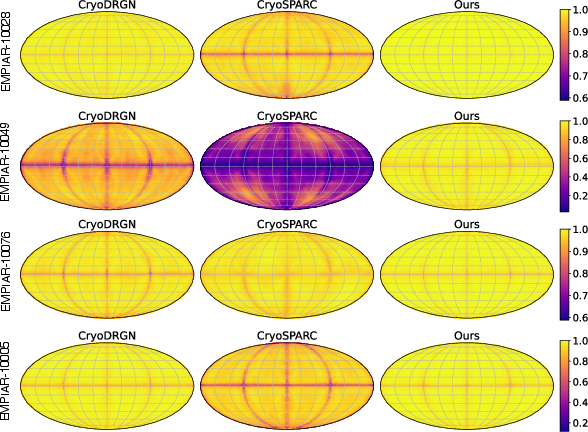
Figure 5: FSLC maps of reconstructions from GEM, CryoSPARC, and CryoDRGN. More uniform yellow regions indicate higher and more isotropic directional resolution.
Ablation Studies
Ablations on per-Gaussian rotation and anisotropic scaling reveal that both are critical for accurate cryo-EM modeling. Enforcing isotropic scaling or removing rotation degrades resolution, with combined constraints causing the most severe performance drop.
Implementation Considerations
- Parameterization: Each Gaussian requires 11 parameters (center, rotation, scale, density), enabling compact representation.
- CUDA Kernels: Closed-form projection and thresholded accumulation are amenable to efficient GPU implementation.
- Memory Management: The number of active Gaussians can be tuned to balance resolution and resource usage.
- Gradient Localization: Restricting gradient computation to contributing Gaussians is essential for scalability.
- Compatibility: GEM integrates with standard cryo-EM pipelines and evaluation protocols, facilitating adoption.
Implications and Future Directions
GEM establishes 3DGS as a practical and scalable paradigm for cryo-EM reconstruction, unifying speed, efficiency, and high-resolution accuracy. The explicit, sparse, and local nature of the representation enables tractable modeling of large macromolecules and heterogeneous structures. The approach is extensible to heterogeneous reconstruction, multi-state modeling, and integration with downstream tasks such as atomic model fitting and conformational analysis. Future work may explore adaptive Gaussian placement, joint optimization with imaging latents, and extension to time-resolved or multi-modal datasets.
Conclusion
GEM leverages 3D Gaussian Splatting to overcome the limitations of both Fourier-space and neural field-based cryo-EM reconstruction methods. By explicitly modeling protein density with compact, parameter-efficient Gaussians and introducing efficient gradient computation, GEM achieves state-of-the-art resolution, speed, and memory efficiency. The framework approaches the physical resolution limits of cryo-EM, offering substantial practical and theoretical benefits for structural biology and computational imaging.





