- The paper introduces HELM, an equivariant graph neural network that pretrains on Hamiltonian matrices to reduce energy prediction error by up to 2× in low-data regimes.
- It employs an SE(3)-equivariant backbone with parity expansions and Clebsch-Gordan decompositions, effectively handling complex orbital and heavy-element interactions.
- The study demonstrates that leveraging detailed electronic-structure labels greatly enhances molecular representation and generalization across diverse atomic configurations.
Learning Molecular Electronic Structure Across the Periodic Table with HELM
Introduction
This work presents HELM (Hamiltonian-trained Electronic-structure Learning for Molecules), an equivariant graph neural network architecture for scalable Hamiltonian matrix prediction, specifically designed to bridge the gap between electronic-structure learning and current state-of-the-art machine-learned interatomic potentials (MLIPs). The authors also introduce OMol_CSH_58k, a diverse and extensive molecular Hamiltonian matrix dataset, and demonstrate that pretraining on electronic structure enables significant improvements in downstream energy prediction, particularly in low-data regimes. The approach leverages the vastly richer supervision available from quantum chemical Hamiltonians compared to conventional training solely on energies and forces, thereby enabling both data efficiency and generalization over unprecedented molecular and chemical complexity.
HELM: A Scalable Architecture for Hamiltonian Learning
HELM integrates symmetry principles and efficient message passing to predict Hamiltonians of large molecular systems with high elemental and structural diversity. The model employs an SE(3)-equivariant backbone encoding atomic and inter-atomic features as spherical harmonic coefficient embeddings, processed through layers incorporating SO(2) convolutions, parity-expansion blocks for diagonal (intra-atomic) interactions, and gated nonlinearities to manage high-orbital multiplicities and heavy elements.
The backbone simultaneously produces atom- and edge-specific vector-valued features; these are mapped to irreducible representations (irreps) corresponding to blocks of the Hamiltonian matrix, decomposed using Clebsch-Gordan coefficients. The architecture supports arbitrary atomic configurations and basis set expansions up to high angular momentum (e.g., f-orbitals), facilitating application to large organic and inorganic molecules.
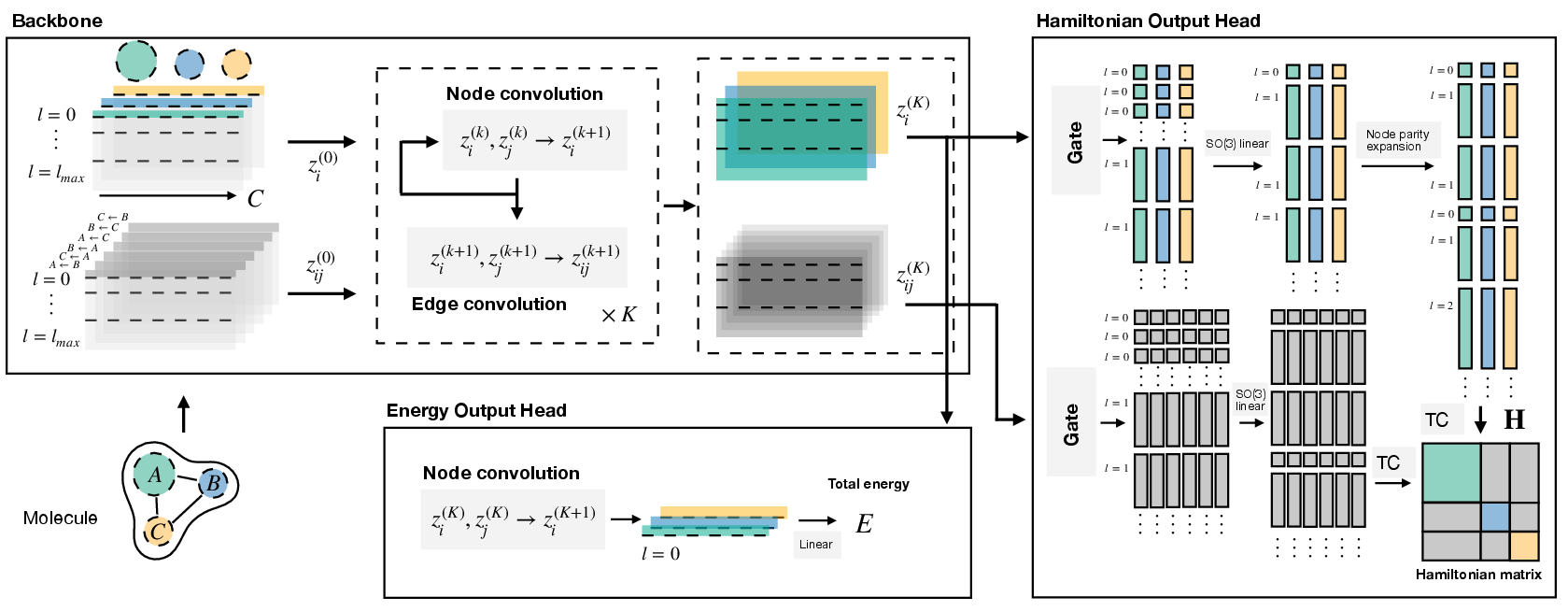
Figure 1: Architecture of HELM, displaying atom- and inter-atomic interaction embeddings parameterized by lmax and channel count, with separate heads for Hamiltonian and energy prediction.
The OMol_CSH_58k Dataset
To match the increasing chemical diversity in force/energy datasets, OMol_CSH_58k was curated to provide high-quality supervised targets for Hamiltonian learning. It contains over 56,000 molecular structures with up to 150 atoms, spanning 58 elements (first 83 elements excluding 3d transition metals and lanthanides for computational tractability with f-orbitals and basis set size), with each structure annotated by a Kohn-Sham Hamiltonian in the large def2-TZVPD basis. This dataset ensures broad coverage of atomic interactions, orbital configurations, interatomic distances, and element-element interactions, providing a challenging benchmark for generalization and robust representation learning.
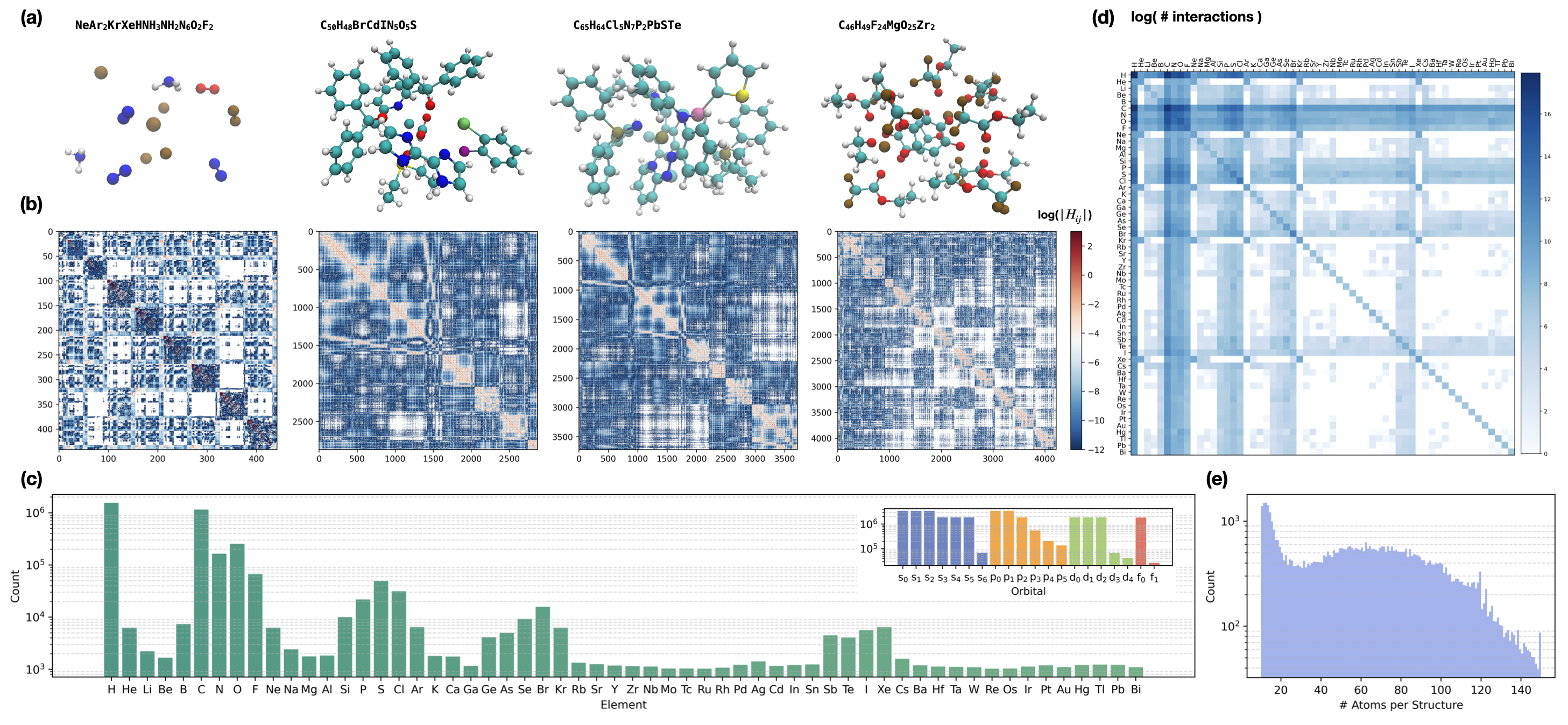
Figure 2: OMol_CSH_58k visualizes molecular structures, Hamiltonian element distributions, element coverage histograms, and interaction diversity across atomic pairs and structure sizes.
Hamiltonian Pretraining: Transfer to Energy Prediction
The central algorithmic contribution is Hamiltonian pretraining: HELM’s backbone is first trained to predict matrix elements of the Kohn-Sham Hamiltonian, leveraging the O(N2) matrix element labels per structure, and then the shared embedding space is repurposed as input for a separate head trained/fine-tuned on total energy data. This inductive transfer is shown to yield superior atomic environment descriptors, particularly for under-represented elements and large molecules. Three regimes are considered:
- Direct energy training—the network is optimized solely on energy labels.
- Pretrained-frozen—the backbone is frozen after Hamiltonian training, with only the energy head trained.
- Pretrained-finetuned—the entire network is fine-tuned for energy after Hamiltonian pretraining.
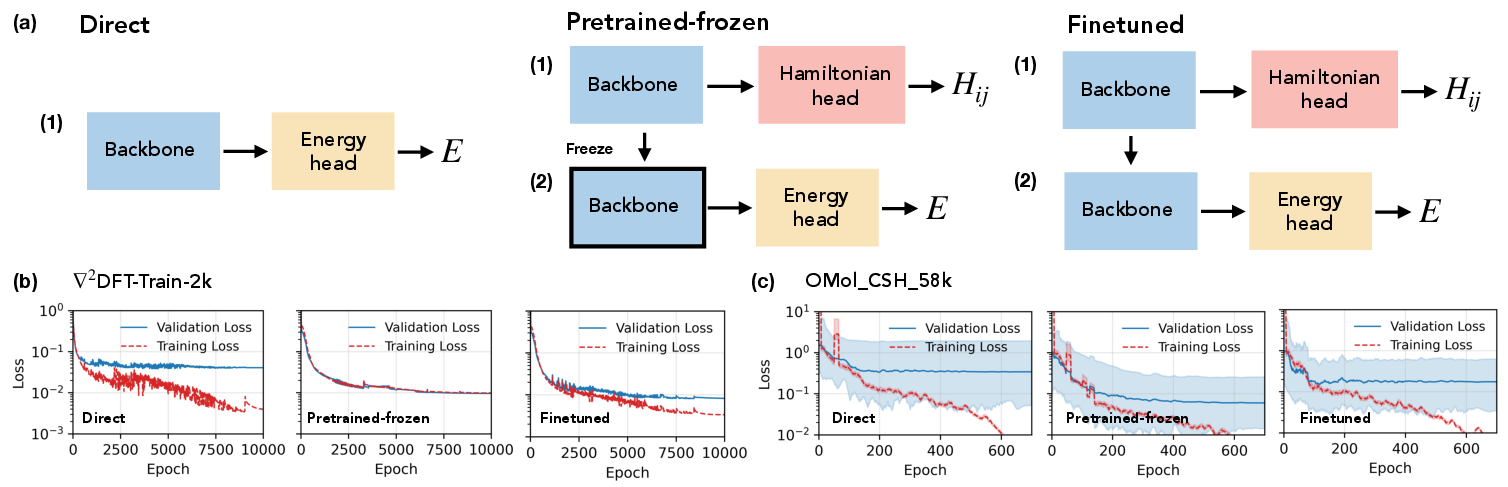
Figure 3: Comparison of direct, pretrained-frozen, and finetuned training schemes, with corresponding energy loss curves demonstrating accelerated and stabilized convergence in pretraining setups.
Empirically, pretraining results in up to a 2× reduction in energy test MAE on low-data splits compared to direct energy regression with identical model capacity—a gain otherwise requiring an order-of-magnitude increase in energy/force training data volume. Improvements are most pronounced in few-shot settings and for predicting properties associated with heavy atoms and long-range interactions.
Representation Analysis
To elucidate the improved transfer, the authors analyze the atomic embeddings produced by the different training paradigms using UMAP. Models benefitting from Hamiltonian pretraining exhibit greater cluster diversity and atomic-type resolution, especially for minority and heavy elements, compared to direct energy regression, which primarily clusters the most frequent elements (e.g., H and C). This demonstrates the utility of Hamiltonian supervision for constructing generalizable and element-discriminative atomic representations.
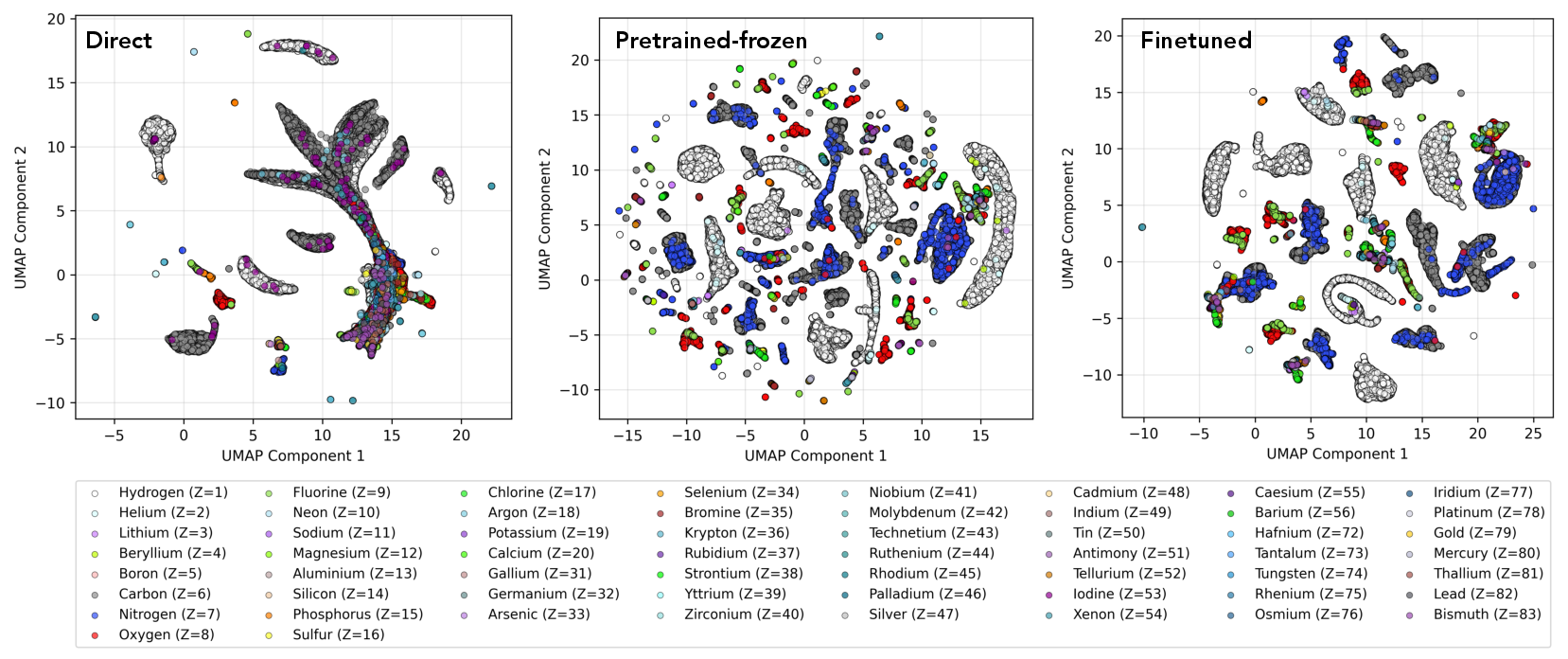
Figure 4: UMAP visualization of the HELM node embedding space following Hamiltonian pretraining, colored by element, indicating enhanced atomic-type separation and representation richness relative to direct energy training.
HELM achieves state-of-the-art accuracy on Hamiltonian prediction benchmarks (MD17, ∇2DFT, OMol_CSH_58k), outperforming recent methods including SchNOrb, PhiSNet, QHNet, and SPHNet, both in terms of global matrix element errors and energies derived from the predicted Hamiltonian. Error ablations show the critical impact of parity-constrained output mapping on diagonal matrix blocks and of per-element normalization for scalar irreps, especially for heavy elements with disparate Hamiltonian component magnitudes. The model's performance is robust with respect to dataset size, molecule size, chemical diversity, and angular momentum complexity.
Implications and Future Directions
This work demonstrates that supervised learning on electronic-structure labels offers a scalable path to improved data efficiency and transferability for MLIPs and quantum machine learning models, especially in low-data and out-of-distribution settings. The use of Hamiltonians as supervision targets provides much richer information than energies/forces, enabling the extraction of transferable physical structure from smaller datasets and facilitating generalization to novel atomic environments.
The approach also suggests promising extensions: integrating more complete quantum observables (electron densities, multipoles, response functions), pretraining on even larger datasets, application to materials and condensed matter systems, and improvement of loss functions via spectral/eigenvalue alignment and symmetry-based constraints. Further advances in model design will be required to address the computational and memory challenges posed by even larger and more complex orbital bases.
Conclusion
By unifying equivariant neural architectures with explicit electronic structure supervision, HELM and OMol_CSH_58k set a new standard for atomistic property prediction models, demonstrating that leveraging the quantum Hamiltonian enables more efficient and general representation learning across the periodic table. Hamiltonian pretraining provides significant improvements in energy prediction accuracy with orders-of-magnitude less data, and the learned backbone captures fine-grained, element-specific chemical environments. These results point to the necessity of integrating electronic-structure data as a core component in the ongoing advancement of high-fidelity atomistic machine learning.



