- The paper presents BioX-Bridge, a model bridging method that efficiently transfers knowledge across biosignal modalities using a prototype-based low-rank network.
- It achieves state-of-the-art results by reducing trainable parameters by 88–99% while outperforming baselines by 1–2% in key metrics.
- Practical evaluations on datasets like WESAD, FOG, and ISRUC demonstrate robustness, efficiency, and flexibility across different biosignal types.
BioX-Bridge: Model Bridging for Unsupervised Cross-Modal Knowledge Transfer across Biosignals
Motivation and Problem Setting
The paper addresses the challenge of unsupervised cross-modal knowledge transfer in biosignal analysis, where models trained on one modality (e.g., EEG, ECG, PPG, EMG) are leveraged to enable predictive modeling on another modality with limited or no labeled data. This is motivated by the practical constraints of biosignal acquisition: different modalities vary in sensor comfort, cost, and signal fidelity, and large labeled datasets are often unavailable for new or underrepresented modalities. Existing approaches—primarily knowledge distillation and data translation—are either computationally expensive or limited in their applicability across diverse biosignal pairs.
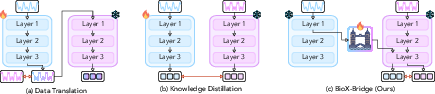
Figure 1: Comparison of unsupervised cross-modal knowledge transfer methods for biosignals. The red arrow indicates loss computation.
BioX-Bridge Framework
BioX-Bridge introduces a model bridging paradigm, constructing a lightweight bridge network that projects intermediate representations from a new modality model into the representation space of an old modality model. This enables the use of powerful, pre-trained foundation models for inference on new modalities without retraining large models or requiring extensive paired data.
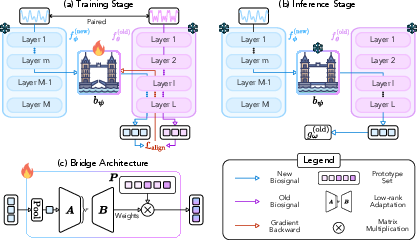
Figure 2: Overview of BioX-Bridge. (a) Training: the bridge learns to project intermediate representations from the new modality to the old modality, mimicking the output of the old modality model. (b) Inference: the bridge enables predictions on new modality data. (c) The bridge consists of a low-rank approximation module and a prototype set.
Bridge Position Selection
A critical aspect of BioX-Bridge is the selection of optimal input and output positions for the bridge within the respective models. The framework employs a two-stage strategy:
- Input Position (New Modality): Linear probing is used to identify the layer whose representations are most discriminative with respect to pseudo-labels generated by the old modality model.
- Output Position (Old Modality): Linear CKA is used to select the layer whose representations are most similar to those of the selected input layer in the new modality model.
This approach avoids brute-force search over all possible layer pairs, significantly reducing computational overhead while maximizing transfer performance.
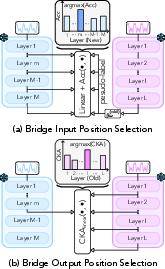
Figure 3: BioX-Bridge learning procedure.
Bridge Architecture
The bridge network is designed for parameter efficiency and flexibility. Instead of a full-rank linear projection (which would require billions of parameters for high-dimensional representations), BioX-Bridge employs a prototype network comprising:
- Prototype Set: Learnable vectors initialized from the old modality model's representations.
- Low-Rank Approximation Module: Factorized matrices generate aggregation weights for the prototypes, enabling effective high-dimensional projection with orders-of-magnitude fewer parameters.
This architecture supports both CNN and transformer-based foundation models and is compatible with arbitrary biosignal modalities.
Experimental Evaluation
BioX-Bridge is evaluated on three biosignal datasets (WESAD, FOG, ISRUC) spanning four modalities and six transfer directions. The backbone models include LaBraM (EEG), HuBERT-ECG (ECG), PaPaGei (PPG), and NormWear (EMG), all initialized with pre-trained weights.
Main Results
BioX-Bridge achieves comparable or superior performance to state-of-the-art baselines (knowledge distillation, contrastive distillation, CardioGAN) while reducing the number of trainable parameters by 88–99%. For example, in WESAD (PPG→ECG), BioX-Bridge uses only 1.3% of the parameters required by KD and outperforms it by 1–2% across balanced accuracy, F1-macro, and F1-weighted metrics.
Notably, in several cases, BioX-Bridge and KD-Contrast surpass the supervised Oracle in balanced accuracy, indicating improved recall at the expense of precision—a consequence of the transfer learning setup.
Ablation Studies
Implementation Considerations
BioX-Bridge requires only the bridge network to be trained, with all backbone models frozen. The framework is compatible with PyTorch's forward hooks for extracting and replacing intermediate representations. Hyperparameters (bridge rank, prototype set size) are selected via grid search. Training is feasible on commodity GPUs (V100, 32GB VRAM), with single runs ranging from minutes to a few hours depending on model size and dataset.
The method assumes the availability of pre-trained models for each modality and paired data for bridge training. While this is a limitation for emerging biosignals, the approach is extensible to unpaired data via future work in unsupervised domain adaptation.
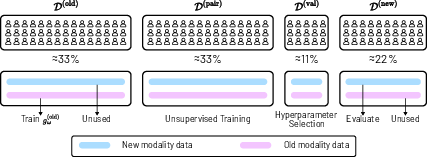
Figure 5: Illustration of Dataset Split. The dataset is divided into four subject-independent subsets for training, validation, and testing.
Theoretical and Practical Implications
BioX-Bridge demonstrates that model bridging is a viable alternative to knowledge distillation and data translation for cross-modal transfer in biosignals. The prototype-based, low-rank architecture enables efficient alignment of high-dimensional representations, facilitating interoperability between foundation models. The two-stage bridge position selection strategy is empirically validated to maximize transfer performance.
Practically, BioX-Bridge enables the deployment of biosignal models in resource-constrained environments, supporting real-world health monitoring applications where labeled data and computational resources are limited. The framework is modality-agnostic and compatible with both traditional and foundation models.
Future Directions
Potential extensions include:
- Transfer learning with unpaired data via adversarial or contrastive objectives.
- Application to emerging biosignal modalities lacking pre-trained models.
- Integration with multimodal foundation models and connector architectures for general-purpose biosignal analysis.
- Exploration of bridge architectures for other domains (e.g., vision-language, sensor fusion).
Conclusion
BioX-Bridge provides an efficient, flexible framework for unsupervised cross-modal knowledge transfer in biosignal analysis. By leveraging model bridging and prototype-based low-rank projection, it achieves strong transfer performance with minimal computational overhead. The approach is robust across modalities, tasks, and model architectures, and offers a scalable solution for real-world biosignal applications where data and resources are limited.