OpenTSLM: Time-Series Language Models for Reasoning over Multivariate Medical Text- and Time-Series Data
Abstract: LLMs have emerged as powerful tools for interpreting multimodal data. In medicine, they hold particular promise for synthesizing large volumes of clinical information into actionable insights and digital health applications. Yet, a major limitation remains their inability to handle time series. To overcome this gap, we present OpenTSLM, a family of Time Series LLMs (TSLMs) created by integrating time series as a native modality to pretrained LLMs, enabling reasoning over multiple time series of any length. We investigate two architectures for OpenTSLM. The first, OpenTSLM-SoftPrompt, models time series implicitly by concatenating learnable time series tokens with text tokens via soft prompting. Although parameter-efficient, we hypothesize that explicit time series modeling scales better and outperforms implicit approaches. We thus introduce OpenTSLM-Flamingo, which integrates time series with text via cross-attention. We benchmark both variants against baselines that treat time series as text tokens or plots, across a suite of text-time-series Chain-of-Thought (CoT) reasoning tasks. We introduce three datasets: HAR-CoT, Sleep-CoT, and ECG-QA-CoT. Across all, OpenTSLM models outperform baselines, reaching 69.9 F1 in sleep staging and 65.4 in HAR, compared to 9.05 and 52.2 for finetuned text-only models. Notably, even 1B-parameter OpenTSLM models surpass GPT-4o (15.47 and 2.95). OpenTSLM-Flamingo matches OpenTSLM-SoftPrompt in performance and outperforms on longer sequences, while maintaining stable memory requirements. By contrast, SoftPrompt grows exponentially in memory with sequence length, requiring around 110 GB compared to 40 GB VRAM when training on ECG-QA with LLaMA-3B. Expert reviews by clinicians find strong reasoning capabilities exhibited by OpenTSLMs on ECG-QA. To facilitate further research, we provide all code, datasets, and models open-source.
Paper Prompts
Sign up for free to create and run prompts on this paper using GPT-5.
Top Community Prompts
Explain it Like I'm 14
What is this paper about?
This paper introduces OpenTSLM, a new kind of AI model that can read and reason about time-series data (numbers that change over time, like heart rate every second) together with text. The goal is to help with medical tasks that depend on how things change over time—like recognizing activities from wearable sensors, figuring out sleep stages from brain waves, or answering questions about ECG heart signals—using normal, plain-language prompts and explanations.
What questions did the researchers ask?
They focused on a few simple but important questions:
- Can we teach a LLM (like ChatGPT) to understand time-series data natively, not just text or pictures?
- What’s the best way to mix time-series with text inside the model?
- Method 1: Treat time-series like extra “words” and stick them into the sentence.
- Method 2: Keep time-series separate and let the model “look at” them when needed.
- Will these methods work on real medical tasks (activity recognition, sleep staging, ECG question answering)?
- Can small, open-source models beat much larger closed models (like GPT-4o) on these time-based tasks?
- How much computer memory do these methods need, and which scales better to long signals?
How did they try to answer these questions?
They built two versions of OpenTSLM on top of well-known LLMs (LLaMA and Gemma). Think of a LLM as a very smart reader that understands and writes text.
- A quick analogy: Time-series are like a movie made of numbers over time. The model needs a way to “watch” this movie and explain what’s happening.
Here’s how the two versions differ:
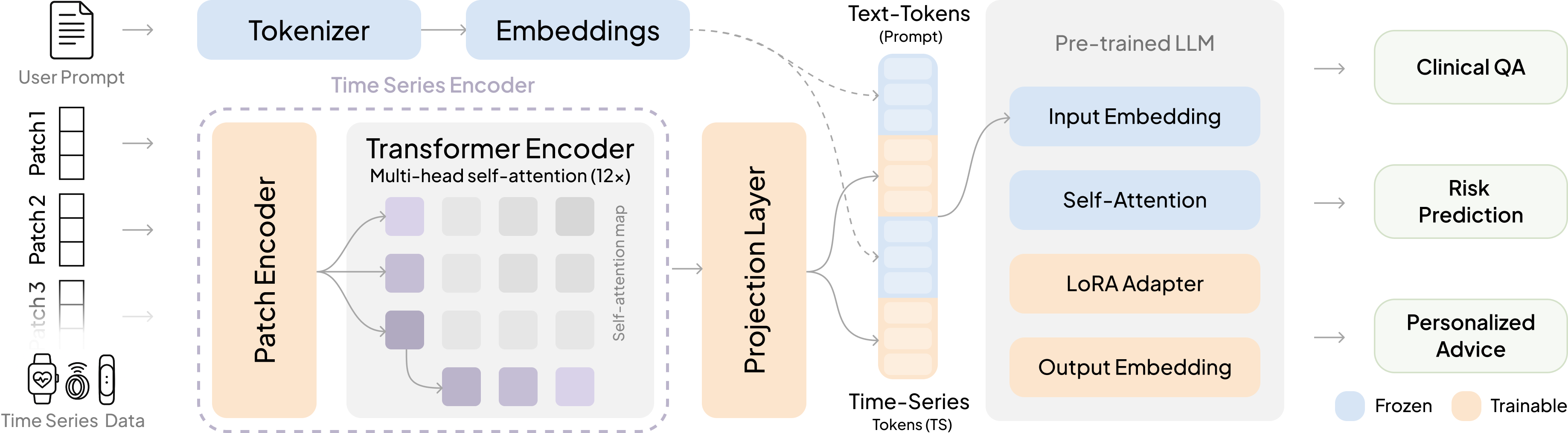
- OpenTSLM-SoftPrompt
- Idea: Turn chunks of the time-series into special “tokens” (like invented words) and insert them directly into the text the model reads.
- Analogy: Imagine adding lots of sticky notes into a book. The model reads the original text plus all these sticky notes.
- Pros: Simple and efficient for short signals.
- Cons: The longer the signal, the more sticky notes you add—this can blow up memory and slow everything down.
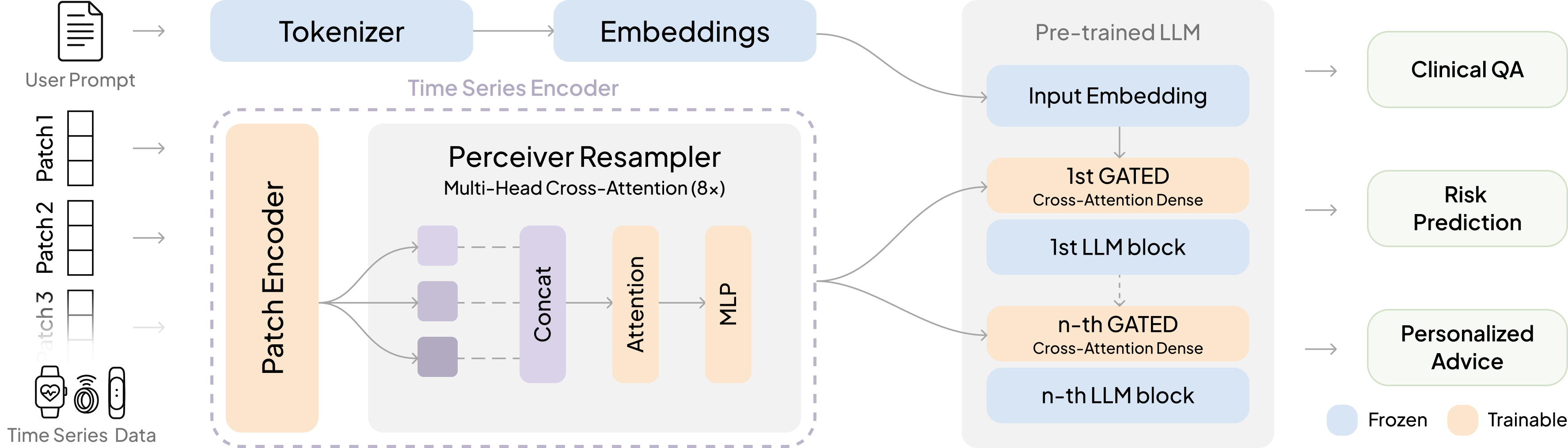
- OpenTSLM-Flamingo (cross-attention)
- Idea: Keep the time-series as a separate stream. When the text says “look at the signal now,” the model glances at that data through a special “cross-attention” mechanism.
- Analogy: The model reads a story on one screen and has the time-series on a second screen. When it sees a cue (like a special token), it looks at the second screen to gather the right details.
- Pros: Handles long signals much more smoothly; memory stays stable.
- Cons: Slightly more complex to set up.
How the numbers become model-friendly:
- The time-series are chopped into small patches (like cutting a long song into short clips) and turned into vectors (numbers the model understands).
- The model also gets a simple text note about scale and timing (e.g., “sampled at 50 Hz, mean = 61, std = 12”) so it doesn’t lose real-world meaning.
Training approach (in simple steps):
- Step 1: Practice the basics on easier, synthetic time-series tasks (like spotting rising or falling trends) and time-series captions, to “warm up” the model.
- Step 2: Train on three real medical-style datasets where the model must explain its reasoning step-by-step (called chain-of-thought) and then give a final answer:
- HAR-CoT: human activity recognition from motion sensors (e.g., walking, running).
- Sleep-CoT: sleep stage detection from EEG (brain waves).
- ECG-QA-CoT: ECG question answering with medical context.
How they checked performance:
- They compared OpenTSLM to several baselines:
- Treating time-series as plain text.
- Turning time-series into images (plots) and feeding those to an image+text model.
- Fine-tuning text-only models.
- GPT-4o (a strong proprietary model).
- They measured accuracy and F1 score (a balanced measure of how often the model is right across all classes).
- They also measured computer memory (VRAM) usage to see what scales to longer signals.
- For ECG, cardiologists reviewed the model’s explanations to judge whether its reasoning made clinical sense.
What did they find?
Big picture:
- Both OpenTSLM versions beat the baselines across tasks. Even small OpenTSLM models outperformed GPT-4o on these time-series challenges.
Key results (high-level numbers):
- Sleep staging: OpenTSLM reached about 69.9% F1, while a fine-tuned text-only baseline was around 9.05%.
- Human activity recognition: OpenTSLM reached about 65.4% F1, beating fine-tuned text-only models and GPT-4o.
- ECG-QA: OpenTSLM-Flamingo (with cross-attention) did best here, with stronger performance and more stable memory use.
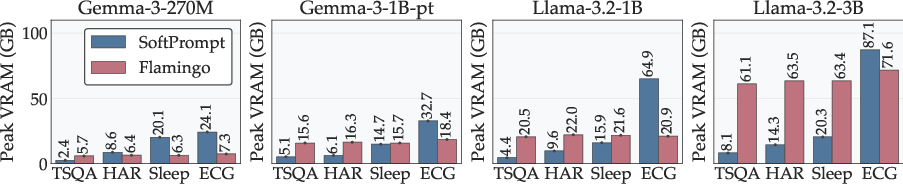
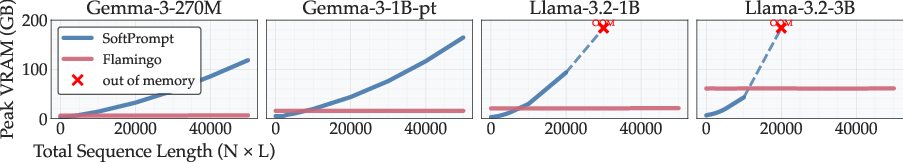
Memory and scaling:
- SoftPrompt works well for short signals but can require huge memory as signals get longer (it might jump to ~110 GB VRAM on some ECG training cases).
- Flamingo (cross-attention) stays much more stable (around ~40 GB in similar settings) and is better for long or multiple time-series.
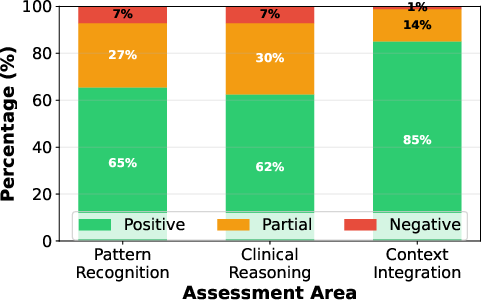
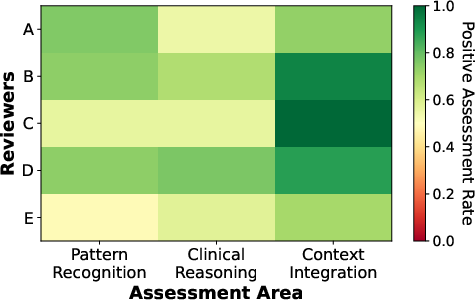
Expert review (ECG explanations):
- Cardiologists said the model’s ECG reasoning was correct or partially correct in about 92.9% of cases they checked.
- The model was especially good at using clinical context (like patient age and artifacts) to explain its answers.
Why does this matter?
- Medicine is all about changes over time—heartbeats, sleep cycles, activity patterns. A model that can “read” these signals and explain them in plain language could help doctors, nurses, and even patients understand what’s going on.
- OpenTSLM shows that small, open models (which are cheaper and can run on modest hardware) can beat much larger closed models on time-series tasks. This is promising for hospitals, research labs, and even phones or wearables.
- Because the models generate step-by-step explanations, clinicians can see how an answer was reached, which builds trust.
- The team released code, datasets, and models as open-source, so others can use and improve them.
- Beyond healthcare, the same ideas could help in finance (stock trends), factories (machine sensors), and logistics (supply chains), where time-series data is everywhere.
Simple takeaway
OpenTSLM teaches LLMs to “watch the movie” of data over time and talk about it clearly. Two ways work: stuffing the data into the text (good for short clips) or looking at it on a separate screen when needed (better for long clips). This makes powerful, explainable time-series reasoning possible—even with small, open models.
Knowledge Gaps
Knowledge gaps, limitations, and open questions
Below is a concise list of concrete gaps and open questions that remain unresolved and could guide future work:
- External validity: No evaluation on out-of-distribution generalization across institutions, devices, or populations (e.g., train on SleepEDF, test on another sleep dataset; train on one HAR corpus, test on a different one; external ECG cohorts).
- Irregular, missing, and asynchronous sampling: The encoder assumes uniform patches and normalized inputs; robustness to irregular sampling, missingness, sensor dropout, and asynchronous multi-sensor streams is not assessed.
- Multi-channel fusion at scale: How best to jointly encode and fuse many synchronized channels/leads (e.g., 12‑lead ECG, multi-sensor wearables) is unclear; no ablations on cross-sensor interactions, channel attention, or shared-vs-separate encoders.
- Extremely long horizons and streaming: Support for day-to-week scale signals, streaming/online inference, and retrieval over very long histories is not demonstrated (only up to 10,000 steps in simulation).
- Inference-time efficiency: Memory and latency are reported for training but not for inference; edge-device feasibility (CPU-only, quantized, mobile NPUs), throughput under multiple concurrent series, and energy use are unreported.
- Safety and calibration: No analysis of calibration, selective prediction, uncertainty estimation, or risk-aware thresholds for clinical deployment.
- Robustness: No stress tests for noise, motion artifacts, sampling-rate mismatches, unit errors, adversarial perturbations, or out-of-domain signals.
- Faithfulness of rationales: While clinicians rated many ECG rationales positively, faithfulness is not tested (e.g., feature occlusion/ablation, counterfactuals, input perturbations) to verify that cited features causally drive predictions rather than being post-hoc.
- CoT supervision efficacy: No comparison to training without CoT, with weak/concise rationales, or with alternative supervision (e.g., step labels, evidence tags); impact of CoT style/length/quality is unknown.
- Synthetic/pseudo-labeled pretraining: Stage-1 relies on synthetic TSQA and ChatGPT-generated M4 captions; the effect of this curriculum and its potential biases are not ablated against curated, expert-verified pretraining or self-supervised time-series pretraining.
- Baseline coverage: Missing comparisons with strong time-series foundation and 1D CNN/Transformer baselines (e.g., InceptionTime, TimesNet/TimesFM, TST, TS2Vec, ROCKET/miniROCKET) and recent multimodal TS-text models under matched training data.
- Task breadth: Focus is on multiple-choice classification/QA; regression, forecasting, anomaly/event detection, segmentation, and time-to-event analyses are not evaluated.
- Multi-turn and interactive use: The ability to handle follow-up queries, refer to prior time windows, and perform dialogue-based exploration of long signals is not assessed.
- Modality fusion beyond time series: Integration with structured EHR (labs, meds, diagnoses), medical images, audio (stethoscope), or video is not explored.
- Scale/unit handling: Encoding mean/std/time scale as text is brittle; learned unit/scale embeddings, per-sensor calibration layers, and automatic unit validation are not investigated.
- Alignment and chunking: The impact of prompt formatting, special tokens (<TS>, <endofchunk>), ordering of text/series chunks, and potential misalignment between descriptions and series is not quantified.
- Architectural ablations: No ablations on patch size, encoder depth, Perceiver latent size, cross-attention frequency/placement, gating parameter γ, LoRA ranks (for SoftPrompt), or projection MLP design.
- Training strategy choices: Consequences of freezing vs partially/fully fine-tuning the LLM, and trade-offs in parameter-efficient tuning strategies, are not explored.
- Scaling laws: Relationships between backbone size, time-series encoder capacity, dataset size/quality, and performance (and compute/memory) are not characterized.
- Multi-language generalization: Rationales and prompts are in English; performance and clinical terminology coverage in other languages are unknown.
- Data contamination: No audit ensures that pretraining corpora or GPT-4o-generated content did not leak task-specific rationales/templates; potential data leakage remains unaddressed.
- ECG-QA scope: Comparison templates were removed; how the model handles comparison/temporal-difference ECG questions or free-form ECG interpretations remains open.
- Clinician study limitations: Small sample (84 items), variable reviewer stringency, no inter-rater reliability statistics (e.g., Cohen’s/Fleiss’ kappa), and no error-severity stratification; scaling the evaluation is needed.
- Answer correctness enforcement: The paper notes that generative training does not guarantee correct final labels; constrained decoding, answer-verification losses, or discriminative auxiliary heads are not explored.
- Fairness and subgroup performance: No breakdown by demographics (age, sex, race), device type, or clinical subgroups; potential biases and fairness constraints are unexamined.
- Continual/online learning: Handling distribution shift and concept drift in longitudinal monitoring (e.g., disease progression, sensor upgrades) is not studied.
- Explainability beyond text: Lack of temporal saliency/attribution (e.g., patch/lead importance) tied to rationales; no evaluation of alignment between highlighted segments and clinical features.
- Data preprocessing sensitivity: No analysis of sensitivity to window length, overlap, filtering, resampling, or normalization choices.
- Data privacy: No treatment of differential privacy, federated learning, or on-device training for sensitive medical time series.
- Multi-series relational reasoning: Although multiple <TS> chunks are supported, explicit reasoning over relationships across simultaneous series (e.g., ECG vs PPG timing, actigraphy vs HRV) is not benchmarked.
- Error analysis: No detailed failure taxonomy (e.g., specific ECG patterns missed, sleep-stage confusions), nor guidance for targeted improvements.
- Clinical endpoints: Metrics are macro-F1/accuracy; clinically meaningful endpoints (e.g., sensitivity for dangerous arrhythmias, alarm fatigue, time-to-diagnosis) are not reported.
- Reproducibility of GPT-4o baselines: Prompt design and sensitivity analyses for GPT-4o and image-based baselines are limited; potential underestimation of strong proprietary baselines remains an open question.
Glossary
- Autoregressive language modeling: A training setup where the model predicts the next token given previous tokens, enabling sequence generation. "we frame the task as an autoregressive language modeling problem."
- Chain-of-Thought (CoT): A prompting and training paradigm where models generate intermediate reasoning steps before final answers. "We introduce three time-series Chain-of-Thought (CoT) datasets: HAR-CoT (human activity recognition), Sleep-CoT (sleep staging), and ECG-QA-CoT (ECG question answering)."
- Contrastive loss: A learning objective that pulls matched pairs together and pushes mismatched pairs apart in representation space. "apply cross-attention between a time series encoder and a text encoder, aligned with contrastive loss, to extract statistical summaries (e.g., mean, max) from a single sensor."
- Cross-attention: An attention mechanism that conditions one sequence (e.g., text) on another (e.g., time series) by attending over the other sequence’s representations. "We thus introduce OpenTSLM-Flamingo, which integrates time series with text via cross-attention."
- Curriculum learning: A training strategy that presents tasks or data in a meaningful order from easier to harder to improve learning efficiency. "Multi-Stage Curriculum Learning – Teaching LLMs Time Series"
- Electronic health record (EHR): Digitized patient health information systems used for clinical data storage and retrieval. "As time-series data from electronic health records and continuous monitoring proliferate"
- Electrocardiogram (ECG): A cardiac diagnostic signal measuring the heart’s electrical activity over time. "which provides 12-lead 10s ECGs and clinical context"
- Electroencephalogram (EEG): A neurological signal measuring brain electrical activity over time. "using 30s EEG segments for sleep staging."
- Few-shot prompting: Guiding an LLM with a handful of examples in the prompt to perform a new task without full fine-tuning. "to enable LLMs to infer clinical and wellness information through few-shot prompting."
- Flamingo: A multimodal architecture that fuses non-text modalities with text via gated cross-attention in a frozen LLM backbone. "using a cross-attention mechanism inspired by Flamingo~\cite{flamingo} to fuse time-series and text."
- Gated cross-attention: Cross-attention augmented with a learnable gating parameter to modulate how much one modality influences another. "we add gated cross-attention layers every N (hyperparameter) transformer blocks"
- Human Activity Recognition (HAR): Automated classification of human physical activities from sensor signals (e.g., accelerometers). "HAR-CoT (human activity recognition)"
- LoRA (low-rank adaptation): A parameter-efficient fine-tuning method that inserts low-rank trainable adapters into large models. "a pretrained LLM, fine-tuned using LoRA adapters~\cite{lora}"
- Multi-layer perceptron (MLP): A feed-forward neural network consisting of multiple linear layers with nonlinear activations. "and subsequently project the resulting tokens with an MLP to align them with the embedding space"
- Perceiver Resampler: A module that converts variable-length inputs into a fixed-size latent representation via cross-attention-style layers. "We use a PerceiverResampler inspired by Flamingo~\cite{open_flamingo} as Encoder for the time series patches, yielding a fixed-size latent representation:"
- Positional encoding: Additive signals injected into embeddings to represent token positions and preserve order information. "and added with a positional encoding~\cite{a_time_series_is_64_words}"
- Projection head: A task-specific mapping (often a small neural network) from shared embeddings to outputs such as class logits. "for classification via a projection head; however this disables free-form text generation."
- Self-attention: A mechanism that lets each token attend to all other tokens in the same sequence, with quadratic compute in sequence length. "increasing context length and compute due to the quadratic cost of self-attention"
- Sleep staging: Classification of sleep into standard stages (e.g., Wake, REM, NREM) from physiological signals. "using 30s EEG segments for sleep staging."
- Soft prompting: Conditioning an LLM by prepending or interleaving learned continuous embeddings (prompts) instead of only discrete tokens. "concatenating learnable time series tokens with text tokens via soft prompting."
- Time Series Question Answering (TSQA): A task/dataset for answering questions about temporal patterns in time series. "We present performance on the test splits of TSQA, HAR–CoT, Sleep–CoT, and ECG–QA–CoT"
- Time-Series LLM (TSLM): An LLM extended to natively process and reason over time-series inputs alongside text. "we present OpenTSLM, a family of TSLMs created by integrating time series as a native modality to pretrained LLMs"
- Tokenization (of time series as text): Converting numeric time-series values into sequences of text tokens for LLM input. "we tokenize time series into text inputs and report zero-shot performance on the test set."
- Transformer encoder: The stack of self-attention and feed-forward layers used to encode sequences before downstream processing. "We apply the patch embeddings to a transformer encoder"
- VRAM (video random access memory): GPU memory used to store model parameters, activations, and intermediate tensors during training/inference. "requiring ~110 GB compared to ~40 GB VRAM when training on ECG-QA with LLaMA-3B."
Collections
Sign up for free to add this paper to one or more collections.