- The paper presents EnzyControl, a novel framework that achieves a 13% improvement in native enzyme design using guided generative modeling.
- It employs a two-stage training strategy with a pretrained base network and EnzyAdapter to inject substrate information via cross-modal projection.
- Benchmarking on the EnzyBind dataset shows enhanced catalytic efficiency and substrate specificity, with zero-shot generalization on unseen enzymes.
EnzyControl: Adding Functional and Substrate-Specific Control for Enzyme Backbone Generation
EnzyControl is an innovative framework designed to address limitations in current computational protein engineering techniques by allowing for functional and substrate-specific control in enzyme backbone generation. It is a promising approach for generating enzyme structures customized for specific interactions with substrates, integrating functional site conservation and substrate-awareness into the generative process.
Introduction
Designing enzyme backbones that exhibit substrate-specific functionality is a critical challenge due to the stringent requirements for substrate binding, functional site preservation, and sensitive catalytic conformations. Traditional protein design methods are inadequate for enzyme design because they often neglect these aspects. EnzyControl leverages a curated dataset, EnzyBind, containing 11,100 enzyme-substrate pairs, focusing on utilizing multiple sequence alignments (MSA) for functional site annotation and incorporating substrate information via a modular component called EnzyAdapter.
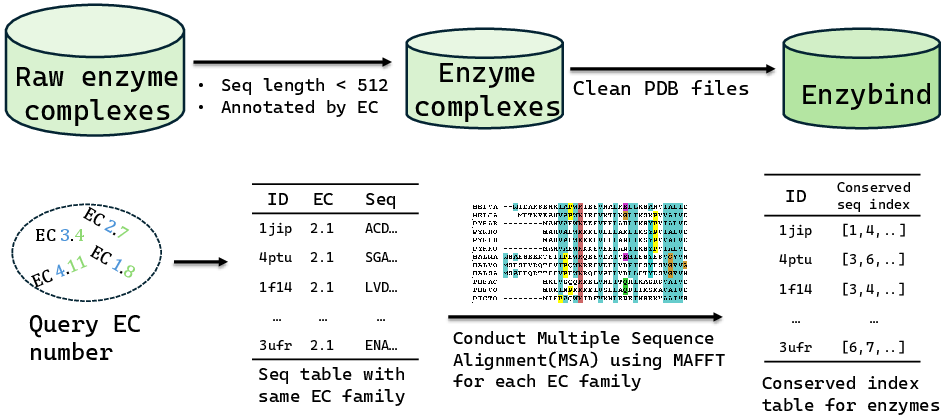
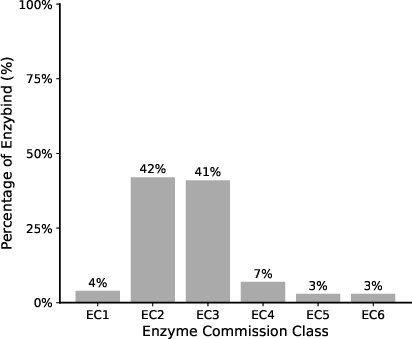
Figure 1: Dataset collection and preprocessing.
Methodology
EnzyControl Architecture
EnzyControl consists of three key components:
- Base Network: Pretrained for motif-scaffolding, integrating functional site conservation using MSA.
- EnzyAdapter: A modular addition that injects substrate information into the network, employing a cross-modal projector to bridge the modality gap between substrates and enzyme backbones.
- Two-Stage Training Strategy: Initial training aligns substrate features with enzyme structures without altering base network parameters, followed by fine-tuning using Low-Rank Adaptation (LoRA) methods.
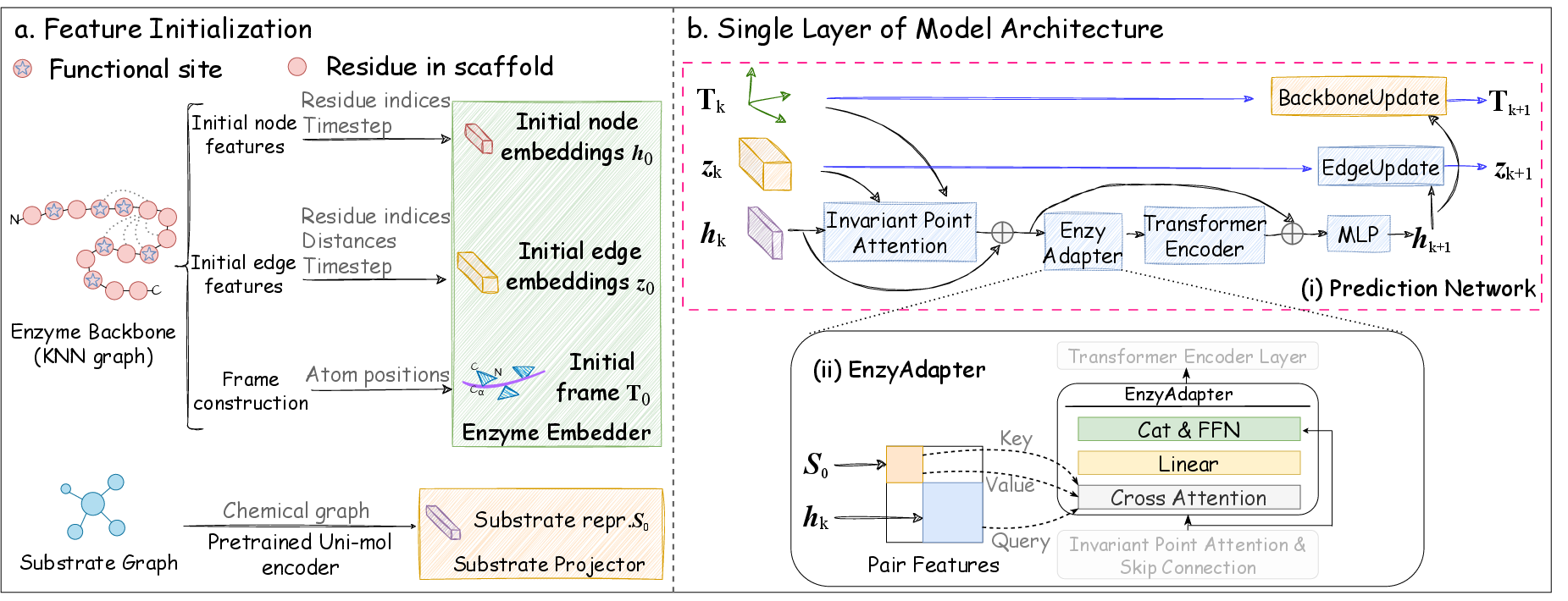
Figure 2: EnzyControl is a flexible approach for the conditional backbone generation of enzymes.
Flow Matching
EnzyControl utilizes Flow Matching (FM), a generative modeling technique, which provides efficient and stable sampling processes. This technique estimates vector fields describing the evolution between data and noise distributions for generating enzyme backbones. FM formulates the generative task as solving an Ordinary Differential Equation (ODE) enabling backward sampling from noise.
Experimental Results
EnzyControl was benchmarked on the EnzyBind dataset across multiple structural and functional metrics. Significant findings include:
- Designability: Achieving a 13% improvement over baseline models, signifying enhanced alignment to native enzyme structures.
- Functional Performance: Improvements of 13% in catalytic efficiency ($k_{\text{cat}$) and 10% in EC match rates, demonstrating strong functionality preservation during backbone generation.
- Substrate Affinity: Enhanced binding affinity and substrate specificity scores compared to other models.
EnzyControl also exhibits zero-shot generalization capabilities, maintaining strong binding affinities on previously unseen substrates and enzyme categories.
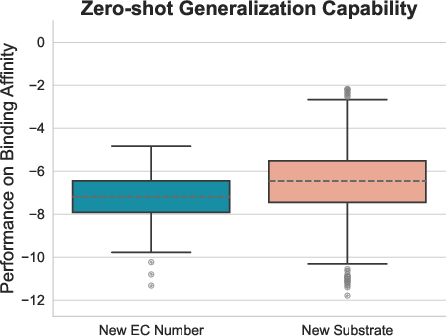
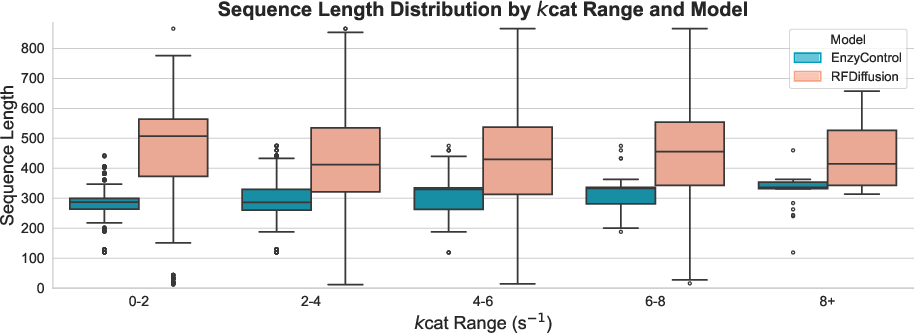
Figure 3: Zero-shot generalization.
Case Study
A targeted case study on enzyme 2cv3 demonstrated that EnzyControl-generated backbones achieve better substrate-specificity and interaction characteristics than existing models like RFDiffusion.
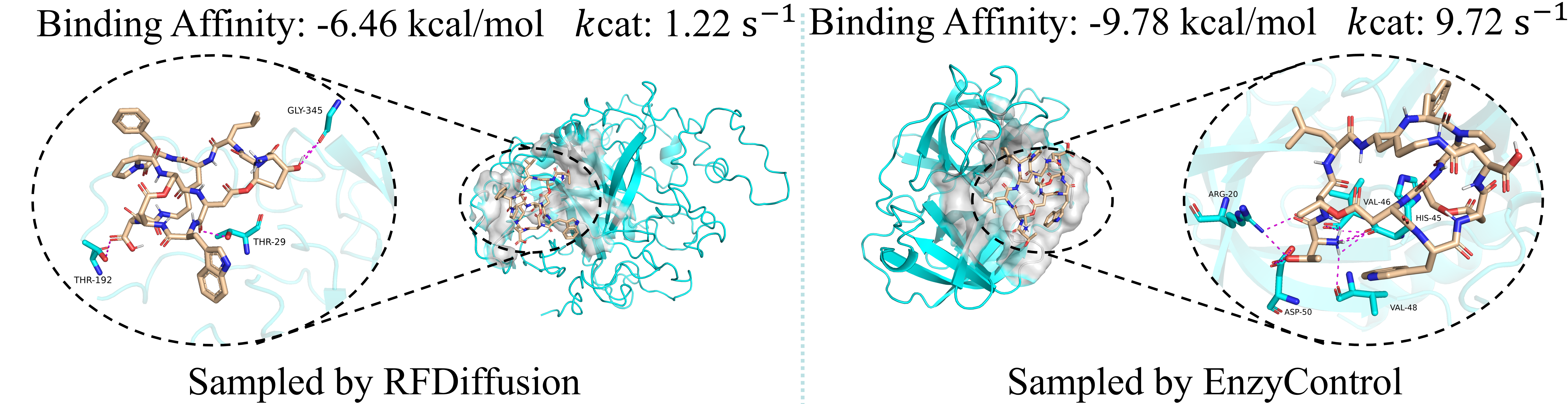
Figure 4: Comparison of docking results between EnzyControl and RFDiffusion on the 2cv3 enzyme.
Conclusion
EnzyControl effectively integrates functional site conservation and substrate-specific control, pioneering a nuanced approach to enzyme design that extends current motif-scaffolding models. The framework not only advances structural accuracy but also enhances functional relevance in enzyme backbone generation, offering a robust tool for computational protein engineering and potential practical applications in pharmaceuticals, specialty chemicals, and biotechnology. Future directions include refining substrate-conformation modeling and expanding capabilities for multi-substrate or multi-chain enzyme systems.







