- The paper demonstrates the impact of U-Net architecture choices and loss functions on overcoming class imbalance in AMD lesion segmentation.
- It leverages weighted BCE and Tversky loss to optimize both pixel-wise segmentation and image-level detection across diverse lesion types.
- It establishes new benchmarks on the ADAM challenge dataset, paving the way for improved automated AMD screening in clinical settings.
Surpassing State of the Art on AMD Area Estimation from RGB Fundus Images: U-Net Architectures and Loss Functions for Class Imbalance
Introduction
This work addresses the problem of automated age-related macular degeneration (AMD) lesion segmentation in RGB fundus images, a clinically significant task due to the prevalence and impact of AMD on vision loss in the elderly population. The study leverages the ADAM challenge dataset as a benchmark and systematically investigates the impact of U-Net architectural choices and loss functions tailored for class imbalance. The primary contribution is a segmentation framework that achieves superior performance compared to all prior ADAM challenge submissions, with a focus on both pixel-wise segmentation and image-level lesion detection.
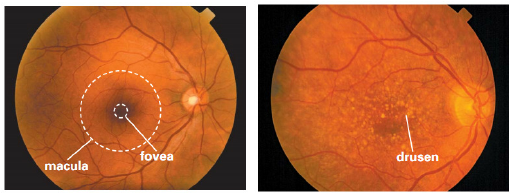
Figure 1: A healthy retina (left) and an AMD-affected retina with yellow drusen (right).
Background and Motivation
AMD is characterized by the presence of lesions such as drusen, exudates, hemorrhages, scars, and other abnormalities in the macula, as visualized in RGB fundus images. Early detection is critical, but manual assessment is labor-intensive and subject to inter-observer variability. Automated segmentation methods, particularly those based on deep learning, offer a scalable solution. However, the task is complicated by severe class imbalance: the majority of images lack lesions, and lesion pixels are sparse even within positive cases.
The ADAM challenge dataset provides a comprehensive benchmark, with 1200 high-quality RGB fundus images and pixel-wise annotations for five lesion types. The evaluation metric is a weighted combination of the Dice coefficient (segmentation) and F1 score (detection), emphasizing the clinical importance of both accurate localization and reliable detection.
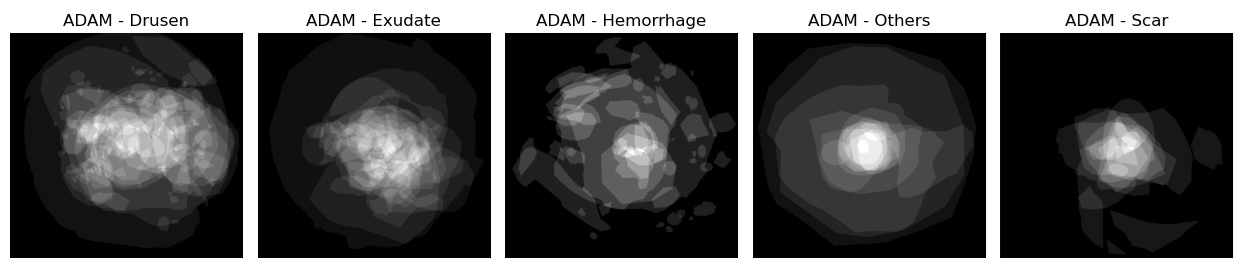
Figure 2: Average of the masks over different types of lesions for the ADAM dataset, illustrating the sparsity and variability of lesion distributions.
Methodology
Data Preprocessing and Augmentation
The pipeline constructs a multi-channel binary mask for each image, with each channel corresponding to a specific lesion type. Images and masks are resized to 320×320 to balance computational efficiency and segmentation accuracy. Data augmentation includes rotations, cropping, scaling, and photometric adjustments, applied identically to images and masks to preserve spatial correspondence.

Figure 3: Example of an input image and its corresponding multi-channel ground truth mask.
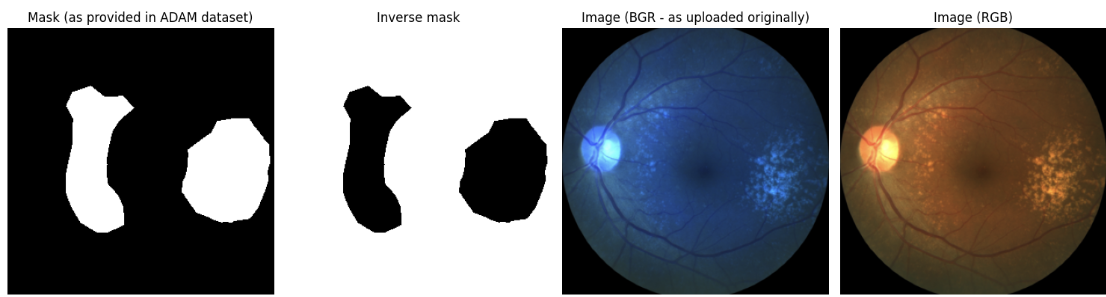
Figure 4: Original and preprocessed input image and ground truth mask after data loading and transformation.
Model Architecture
The core architecture is a U-Net with an encoder-decoder structure and skip connections, implemented using the Segmentation Models PyTorch library. The encoder is based on EfficientNet (B0 or B2), initialized with ImageNet-pretrained weights via the timm library, which provides superior pretraining compared to standard implementations. Each lesion type is modeled independently, with a dedicated U-Net trained per class.
Loss Functions for Class Imbalance
Given the extreme foreground-background imbalance, several loss functions are evaluated:
- Weighted Binary Cross-Entropy (BCE): Class weights are computed as the ratio of negative to positive pixels per lesion type, directly addressing imbalance.
- Dice Loss: Optimizes overlap but is unstable for sparse targets and can lead to oversegmentation.
- Tversky Loss: Generalizes Dice by introducing tunable penalties for false positives and false negatives.
- Focal Loss: Downweights easy examples, focusing on hard-to-classify pixels, with tunable focusing parameter γ.
Parameter tuning for Tversky and Focal losses is performed per lesion type to optimize performance.
Training and Evaluation
Each model is trained for 100 epochs with Adam optimizer and learning rate scheduling. The best model is selected based on validation Rank (weighted Dice and F1). Evaluation includes both quantitative metrics and qualitative inspection of predicted masks.

Figure 5: Input image, ground truth mask, model output, and binarized segmentation mask (threshold = 0.5).
Experimental Results
Encoder Selection
Empirical results demonstrate that the choice of encoder and pretraining strategy significantly impacts performance. EfficientNetB2 (timm) achieves the highest overall Rank, with deeper encoders favoring classification (F1) but not always improving segmentation (Dice). The timm library's advanced pretraining yields consistent gains over standard implementations.
Loss Function Analysis
Weighted BCE and Tversky loss provide the most balanced performance across segmentation and detection metrics, as evidenced by the highest Rank scores. Dice loss, while effective for segmentation, leads to excessive false positives in negative images, degrading classification. Focal loss excels in detection but underperforms in pixel-wise segmentation due to the nature of the imbalance.
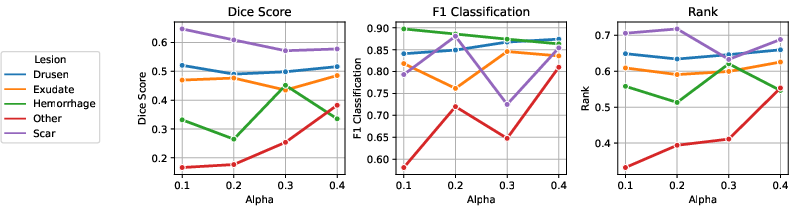
Figure 6: Results of parameter tuning for Tversky loss; α values control the trade-off between false positives and false negatives.
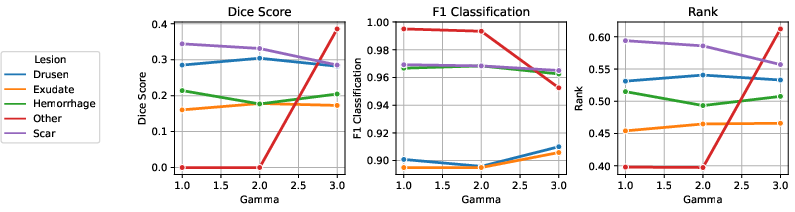
Figure 7: Results of parameter tuning for Focal loss; γ values modulate the focus on hard examples.
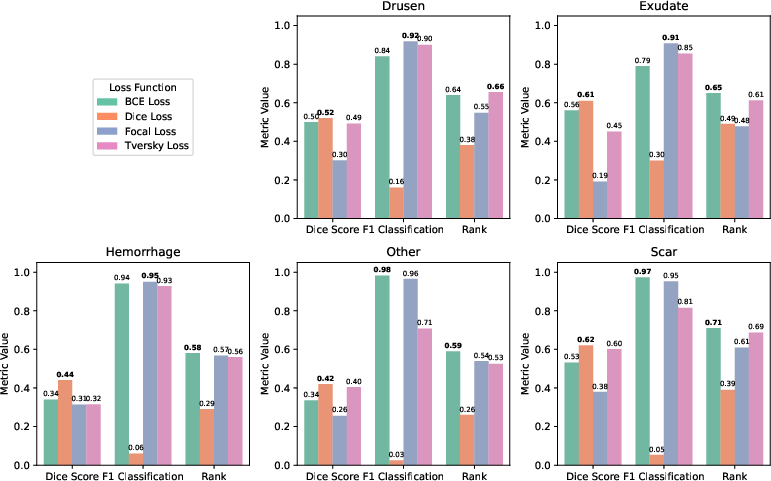
Figure 8: Comparative results for different loss functions across lesion types, highlighting the trade-offs between segmentation and detection.
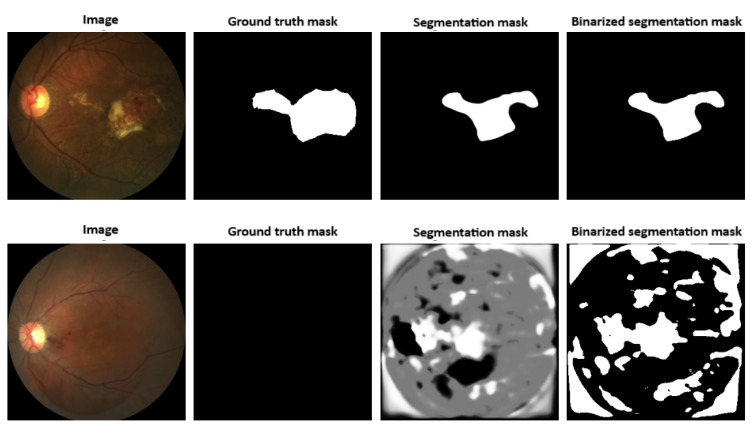
Figure 9: Example outputs from a model trained with Dice loss, illustrating oversegmentation and false positives in negative images.
Final Configuration and Benchmarking
The final framework employs a U-Net with EfficientNetB2 (timm) encoder, weighted BCE loss, and extensive data augmentation. This configuration achieves the highest or second-highest Dice and F1 scores for all lesion types compared to ADAM challenge submissions, and the highest overall Rank. Notably, the method demonstrates robust detection (F1) even for rare lesion types where prior methods failed, and maintains balanced performance across all classes.
Discussion
The study provides a rigorous analysis of architectural and loss function choices for semantic segmentation under severe class imbalance. The results underscore the importance of advanced encoder pretraining and loss functions that directly address imbalance at both the image and pixel levels. Weighted BCE offers a practical and effective solution, requiring no parameter tuning and yielding consistent results. The findings also highlight the limitations of Dice and Focal losses in joint segmentation-classification tasks, particularly in the presence of negative images.
From a practical perspective, the proposed framework is well-suited for deployment in clinical settings, offering reliable detection and localization of AMD lesions from non-invasive RGB fundus images. The modular design allows for extension to other retinal pathologies or imaging modalities.
Implications and Future Directions
The demonstrated improvements in both segmentation and detection metrics have direct implications for automated screening and monitoring of AMD. The methodology is generalizable to other medical image segmentation tasks characterized by class imbalance and sparse targets. Future work should focus on further enhancing boundary precision, possibly via boundary-aware losses or post-processing, and exploring multi-task learning to jointly model all lesion types. Integration with clinical decision support systems and prospective validation in real-world settings are also warranted.
Conclusion
This work establishes a new state of the art for AMD lesion segmentation in RGB fundus images, surpassing all prior ADAM challenge results through careful selection of U-Net architectures and loss functions tailored for class imbalance. The findings provide actionable insights for the design of robust medical image segmentation pipelines and contribute to the broader field of automated ophthalmic diagnosis. The open-source codebase facilitates reproducibility and further research.








