- The paper introduces a BN-LTE framework that leverages latent pseudotime to dynamically order patients and enhance causal inference in Alzheimer’s disease.
- It demonstrates that pseudotime outperforms chronological age, achieving an AUC of 0.82 versus 0.59 for AD status prediction.
- The study reveals both established and novel causal edges between biomarkers, suggesting new opportunities for therapeutic stratification.
Dynamic Causal Discovery in Alzheimer's Disease via Latent Pseudotime Modeling
Introduction and Motivation
Alzheimer's disease (AD) research is fundamentally impeded by the complexity and heterogeneity of its progression, with attempts to model causality in AD hampered by a reliance on static, time-invariant causal graphs. The presented work applies Bayesian Networks with Latent Time Embedding (BN-LTE) to infer a pseudotime as a latent variable, dynamically ordering patients along a data-driven disease trajectory. Unlike chronological age, pseudotime serves as a surrogate for the unobserved progression and underlying modulators of AD, thereby enabling the modeling of dynamic causal interactions. This method leverages both established AD markers and emerging plasma biomarkers—specifically neurofilament light (NfL) and glial fibrillary acidic protein (GFAP)—to characterize the evolving interdependencies as disease advances.
Methodological Framework
The analytic pipeline incorporates real-world data from the ADNI dataset, encompassing demographic variables, region-specific volumetric measures, plasma biomarkers, and cognitive scores. BN-LTE, as proposed in Zhou et al. (2023), provides a mechanism to order cross-sectional samples (patients) along a continuous pseudotime, reflecting disease state rather than chronological time. Causal relationships between variables are expressed as time-dependent functionals, parameterized by cubic b-splines. Model parameters are estimated via MCMC, with convergence diagnostics verifying posterior stability.
Background knowledge integration is implemented agnostically to disease specifics, constraining only immutable demographic factors (sex, genotype) as root nodes and cognitive scores as sinks. This mitigates orientation bias and improves identifiability, with causal sufficiency and faithfulness assumed in the presence of pseudotime.
Empirical Results
Pseudotime Versus Age in Disease Prediction
Pseudotime demonstrates greater discriminability in predicting AD status (AUC = 0.82, 95% CI: 0.81–0.82) compared to age alone (AUC = 0.59, p≪0.01). Patients are arranged by increasing disease severity along the pseudotime axis, with AD phenotypes clustering at late pseudotime, MCI in the midrange, and healthy controls at early stages.
Causal Graph Recovery and Edge Dynamics
Causal edges with high posterior inclusion probability (PIP) replicate both canonical and novel AD pathways. The BN-LTE model, especially when augmented with background knowledge, robustly recovers literature-supported links such as NfL → hippocampal volume decrease and pTau → NfL elevation. The addition of background constraints markedly improves edge detection precision (from 0.80 to 0.88) and orientation accuracy (from 0.62 to 0.96).
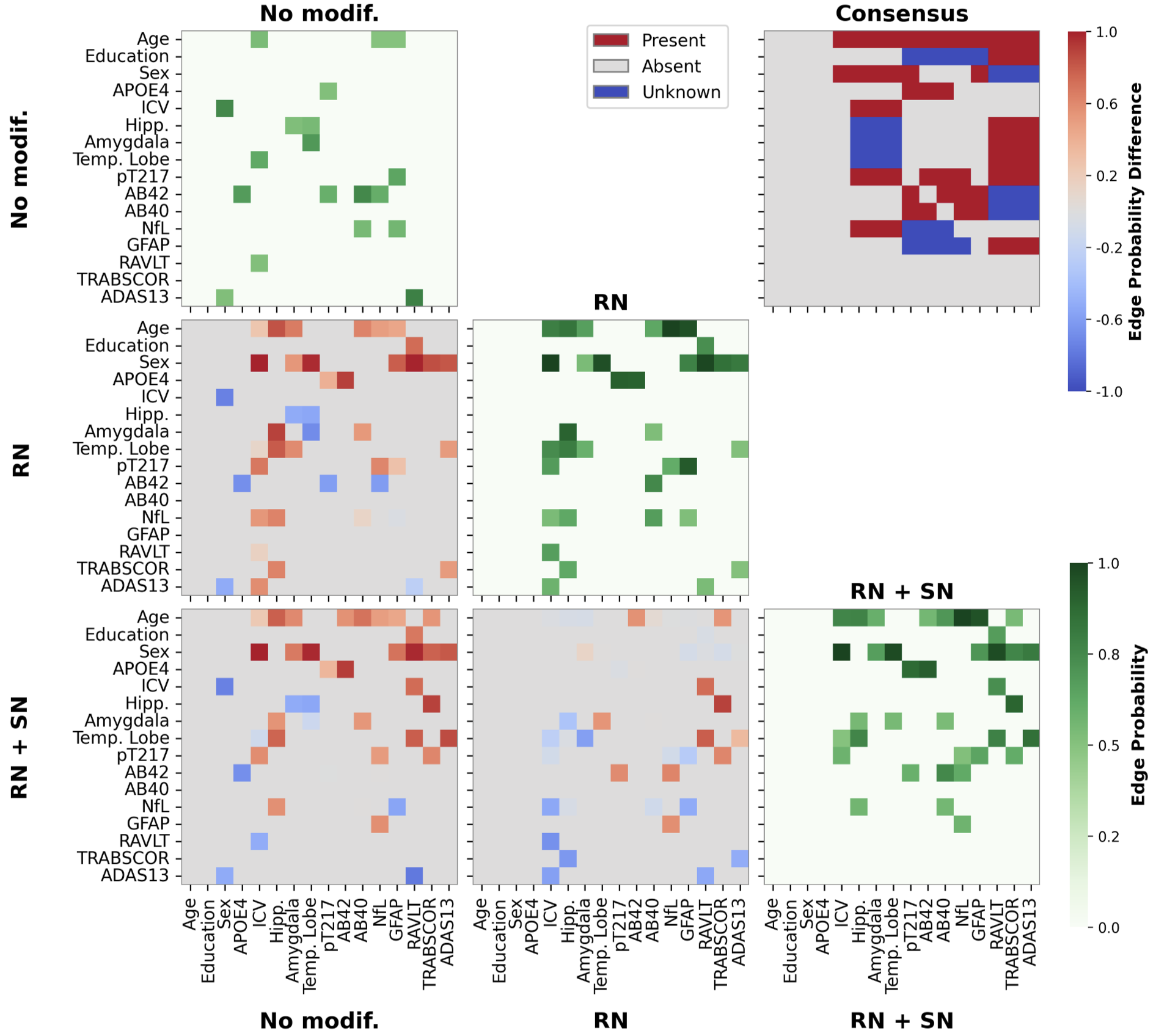
Figure 1: Matrices of edge inclusion probabilities highlight the improvement in graph recovery under background knowledge constraints, with comparisons to the literature consensus graph.
Contradictory and Novel Findings
Certain identified edges challenge established pathophysiological models—e.g., pTau → GFAP and NfL → A$\upbeta$40—contradict the theorized sequence where amyloid pathology precedes neurodegeneration and glial activation. While these conflicting causal inferences necessitate cautious interpretation, they may also signal limitations of cross-sectional data or model assumptions, such as the exclusion of feedback cycles.
Dynamic causal effects manifest along pseudotime; the effect of pTau on NfL is restricted to early disease, consistent with anticipated biological progression, while the impact of age on GFAP remains temporally invariant. These findings underscore the necessity of modeling time-evolving, non-stationary causality in AD.
Methodological Implications
The operationalization of latent pseudotime as a continuous modulator provides several key improvements over conventional approaches:
- Enhanced Causal Identifiability: Modeling time-evolving graphs circumvents violations arising from cross-sectional sampling heterogeneity and unobserved confounders.
- Translational Potential: Stratification by disease stage unveils new therapeutic windows and rationalizes combination therapy sequencing.
- Robustness to Violated Assumptions: Background knowledge incorporation serves as a practical workaround for cyclical or confounded graph structures that would otherwise undermine inference validity.
The work highlights that inclusion of emerging plasma biomarkers alongside established features captures both classical and previously undocumented causal interactions. Computational implementation via MCMC and cubic-spline parameterization achieves tractable inference on consumer hardware (24GB RAM, 30 minutes for 4 chains × 5000 steps), enabling scalability to moderately sized datasets. However, the absence of explicit unobserved confounder modeling and feedback cycle representation restricts completeness.
Limitations and Future Directions
The approach is constrained by sample size, variable selection, and dataset heterogeneity, limiting generalizability across ethnic and demographic strata. Causal graph reconstruction accuracy reduces in graph subsets lacking background constraints, suggesting orientation bias remains a challenge when agnostic background specification is infeasible. Methodological advances, such as multi-dimensional pseudotime (vector-valued latent trajectories) and cross-cohort joint causal inference, are advocated to address AD's extensive heterogeneity.
Further, relaxing DAG constraints to incorporate cyclic or latent confounded relationships may align causal inference with the intricate feedback present in AD biology. Longitudinal data acquisition would substantively validate dynamic causal paths revealed by pseudotime modeling, providing temporal anchoring to inferred relationships.
Conclusion
This study demonstrates that dynamic causal modeling of Alzheimer's disease via latent pseudotime markedly improves the identification and orientation of causal edges compared to traditional age-based stratification or static DAGs. Integrating minimal background knowledge proves critical for causal graph accuracy under real-world violations of model assumptions. The framework is directly extendable to clinical trial stratification, biomarker-driven diagnosis, and ongoing mechanistic research. Future work should broaden the latent modulator space, expand cohorts, and refine model assumptions for deeper insight into AD heterogeneity and progression.
