- The paper introduces a LoRA-driven framework that decomposes models into a frozen backbone and low-rank adapters to achieve targeted unlearning.
- The methodology employs a dual-phase strategy combining gradient ascent for forgetting and supervised losses for data retention to manage privacy.
- Experimental results reveal significant, localized performance drops on forgotten subsets while preserving overall segmentation accuracy in clinical datasets.
Low-Rank Adaptation Guided Selective Unlearning in Medical Segmentation Networks
Introduction and Motivation
Erase to Retain (2511.16574) presents a systematic methodology for selective knowledge removal in medical segmentation networks, driven by the increasing necessity for privacy compliance, ethical AI deployment, and continual revision of clinical datasets. The requirement to "forget" training data is accelerated by regulations such as GDPR, which mandate mechanisms for retrospective deletion. In medical image segmentation, the stakes of secure unlearning are heightened: precise spatial representations entangle sensitive patient features across pixel-level regions, and naive approaches risk inefficiency or inadvertent leakage.
This work addresses core limitations of previous methods—sample-level inversion, influence-based reweighting, and knowledge decomposition—which are primarily designed for classification or NLP and do not address dense, structured representations in medical images. The proposed approach decomposes the model into a frozen backbone and low-rank (LoRA) adapters, wherein selective unlearning is localized, computationally efficient, and does not globally distort anatomical priors essential for clinical deployment.
Methodological Framework
The pipeline comprises four main stages—teacher model inference, LoRA-based student adaptation, retain-guided knowledge distillation, and background-focused forgetting—as visualized in Figure 1.
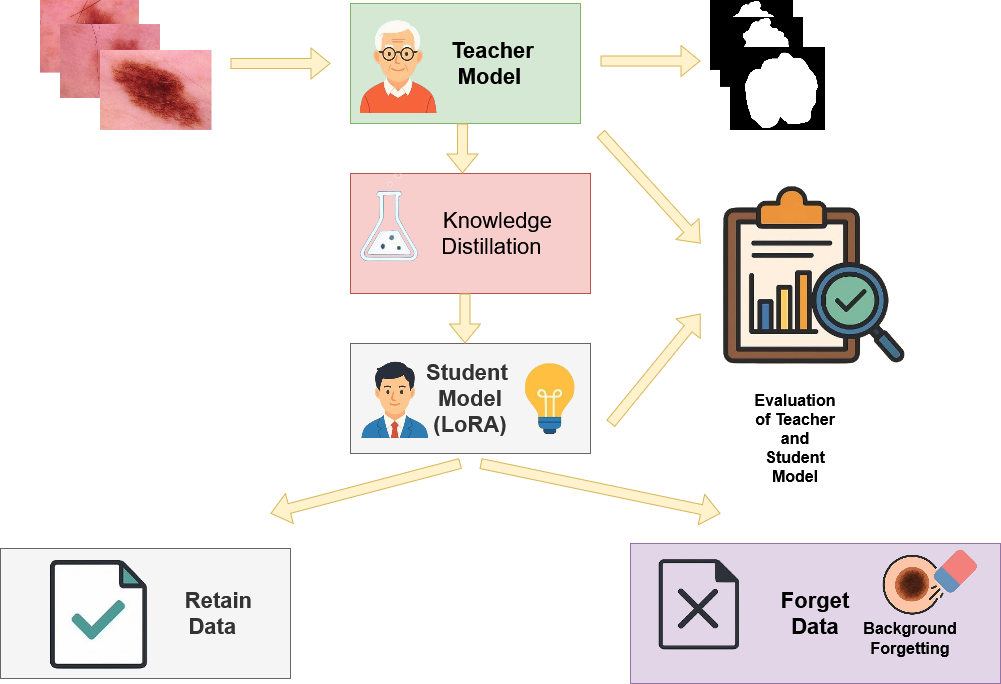
Figure 1: Overview of the proposed unlearning methodology, illustrating the modular teacher-student distillation pipeline with LoRA-driven parameter updates and targeted suppression on the forget subset.
The LoRA adapters act as subspace modulators, constraining updates to low-dimensional projections that enable reversible and fine-grained forgetting. The theoretical construct bifurcates learning objectives:
- Ascent (Forget) Objective: Gradient ascent is performed on the forget set using composite losses—label-flip (forcing complement predictions), teacher contradiction (driving anti-alignment), entropy maximization (enforcing high uncertainty), feature-space repulsion, mean-probability regularization, and total variation regularization. These combine to maximize divergence between student and teacher (∣∣ps(x)−pt(x)∣∣1>δ) for x∈Df, restricting the student's ability to recall lesion evidence.
- Descent (Retain) Objective: On the retain set, supervised Dice/BCE loss is combined with KD and guard regularizers. LoRA's restricted update manifold (≈3% trainable parameters) limits catastrophic drift and preserves the teacher’s boundary on Dr.
This dual-phase strategy mimics a two-player adversarial game—forgetting is maximized only for designated samples while retention and generalization are stably preserved.
Experimental Results
Model Efficiency
LoRA adaptation modifies only 2.93% of overall model parameters, yielding a highly efficient, easily reversible framework. Full retraining is avoided, and base anatomical representations remain largely intact for non-forgotten regions.
On the ISIC 2018 skin lesion benchmark, the teacher exhibits uniform segmentation performance (Dice ≈ 0.93). After targeted unlearning, the student model's Dice on the forget subset falls sharply by nearly 30 points (to 0.63), with retain and validation sets showing moderate, controlled declines (16 and 12 points, respectively). This demonstrates pronounced, deliberate forgetting restricted to specified samples.
Qualitative analysis in Figure 2 displays row-wise input, GT mask, teacher and student predictions, and their absolute difference. In retained images, predictions remain congruent and lesion foreground probabilities are preserved. Conversely, on forgotten samples, the student explicitly suppresses lesion probability, as reflected by concentrated difference maps.
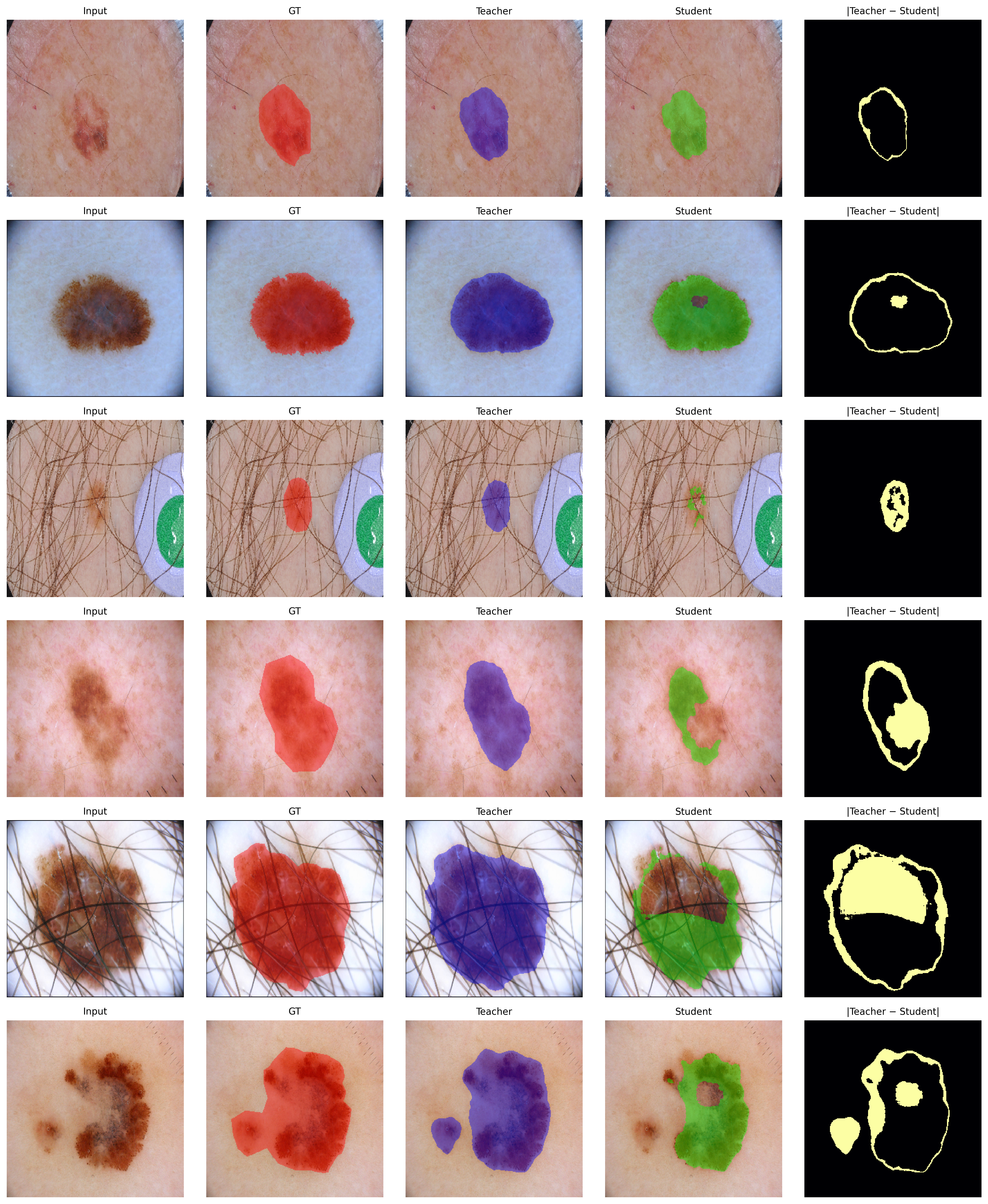
Figure 2: Qualitative visualization of unlearning on ISIC: compared mappings for retain and forget samples, highlighting selective suppression and spatial fidelity across foreground structures.
CHASE_DB1 Retinal Vessel Segmentation
In cross-domain settings (CHASE_DB1), the pattern of sharp, localized forgetting persists. For the 1st Human Observer mask, the student’s Dice on the forget set drops by 19.3 points; retention and validation subsets remain near teacher scores (see Section Results). The trend is confirmed using alternative ground truth (2nd Human Observer mask), showing robustness to annotation variability.
Cross-Task Generalization: ISIC Classification
Applying the same framework to ISIC lesion classification, the teacher's accuracy on the forget subset drops from 87.0% to 64.1%, while retention accuracy improves from 83.9% to 90.6%. Furthermore, generalization to unseen test data remains competitive (student 55.9% vs. teacher 61.9%), indicating effective containment of forgetting without global performance collapse.
Classwise analysis in Figure 3 highlights deliberate accuracy degradation on forgotten classes and stability elsewhere.
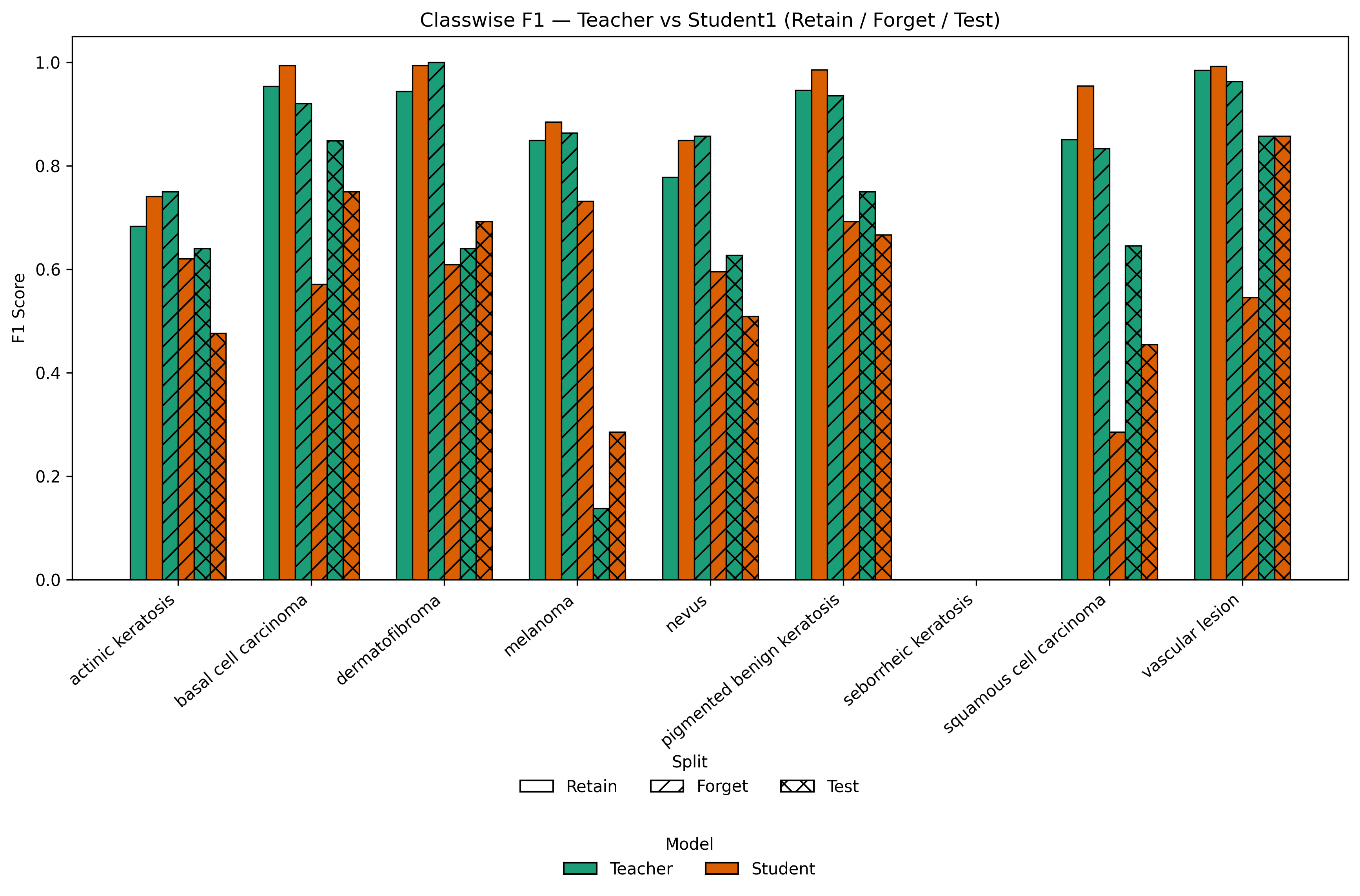
Figure 3: Classwise F1 comparison for Teacher vs LoRA-based Student across Retain, Forget, and Test splits, revealing selective knowledge suppression with persistent generalization.
Implications and Future Directions
From a practical perspective, Erase to Retain provides a feasible pathway for privacy-preserving medical AI, enabling highly controllable knowledge deletion post hoc, with minimal risk to clinically relevant global functionality. The retention-forget trade-off is rendered explicit via low-rank adaptation, and the methodology is computationally scalable across tasks (segmentation, classification) and domains.
Theoretically, the approach delineates boundaries—both spectral and empirical—on catastrophic forgetting, deriving guarantees from the constrained LoRA update subspace. The dual-phase adversarial optimization presents new vistas for certified unlearning, foundational medical models, and multi-modal deletion for clinical decision support.
Speculatively, future work could explore extensions to 3D volumetric data, federated unlearning across distributed institutions, and integration with certified influence-based unlearning for structural guarantees of data deletion. The work also motivates generalization of parameter-efficient selective unlearning in other dense prediction domains, including remote sensing, histopathology, and multi-organ segmentation.
Conclusion
Erase to Retain (2511.16574) delivers a rigorously constructed, LoRA-driven framework for selective unlearning in medical segmentation and classification models. By leveraging low-rank updates and adversarial objectives, it enables removal of patient- or class-specific knowledge with constrained collateral impact, rapid training, and reversible adaptation. The strong empirical results on ISIC and CHASE demonstrate effectiveness, stability, and generalizability, marking a significant step toward practical, responsible unlearning solutions in sensitive clinical AI deployments.


