- The paper introduces a universal two-dimensional phase-plane framework to model protein aggregation and clearance in living cells.
- It employs bifurcation analysis to reveal bistable states driven by aggregate removal saturation, reflecting transitions from healthy to pathological conditions.
- The findings offer actionable insights for therapeutic strategies targeting neurodegenerative diseases through modulation of protein homeostasis.
A Universal Phase-Plane Model for In Vivo Protein Aggregation
Introduction
Protein aggregation is a critical factor in the progression of neurodegenerative diseases such as Alzheimer's and Parkinson's. While in vitro studies have extensively characterized the kinetics of protein aggregation, these models often fall short of explaining the dynamics within living organisms, where active cellular processes such as aggregate removal come into play. In their paper titled "A Universal Phase-Plane Model for In Vivo Protein Aggregation" (2511.18893), the authors propose a theoretical framework utilizing a two-dimensional phase-plane representation to describe the interplay between aggregate formation and removal in cellular environments. This model aims to elucidate the conditions under which cells transition from a stable state dominated by monomeric proteins to a pathological state rife with aggregates.
In Vitro Aggregation Models
The foundational aspect of the paper is the interplay of aggregation mechanisms known from in vitro observations, which include primary nucleation, secondary nucleation, and fragmentation. These processes collectively drive the formation of aggregates from monomeric proteins. Previous studies have demonstrated that these in vitro models reliably predict aggregation rates across various protein types. However, the authors argue that living systems impose additional constraints, particularly related to active removal processes that metastably manage aggregate concentrations.
Extension to In Vivo Environments
In vivo systems differ fundamentally from in vitro environments due to the active processes involving both monomer production and aggregate clearance. The paper introduces kinetics models where cellular homeostasis maintains monomer concentrations over aggregation dynamics but also highlights removal mechanisms like autophagy that contest aggregate buildup. A critical innovation in the authors' model is the inclusion of removal kinetics that saturate beyond a certain aggregate load, forming bistable cellular states—a healthy state with low aggregate concentration and a pathological state characterized by runaway aggregation.
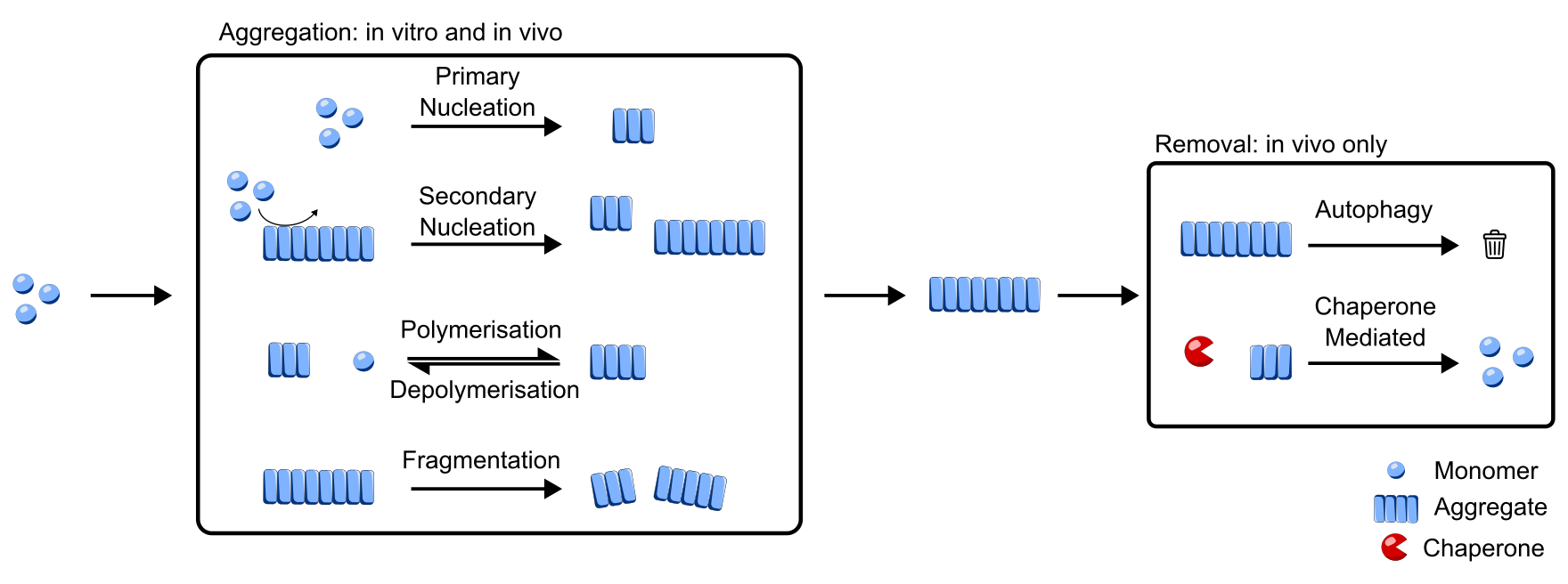
Figure 1: Multiple reactions can convert monomeric protein into an aggregated state. Additionally, in living cells aggregates are removed from cells by active processes.
Bistability and Phase-Plane Analysis
The paper rigorously examines systems where aggregate clearance efficiency declines as aggregate concentration escalates, employing a bifurcation analysis within the proposed phase-plane framework. The findings suggest conditions under which systems exhibit bistability—states where cells toggle between low aggregate loads (health) and high aggregate concentrations (disease). This bistable behavior occurs due to mechanisms like saturation kinetics in aggregate removal processes, exemplified by enzyme kinetics models for clearance.
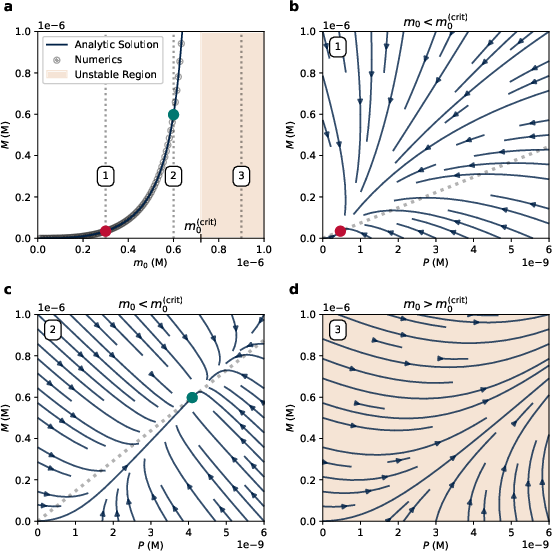
Figure 2: Exact steady states and dynamics for the unbound removal model. (a) The steady state aggregate mass diverges as the monomer concentration approaches the critical value.
Broader Implications and Future Directions
The broader implications of this universal modeling framework provide profound insights into therapeutic strategies for neurodegenerative diseases. By understanding the phase transitions between stable and unstable cellular states, the model can guide the development of interventions aimed at altering monomer and aggregate concentrations. Future research may expand upon these models to integrate more complex biological interactions and clinically test predictions regarding the efficacy of aggregate-reducing therapies, such as those employing monoclonal antibodies or antisense technologies.
Conclusion
The universal phase-plane model delineated in this paper represents a significant advancement in understanding protein aggregation within living systems, extending the analytical toolkit used to study in vitro aggregation processes. By integrating aggregate formation and removal dynamics, the researchers offer a robust framework for predicting cellular behavior and transitions into degenerative states. This theoretical approach promises to inform ongoing therapeutic development and enhance our fundamental understanding of neurodegenerative disease mechanisms.
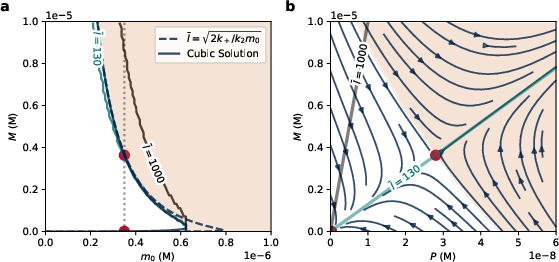
Figure 3: Exact steady states and dynamics for the MM-like removal model. (a) Steady state values of M are plotted for the full cubic system.


