- The paper introduces a robust ABC-SMC method using PyMC to perform likelihood-free inference in dynamical models.
- It validates the approach through case studies like Lotka–Volterra and SEIR, emphasizing accurate parameter recovery and uncertainty quantification.
- It offers practical guidance on diagnostics, adaptive tolerances, and hierarchical modeling to enhance inference in partially observed systems.
Practical Approximate Bayesian Computation with ABC-SMC Using PyMC: A Technical Overview
Introduction and Context
The increasing complexity and realism of mechanistic models in ecological and epidemiological research have amplified the demand for robust statistical inference methods capable of handling intractable likelihoods. The reviewed work, "Approximate Bayesian Computation Made Easy: A Practical Guide to ABC-SMC for Dynamical Systems with pymc" (2511.21587), systematically addresses this challenge by offering an accessible and reproducible approach to Approximate Bayesian Computation with Sequential Monte Carlo (ABC-SMC), leveraging modern probabilistic programming through PyMC. The tutorial-driven exposition systematically escalates case complexity, highlighting critical methodological, practical, and diagnostic facets of likelihood-free inference in dynamical systems governed by ODEs and stochastic processes.
ABC-SMC: Methodological Framework
Approximate Bayesian Computation circumvents the explicit evaluation of likelihood functions, instrumental for systems where analytic or computational evaluation is infeasible due to latent variables, stochastic generative models, or high-dimensional outputs. The ABC framework substitutes likelihood with a simulation-comparison paradigm: parameter vectors are sampled from the prior and accepted if simulated data are sufficiently similar to observed data under a defined distance metric and tolerance threshold.
The Sequential Monte Carlo variant of ABC (ABC-SMC) introduces a population-based inference scheme with adaptive tolerances, ensuring efficient exploration and progressive refinement of parameter distributions across intermediate distributions. This yields substantial efficiency gains compared to rejection ABC, particularly relevant for models of moderate to high dimensionality with pronounced complexity.
PyMC Integration and Implementation
A core contribution of the paper is the demonstration of PyMC's pm.Simulator interface, which seamlessly integrates user-defined simulators into the Bayesian workflow. This unifies likelihood-based and likelihood-free paradigms, facilitating implementation, diagnostics, and visualization within a single, extensible framework. The paper details the implementation of ABC-SMC across four instructive case studies, each leveraging the concise API and high-level abstractions of PyMC.
Case Study Analyses
Lotka–Volterra with Full Observability
ABC-SMC is first applied to parameter inference in the classical predator-prey Lotka–Volterra system, under the setting where both species are observed across time. The results feature tight unimodal marginal posteriors for growth and predation rates, accurately centered on ground-truth parameters. The 2D joint posterior reveals negative correlation, illustrating compensatory trade-offs between growth and predation in maintaining oscillatory dynamics.
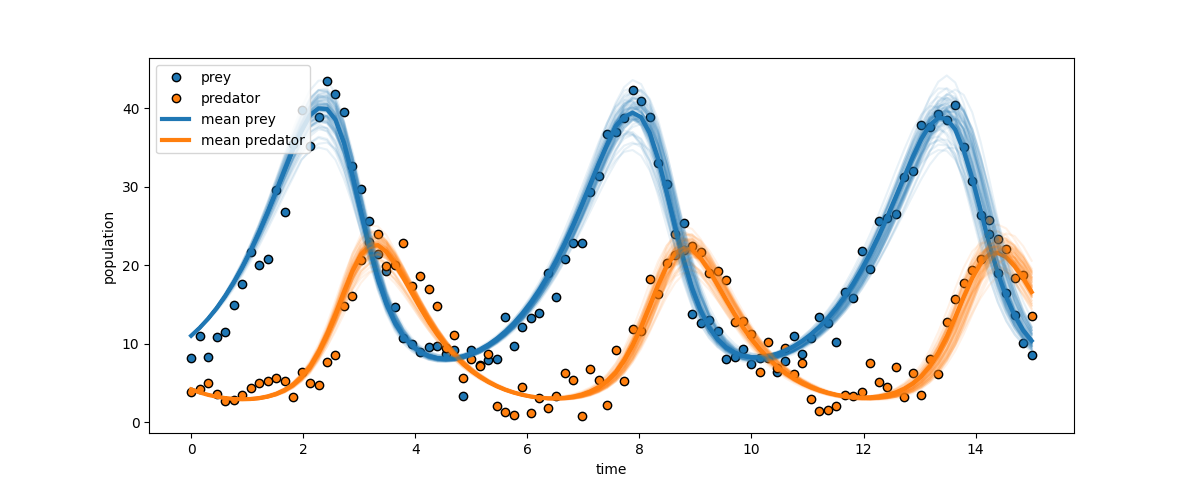
Figure 2: The close alignment between observed predator-prey trajectories and posterior predictive simulations demonstrates accurate parameter recovery and effective uncertainty quantification.
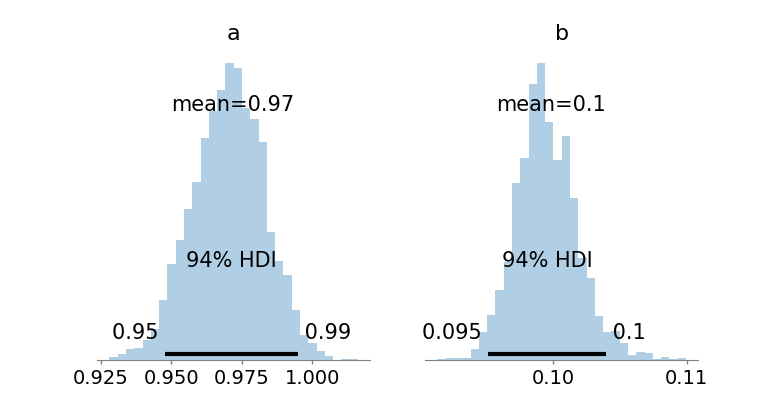
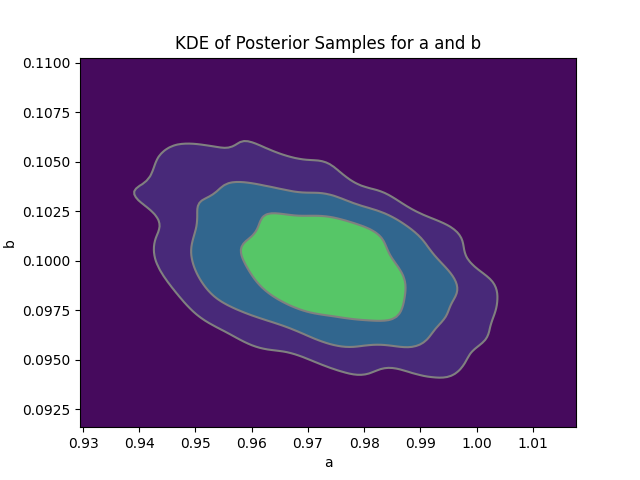
Figure 4: Posterior densities for a (prey growth) and b (predation) are highly concentrated near true values, with the joint posterior contour reflecting parameter compensability.
Lotka–Volterra with Partial Observability
A reduction in observability—where only the prey population is measured—results in a notable expansion of uncertainty regarding predator dynamics. Although prey predictions remain tightly constrained, posterior uncertainty for latent predator trajectories increases, and parameter identifiability is reduced. Nevertheless, the method maintains dynamic consistency due to the mechanistic model structure.
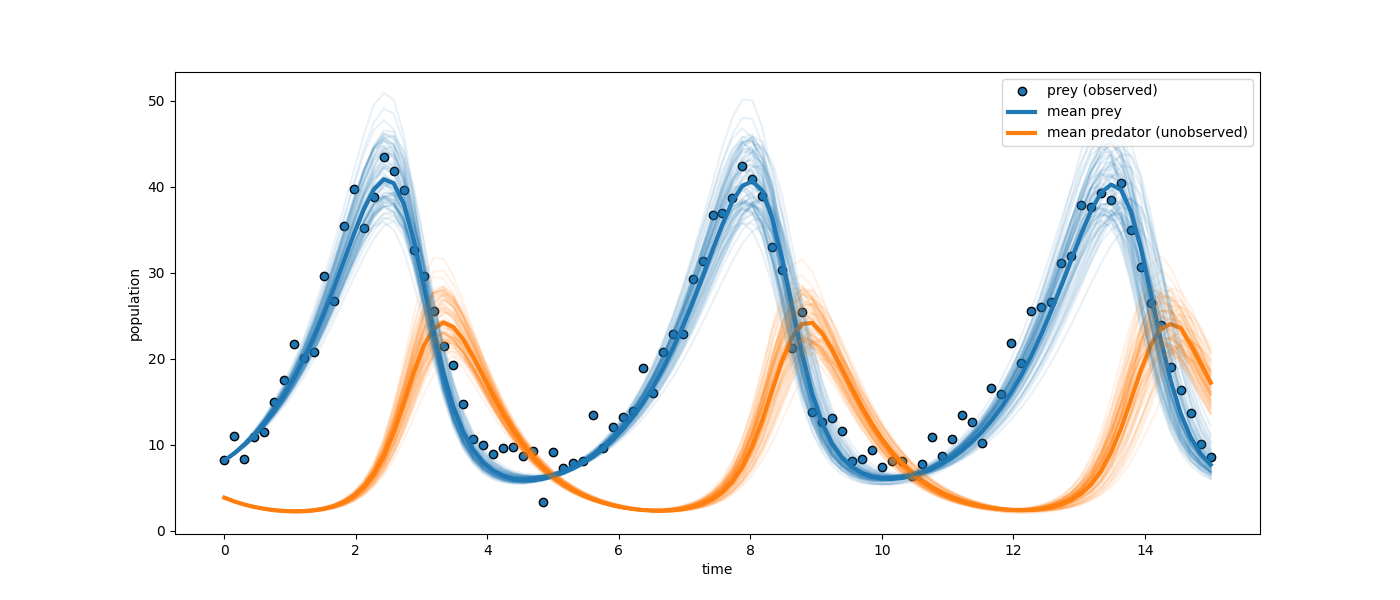
Figure 1: Prey dynamics (observed and reconstructed in blue) display much lower uncertainty than latent predator trajectories (orange), illustrating information loss from partial observability.
SEIR Model with Partially Observed Compartments
The SEIR compartmental model is considered next, reflective of epidemic processes with unmeasured latent variables (exposed but not yet infectious individuals). ABC-SMC recovers posterior estimates for transmission, incubation, and recovery rates that are well concentrated, and accurately reconstructs the unobserved exposed dynamics solely through their interaction with observed susceptible, infected, and recovered compartments.
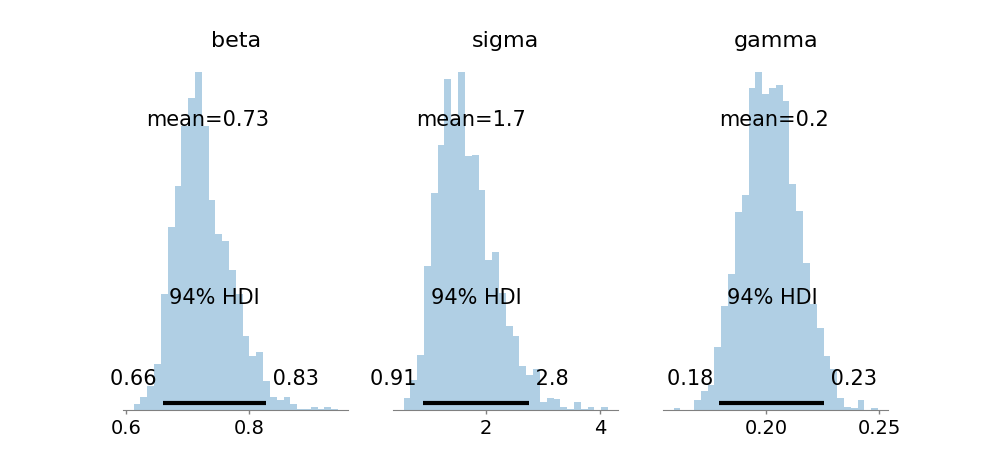
Figure 3: Marginal posteriors for β, σ, and γ in SEIR demonstrate effective parameter identifiability from incomplete observations.
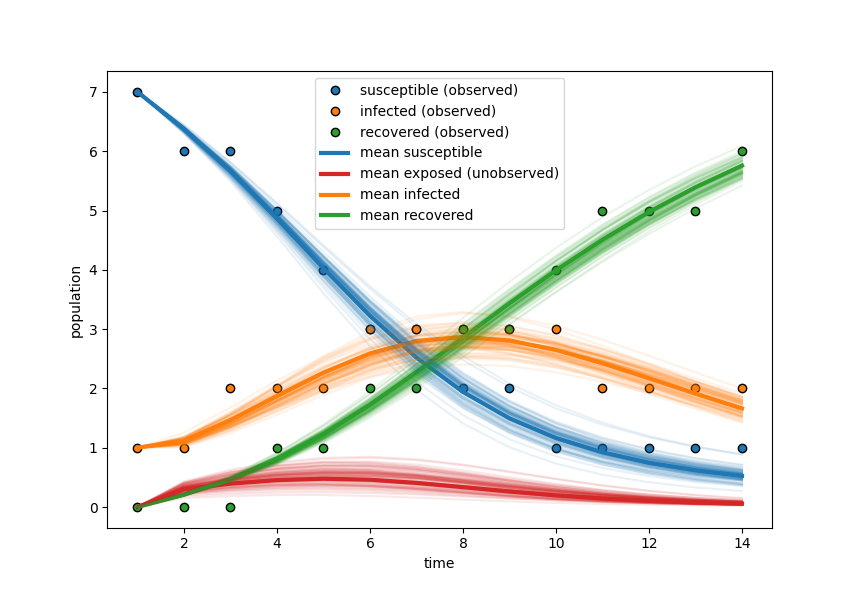
Figure 8: Posterior predictive simulations capture the dynamics of all compartments, reconstructing even the unobserved exposed variable, with uncertainty reflecting latent status.
Hierarchical SEIR Across Subpopulations
The framework is extended to hierarchical models, where subpopulation-specific (grouped) SEIR parameters are drawn from common hyperdistributions, enabling partial pooling. Compared to independent inference, this results in broader posterior spreads for group-specific parameters but quantifies both average effects and heterogeneity. Hyperparameter posteriors elucidate between-group variability and pooling strength.
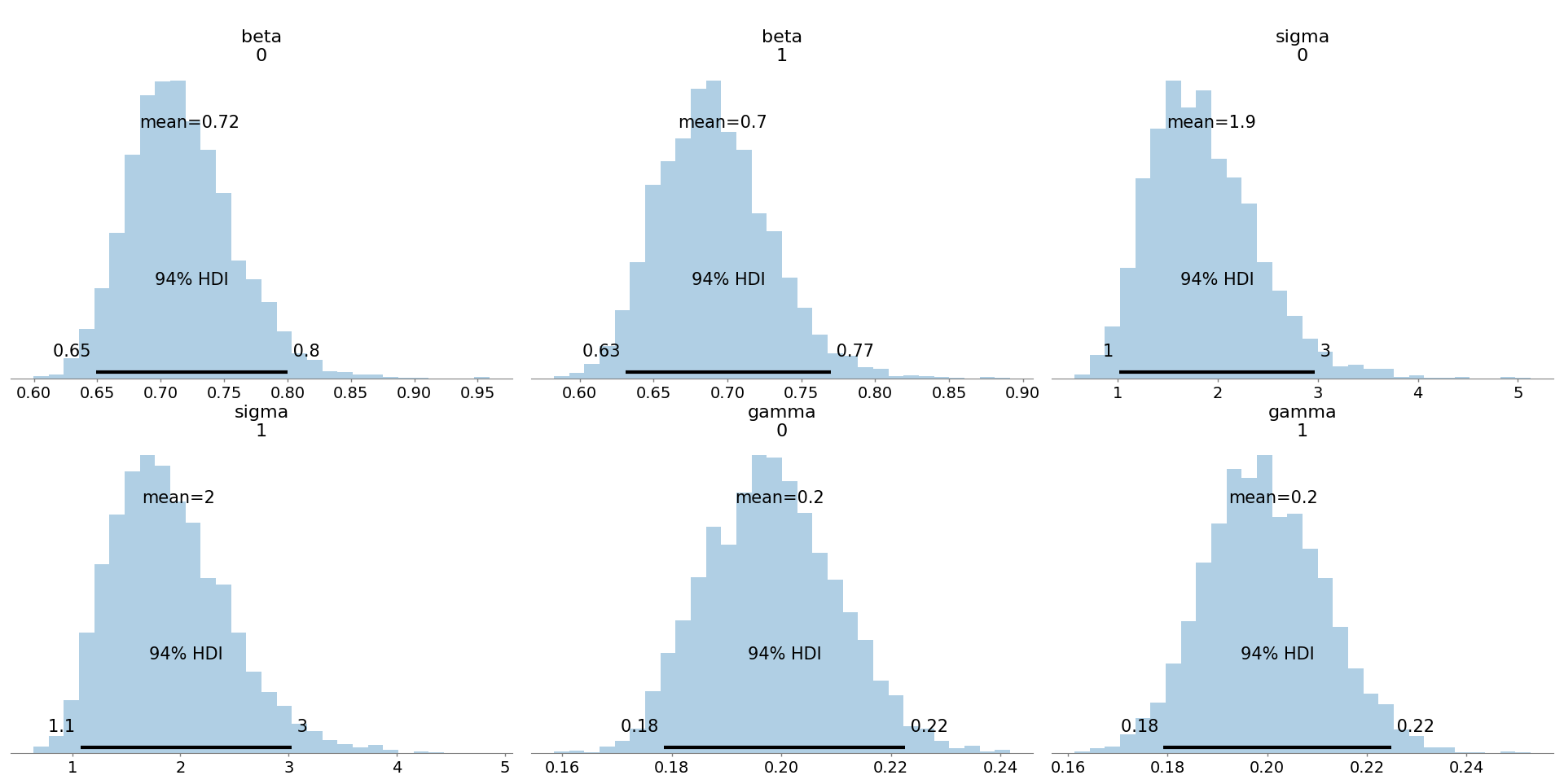
Figure 5: Group-specific posterior marginals are wider than in the single-population model, reflecting increased uncertainty due to hierarchical pooling.
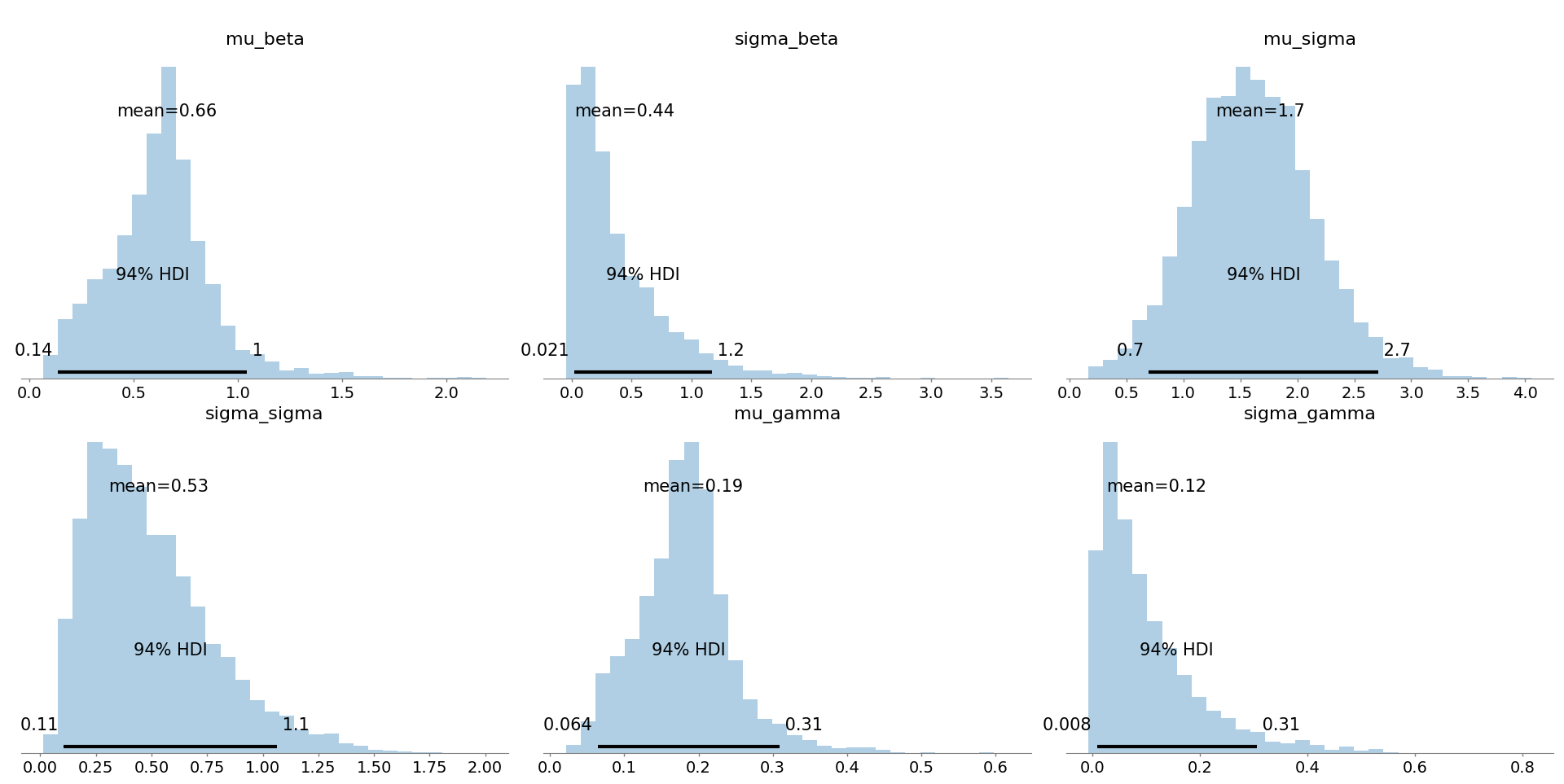
Figure 11: Hyperparameter posteriors quantify the population-level means and variabilities, offering insight into cross-group heterogeneity.
Diagnostics, Best Practices, and Practical Guidance
Diagnostics specific to ABC-SMC, such as monitoring tolerance trajectories, effective sample size (ESS), and distance distributions, are emphasized as critical for rigorous and transparent inference. Posterior predictive checks are identified as essential validations to ensure that the inferred parameter ensemble meaningfully captures key data features.
The paper provides pragmatic guidance: use full datasets over hand-crafted summaries to retain information, select distance metrics appropriate to data scaling, opt for weakly informative priors, and allow adaptive tolerances whenever feasible. These recommendations are grounded in empirical results and repeated in multiple case settings.
Implications and Future Directions
Practically, the demonstrated approach substantially lowers the barrier to entry for likelihood-free Bayesian inference in complex dynamical models, particularly for practitioners whose expertise resides in model-building rather than bespoke statistical coding. Theoretically, the results reinforce that ABC-SMC yields meaningful uncertainty quantification—even under pronounced partial observability and limited data—and facilitates hierarchical modeling for heterogeneous populations.
As advances in simulation-based inference (e.g., neural posterior estimation, synthetic likelihoods) and computational hardware continue, ABC-SMC and its close relatives will be increasingly pivotal. Algorithmic developments—such as improved summarization for very high-dimensional outputs, robust adaptive tolerance schedules, and variance reduction techniques—are likely directions for further research. Furthermore, integration into probabilistic programming ecosystems will foster greater accessibility, reproducibility, and extensibility for domain scientists.
Conclusion
This work provides a technically rigorous, pragmatic roadmap for likelihood-free inference in mechanistic dynamical systems, emphasizing the role of simulation-based methods where traditional Bayesian approaches are infeasible. Through comprehensive examples and clear PyMC implementations, the paper demonstrates that ABC-SMC achieves robust uncertainty quantification and interpretable posterior predictive analyses, even in challenging settings of partial observability and heterogeneous populations. The codebase and workflow are well-positioned to serve as foundational tools for future research in computational ecology, epidemiology, and applied dynamical modeling.







