FALCON: Few-step Accurate Likelihoods for Continuous Flows
Abstract: Scalable sampling of molecular states in thermodynamic equilibrium is a long-standing challenge in statistical physics. Boltzmann Generators tackle this problem by pairing a generative model, capable of exact likelihood computation, with importance sampling to obtain consistent samples under the target distribution. Current Boltzmann Generators primarily use continuous normalizing flows (CNFs) trained with flow matching for efficient training of powerful models. However, likelihood calculation for these models is extremely costly, requiring thousands of function evaluations per sample, severely limiting their adoption. In this work, we propose Few-step Accurate Likelihoods for Continuous Flows (FALCON), a method which allows for few-step sampling with a likelihood accurate enough for importance sampling applications by introducing a hybrid training objective that encourages invertibility. We show FALCON outperforms state-of-the-art normalizing flow models for molecular Boltzmann sampling and is two orders of magnitude faster than the equivalently performing CNF model.
Paper Prompts
Sign up for free to create and run prompts on this paper using GPT-5.
Top Community Prompts
Explain it Like I'm 14
What is this paper about?
This paper introduces a new way to quickly and accurately generate random 3D shapes of molecules that follow the rules of physics at room temperature. These rules are captured by something called the Boltzmann distribution, which says low-energy shapes happen more often than high-energy ones. The method is called FALCON, and it focuses on being both fast (using only a few steps) and precise (so scientists can trust the results).
What questions were the researchers trying to answer?
The paper asks:
- How can we make a model that generates realistic molecule shapes fast, without needing thousands of tiny steps?
- Can we still compute how “likely” each generated shape is (its likelihood), which is needed to correct and trust the samples in scientific tasks?
- Can this model be good enough to beat the best existing methods on real molecule datasets?
How did they do it?
To understand their approach, it helps to picture a landscape of hills and valleys:
- Each possible molecule shape is a point on the landscape.
- The height at that point is its energy.
- The Boltzmann distribution says “low valleys” (low energy) are more common than “high hills.”
Scientists want to randomly sample points in this landscape in a way that matches the Boltzmann distribution. That’s hard because the landscape is rough and high-dimensional.
The challenge with existing methods
A popular approach uses continuous normalizing flows (CNFs), which move points smoothly from a simple starting distribution (like Gaussian noise) to the realistic molecule distribution. But:
- CNFs need to take lots of tiny steps to be accurate.
- Computing the likelihood (how probable a sample is under the model) is extremely expensive, often requiring hundreds to thousands of function calls per sample.
This makes CNFs slow and hard to use at scale.
The FALCON idea
FALCON builds a “map” that transforms simple random inputs into realistic molecule shapes in just a few steps. Think of it like taking a few big strides instead of many tiny steps.
Crucially, the map is designed to be reversible (invertible). That means you can go forward (noise → molecule) and back (molecule → noise). Reversible maps let you quickly compute the likelihood using the change-of-variables formula, which is essential for scientific correction methods like importance sampling.
Training the map (hybrid loss)
They train the map using a hybrid objective:
- A regression part teaches the model the average direction to move from noise toward data (like learning a shortcut).
- An invertibility part (cycle consistency) checks that if you go forward and then back, you return to where you started. This encourages the map to be truly reversible even before perfect training.
In simple terms: the model learns fast jumps that are carefully designed so it can still undo them. That makes the likelihood both accurate and cheap to compute.
Fast sampling and computing likelihoods
Because FALCON uses only a few steps and keeps the map reversible, it:
- Generates samples quickly.
- Computes likelihoods efficiently.
- Supports flexible, powerful neural network architectures (like Transformers), which improve quality without making sampling slow.
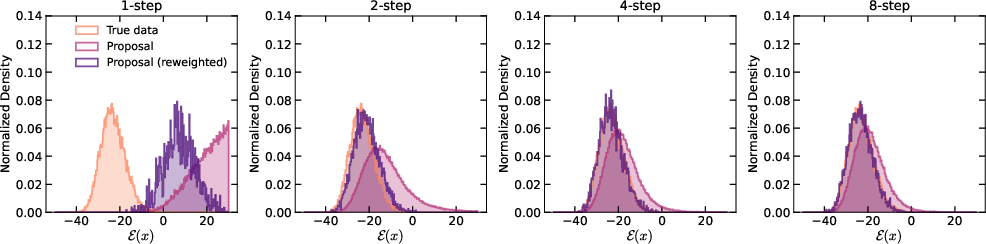
They also found that using smart step schedules (more steps near the end) improves sample quality, similar to strategies used in diffusion models.
What did they find and why it matters?
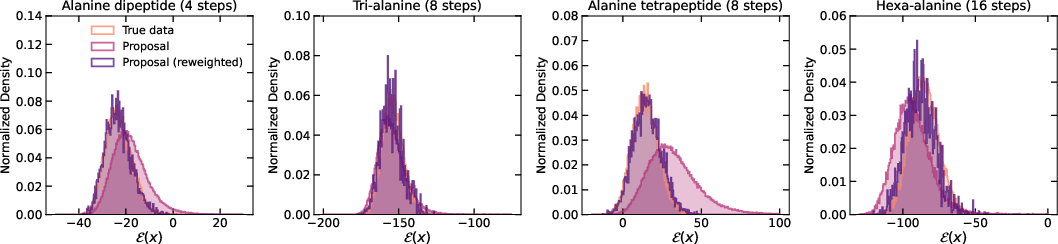
Across several molecule datasets (alanine dipeptide, tri-alanine, tetrapeptide, hexa-alanine), FALCON:
- Produced higher-quality samples than leading continuous-flow models (like ECNF++) when scaled to larger molecules.
- Beat strong discrete-flow baselines (like SBG), even when those baselines used vastly more samples.
- Was around 100 times faster than an equally good CNF when measuring the time needed to generate samples with accurate likelihoods.
They measured performance using:
- Effective Sample Size (ESS): higher is better; it means more of your samples are truly useful after reweighting.
- Wasserstein distance on energy: lower means the generated energy distribution matches the true one more closely.
- Wasserstein distance on dihedral angles (global shape features): lower means better coverage of realistic molecular shapes.
FALCON consistently improved these metrics and made sampling much faster.
Why is this important?
- Faster science: Many tasks in drug design, biomolecules, and materials rely on sampling from the Boltzmann distribution to estimate physical properties. FALCON makes this practical at larger scales.
- Accurate corrections: Because FALCON computes reliable likelihoods, scientists can use importance sampling to correct the generated samples so they match the true distribution, ensuring trustworthy results.
- Better models with fewer steps: The approach shows that you can get the speed of few-step flows without losing the ability to compute precise likelihoods—something many fast generators do not provide.
- Broader impact: The idea of training reversible few-step maps could help other fields that need fast generation with accurate probabilities, from physics to graphics to AI safety.
In short, FALCON combines speed and scientific rigor: it generates realistic molecules quickly, computes how likely they are, and supports scaling to complex systems—pushing Boltzmann Generators closer to real-world use.
Knowledge Gaps
Knowledge gaps, limitations, and open questions
Below is a concise, actionable list of what remains missing, uncertain, or unexplored in the paper, intended to guide future research:
- Theoretical guarantees for invertibility under the proposed cycle-consistency loss
- Provide conditions under which minimizing the invertibility loss yields a globally invertible map in practice (not only at the global minimum and under smoothness assumptions).
- Derive bounds that relate the invertibility loss to the minimum singular value or condition number of the Jacobian, to ensure numerical stability and well-conditioned likelihoods.
- Quantifying likelihood accuracy for importance sampling
- Establish explicit error bounds connecting cycle-consistency error and discretization error to bias/variance of SNIS estimators.
- Calibrate the accuracy of likelihoods by benchmarking against CNF likelihoods or ground-truth (when available) to determine “how accurate is accurate enough” for IS.
- Scalability of Jacobian determinant computation
- Assess how computing det(I + (t − s)∂u/∂x) scales with dimensionality in practice and identify breakpoints where O(d3) operations become dominant.
- Explore architectures with tractable Jacobians (e.g., triangular, block-triangular, coupling-based, or low-rank/sparse structures) to preserve few-step sampling while keeping likelihoods tractable in higher dimensions.
- Extension to larger and more realistic molecular systems
- Evaluate on larger peptides/proteins, nucleic acids, and ligand–protein complexes; quantify how performance, ESS, and runtime scale with system size.
- Investigate feasibility for explicit solvent systems, where dimensionality explodes (orders of magnitude larger than the peptides tested).
- Symmetry handling and invariance gaps
- Incorporate and evaluate strict SE(3) equivariance and permutation invariance of identical atoms (beyond soft augmentation) to assess generalization and sample efficiency.
- Quantify the impact of soft vs. hard equivariance on likelihood calibration and IS variance.
- Mode coverage and proposal robustness
- Systematically study behavior when the biased training dataset misses metastable modes; quantify how missing modes affect SNIS variance, ESS, and observables.
- Develop diagnostics and mechanisms (e.g., exploration schedules, bridging, or adaptive training) for discovering unseen modes.
- Hyperparameter sensitivity and automatic regularization
- Characterize sensitivity to the regularization weight λr; develop adaptive strategies that target a desired invertibility/conditioning threshold without degrading sample quality.
- Provide principled criteria to tune and monitor invertibility (e.g., Jacobian spectral norms, inverse reconstruction error) during training.
- Disentangling architectural gains from methodological gains
- Perform controlled ablations with matched-capacity architectures across FALCON, CNFs, and discrete NFs to isolate contributions from the few-step training objective vs. DiT-scale capacity.
- Inference schedule selection and generalization
- The EDM schedule works well on alanine dipeptide; evaluate across AL3/AL4/AL6 and larger systems, and develop data-driven or error-controlled rules for selecting schedules and step counts per sample.
- Handling of periodic and constrained molecular manifolds
- Address periodicity in dihedral angles and rigid-body or bond-length constraints (e.g., via coordinate reparameterization or constrained flows) to improve physical validity and reduce weight variance.
- Base distribution and path design
- Analyze sensitivity to the choice of base distribution p0 and interpolation path between p0 and p1; design base/path choices that improve conditioning and invertibility.
- Integration with advanced IS/SMC strategies
- Explore combining FALCON with SMC, tempering, or bridging distributions to reduce weight degeneracy and improve ESS without sacrificing few-step efficiency.
- Robustness to different force fields and temperatures
- Assess transferability across force fields (e.g., AMBER, CHARMM, OPLS) and temperatures; quantify how likelihood calibration and ESS degrade under distribution shift.
- Dataset size and bias dependence
- Study sample efficiency with respect to the size, bias, and diversity of the training set; provide learning curves and failure cases.
- Numerical stability and failure modes
- Characterize where the Jacobian becomes ill-conditioned, leading to unstable or unreliable determinants; develop safeguards (e.g., spectral normalization, Lipschitz constraints) or fallback strategies.
- Likelihood computation details and benchmarking
- Provide methodology and profiling for Jacobian/determinant computation (exact vs. approximate), memory usage, and wall-clock breakdown; compare to CNF trace-based computations across dimensions.
- Physical validity and chemical constraints
- Evaluate whether generated conformations respect stereochemistry, chirality, and steric constraints; quantify the fraction of physically invalid samples and their impact on weights and ESS.
- Open-source reproducibility and implementation details
- Release code and detailed configurations for Jacobian/determinant computation, JVP usage, and inverse verification experiments to enable community validation and extension.
- Theoretical connection to continuous flows
- Formalize how average-velocity regression plus cycle-consistency approximates the continuous probability flow; derive error bounds between discrete FALCON maps and the continuous-time flow.
- Generalization beyond molecules
- Test FALCON on other scientific domains with rugged energies (e.g., materials, protein–ligand binding, lattice systems) to assess generality and identify domain-specific adaptations.
Practical Applications
Immediate Applications
The following applications can be deployed now by leveraging FALCON’s few-step, invertible flow maps with accurate likelihoods and SNIS reweighting. Each item includes sector(s), potential tools/workflows, and key feasibility assumptions.
- Faster Boltzmann Generator for small-to-medium molecules
- Sectors: healthcare (drug discovery), materials/chemicals, academia
- What: Replace CNF-based generators with FALCON to produce i.i.d.-like equilibrium samples, compute consistent thermodynamic observables (e.g., free energies, heat capacities) via SNIS, and reduce inference time by orders of magnitude.
- Tools/workflows: “FALCON-BG” Python library; integration adapters for OpenMM/GROMACS; SNIS utilities for observables; DiT-based model configs for peptides/small molecules; EDM sampling schedule presets.
- Assumptions/dependencies: Access to a computable energy function E(x) (force fields or ML potentials) and a small biased dataset; scheduler/step count tuned to target ESS; energy evaluation cost dominates overall throughput; soft SO(3) equivariance is sufficient for target systems.
- Accelerated estimation of thermodynamic observables with SNIS
- Sectors: healthcare (pharmacology), materials/chemistry, academia
- What: Generate ensembles in few steps; compute consistent Monte Carlo estimates of observables using SNIS weights w(x) ∝ exp(−E(x))/pθ(x); rapidly converge by scaling sample count.
- Tools/workflows: “FALCON-Observables” module; ESS tracking; variance diagnostics; automatic sample scaling; report templates for property tables.
- Assumptions/dependencies: Reliable energy evaluations; adequate ESS; correct likelihood evaluation via invertibility regularization; unbiased test set for validation.
- Proposal generation to seed MD/MCMC and improve mixing
- Sectors: materials/chemistry, academia
- What: Use FALCON to propose cross-basin configurations that jump between metastable modes, reducing correlation and burn-in; plug into hybrid MD/MCMC schemes.
- Tools/workflows: “FALCON-Proposals” for hybrid samplers; acceptance-rate monitoring; schedule optimization; partial refresh of MD trajectories from FALCON proposals.
- Assumptions/dependencies: Accurate likelihoods for acceptance tests; acceptance hinges on energy compatibility; mode coverage depends on training data diversity.
- Physics-aware conformer ensembles for ligand and small molecule modeling
- Sectors: healthcare (medicinal chemistry), software (cheminformatics)
- What: Generate conformers with SNIS-corrected weights for docking, alchemical free energy calculations, and QSAR pipelines, improving pose diversity and thermodynamic realism.
- Tools/workflows: RDKit plugin “FALCON-Conformers”; integration with docking (e.g., AutoDock, gnina) and FEP tools; batch pipelines with energy evaluation.
- Assumptions/dependencies: Valid ligand force fields or ML potentials; implicit/explicit solvent modeling consistency; manageable energy evaluation costs at scale.
- Drop-in replacement or complement to discrete normalizing flows (e.g., SBG)
- Sectors: healthcare/materials, academia, software
- What: Use FALCON for few-step sampling with accurate likelihoods; combine with existing NF pipelines to reduce variance and improve E/T metrics; swap modules incrementally.
- Tools/workflows: Adapters/wrappers for SBG/TARFlow interfaces; side-by-side ESS/Wasserstein benchmarking; fallback to discrete NF when invertibility is insufficient.
- Assumptions/dependencies: Interoperability in data formats and APIs; compatibility of equivariance handling; robust regularization tuning.
- Energy-efficient HPC operations and lab productivity
- Sectors: energy/operations, academia/industry R&D
- What: Cut number of function evaluations compared to CNFs, lowering compute cost and wall time while maintaining scientific fidelity, improving throughput on shared clusters.
- Tools/workflows: Cost calculators; throughput dashboards; step-schedule tuning (EDM) for target accuracy/time; resource planning templates.
- Assumptions/dependencies: Availability of GPU/TPU with autograd JVP support; stability of training (regression + cycle-consistency loss); ability to monitor ESS/metrics live.
- Education and reproducible research kits
- Sectors: education, academia
- What: Use FALCON for teaching flow matching, importance sampling, and thermodynamics in hands-on labs; share reproducible notebooks replicating peptide benchmarks.
- Tools/workflows: Colab/Notebook tutorials; datasets (ALDP/AL3/AL4/AL6); comparison scripts (CNF vs FALCON); scheduler ablation modules.
- Assumptions/dependencies: Public datasets; modest GPU access; minimal setup friction for students.
- Sampling with ML potentials (no gradient requirement on E)
- Sectors: materials/chemistry, software
- What: Combine FALCON with ANI/GAP/NequIP-like ML energy models to sample ensembles and compute observables even when energy gradients are unavailable or costly.
- Tools/workflows: “FALCON-MLPotential” wrappers; batched energy inference pipelines; SNIS weighting for ML-based E(x).
- Assumptions/dependencies: ML potential accuracy; inference latency; domain shift correction via SNIS; careful validation against classical force fields.
Long-Term Applications
These applications require further research, validation, scaling, or productization before broad deployment.
- Scaling to large proteins and macromolecular complexes
- Sectors: healthcare (structural biology), academia, biotech
- What: Extend softly equivariant architectures and training to flexible proteins, multi-chain complexes, and solvent effects; generate equilibrium structural ensembles for binding/functional analyses.
- Tools/workflows: “FALCON-Protein” models with stronger equivariance; memory-efficient batching; multi-resolution training; solvent-aware energies.
- Assumptions/dependencies: Larger, diverse training datasets; improved exact/soft SO(3) treatment; robust invertibility at scale; accurate solvent/temperature models.
- High-throughput, physics-aware virtual screening
- Sectors: pharma, materials discovery
- What: Screen millions of candidates with few-step sampling; compute SNIS-weighted thermodynamic properties; integrate active learning loops to prioritize synthesize-worthy molecules.
- Tools/workflows: SaaS pipeline “ThermoScreen”; queue-based sampling/energy evaluation; active retraining; property dashboards; ranking under ESS constraints.
- Assumptions/dependencies: Scalable energy evaluation (force fields/ML/DFT surrogates); cloud/HPC orchestration; careful handling of data bias and distribution shift.
- Real-time interactive molecular design
- Sectors: software (CAD for molecules), biotech/materials R&D
- What: Provide a GUI where researchers set constraints (e.g., dihedral ranges, temperature), and FALCON instantaneously samples and reweights ensembles, enabling rapid hypothesis testing.
- Tools/workflows: Web UI; streaming inference; on-demand energy services; real-time ESS monitoring; caching strategies.
- Assumptions/dependencies: Low-latency energy evaluation; responsive inference backends; robust constraint handling; usability testing.
- FALCON-assisted MCMC/SMC for broader inference tasks
- Sectors: academia (statistics/ML), finance (risk), climate science
- What: Use invertible few-step flows as proposal mechanisms in importance sampling, MCMC, or SMC for complex posteriors (e.g., climate ensemble calibration, tail risk modeling).
- Tools/workflows: “FALCON-IS” library; proposal adapters for probabilistic programming (Pyro/NumPyro/Stan); diagnostics for weight degeneracy and ESS.
- Assumptions/dependencies: A well-defined “energy” or unnormalized log-target; mapping p0→p1 approximations; domain-specific constraints and validation.
- Physics-aware generative foundation models
- Sectors: software/AI, materials/chemistry, healthcare
- What: Train large-scale FALCON models across diverse molecular families to create general-purpose generators with tractable likelihoods for downstream scientific tasks.
- Tools/workflows: “FALCON-Foundation” checkpoints; fine-tuning APIs; cross-domain adapters; evaluation suites (E/T/ESS + domain-specific metrics).
- Assumptions/dependencies: Massive datasets; substantial compute; maintaining invertibility and likelihood accuracy at scale; benchmarking standards.
- Ab initio-quality sampling with DFT/QM energies
- Sectors: materials/chemistry
- What: Pair FALCON with DFT/QM energies (or surrogates) to generate high-fidelity ensembles for small molecules and materials clusters; SNIS corrects surrogate bias toward ab initio ground truth.
- Tools/workflows: “FALCON-DFT” pipelines; hybrid training (surrogate + occasional true QM labels); uncertainty-aware reweighting.
- Assumptions/dependencies: Expensive energy labels; surrogate accuracy; data-efficient retraining; careful extrapolation limits.
- Regulatory-grade thermodynamic property estimation
- Sectors: healthcare (CMC), materials regulation
- What: Produce transparent, reproducible estimations (with ESS and error bars) of properties supporting regulatory submissions for excipients, polymers, and process conditions.
- Tools/workflows: Audit-ready logs; traceable SNIS pipelines; documentation templates; validation against experimental data.
- Assumptions/dependencies: Method validation and standardization; acceptance by regulators; alignment with experimental protocols.
- Cloud-based Boltzmann Sampling as a Service
- Sectors: software/SaaS, enterprise R&D
- What: Offer APIs to submit systems (coordinates, force fields) and receive weighted ensembles and computed observables; flexible cost tiers for sampling depth.
- Tools/workflows: Managed GPU clusters; autoscaling; SLAs; billing and usage analytics; secure data handling.
- Assumptions/dependencies: Reliable multi-tenant performance; failover strategies; integration with customer toolchains; data privacy compliance.
- Materials design for energy technologies (electrolytes, membranes)
- Sectors: energy, materials
- What: Use FALCON to explore thermodynamic stability and transport-relevant conformational ensembles under varied conditions (T, P, solvent), accelerating candidate selection.
- Tools/workflows: Condition-swept sampling; property calculators (e.g., solvation free energies); decision dashboards; lab-to-sim feedback loops.
- Assumptions/dependencies: Condition-dependent energy models; calibration to experimental data; multi-scale validation (molecule → device).
- Few-step invertible generators beyond molecules (3D point clouds, audio, images) with exact/accurate likelihoods
- Sectors: software/AI, robotics (3D), media
- What: Adapt FALCON’s invertibility-regularized flow maps to domains needing tractable likelihoods (compression, anomaly detection, 3D scene sampling), preserving fast inference.
- Tools/workflows: Domain-specific architectures; dataset adapters; evaluation (likelihood, coverage, fidelity); integration with downstream pipelines.
- Assumptions/dependencies: Appropriate “energy” or loss analog; suitable invariances; empirical confirmation of likelihood accuracy and invertibility in new domains.
Cross-cutting assumptions and dependencies
- Energy function E(x): Must be computable quickly and accurately (classical force fields, ML surrogates, or QM). Poor E(x) fidelity directly degrades SNIS correction and observables.
- Training data: Requires a small biased dataset; coverage and quality impact mode coverage and ESS.
- Invertibility and likelihood: Relies on cycle-consistency regularization; hyperparameters (e.g., λr) control the trade-off between sample quality and invertibility; invertibility should be verified empirically (error ~1e−4 in reported experiments).
- Schedules and steps: Performance depends on inference schedule (EDM preferred in experiments) and step count; users should tune for desired accuracy/time trade-offs.
- Scalability: Results demonstrated up to hexa-alanine systems; larger systems will require architectural scaling, stronger equivariance, and more data.
- Compute environment: Autograd with JVP, GPU acceleration, and efficient energy evaluators are important for practical throughput; monitoring ESS and diagnostics is critical for reliability.
Glossary
- 2-Wasserstein distance: A metric measuring the distance between probability distributions based on optimal transport. "the $2$-Wasserstein distance on both the energy distribution (E), and dihedral angles (T)."
- amber-14 force field: A specific molecular mechanics potential used in MD to compute energies and forces. "implicit solvent molecular dynamics (MD) simulations with the amber-14 force field"
- average velocity objective: A training objective that fits the time-averaged velocity between two times to enable few-step generation. "to minimize the average velocity objective:"
- Boltzmann distribution: The probability distribution over molecular states proportional to the exponential of negative energy. "Sampling molecular configurations from the Boltzmann distribution "
- Boltzmann Generators: Generative models combined with importance sampling to produce samples consistent with a target Boltzmann distribution. "Boltzmann Generators tackle this problem by pairing a generative model, capable of exact likelihood computation, with importance sampling"
- change-of-variables formula: The rule for transforming densities through an invertible mapping via the Jacobian determinant. "making the standard change-of-variables formula inapplicable."
- consistency models (CMs): Few-step generative models trained to be consistent across different time steps, enabling fast sampling. "few-step flow models such as consistency models (CMs)"
- continuous normalizing flows (CNFs): Generative models that evolve samples via ODEs and allow exact likelihood computation through continuous change of variables. "Current Boltzmann Generators primarily use continuous normalizing flows (CNFs) trained with flow matching"
- cycle-consistency term: A regularization that encourages a forward map to be invertible by penalizing round trips that don’t return to the original point. "with a cycle-consistency term to encourage invertibility"
- dihedral angles: Torsion angles defining molecular conformations, used as global structural metrics. "the $2$-Wasserstein distance on both the energy distribution (E), and dihedral angles (T)."
- diffusion transformer (DiT): A transformer-based architecture adapted for diffusion/flow models. "we use a standard diffusion transformer (DiT)~\citep{peebles_scalable_2023}"
- Dormand--Prince 4(5) integrator: An adaptive Runge–Kutta ODE solver widely used for integrating continuous-time flows. "integrated over the vector field using the Dormand--Prince 4(5) integrator with "
- EDM: A sampling schedule from the Elucidated Diffusion Model that improves few-step performance by allocating more steps near the data. "EDM substantially outperforms all other schedulers"
- Effective Sample Size (ESS): A measure of the quality of weighted samples in importance sampling reflecting the number of independent samples. "We report Effective Sample Size (ESS), and the $2$-Wasserstein distance"
- equivariant: A property of models whose outputs transform predictably under symmetry operations (e.g., rotations/translations). "a simple and scalable, softly equivariant continuous flow architecture"
- equilibrium conformation sampling: The task of generating molecular structures distributed according to thermodynamic equilibrium. "We evaluate the performance of FALCON on equilibrium conformation sampling tasks"
- flow map: A discrete-time mapping that transports samples between times, used for few-step generation. "flow map models have only been applied for fast generation"
- flow matching: A regression-based training framework that learns a continuous vector field transporting a base distribution to data. "Flow matching models~\citep{lipman_flow_2022,albergo_building_2023,liu_rectified_2022,peluchetti2021} are probabilistic generative models that learn a continuous interpolation"
- force field: A parameterized energy function defining molecular interactions for simulation and evaluation. "implicit solvent molecular dynamics (MD) simulations with the amber-14 force field"
- Geometric Vector Perceptrons (GVPs): Neural modules that process scalar and vector features with geometric constraints, used in equivariant architectures. "leveraging geometric vector perceptrons (GVPs)"
- Hutchinson's trace estimator: A stochastic estimator for the matrix trace using random probe vectors, reducing computational cost. "approximated using Hutchinson's trace estimator "
- Jacobian: The matrix of partial derivatives of a transformation, whose determinant governs density change under mapping. "full Jacobian calculations necessary for each step along the flow"
- Jacobian vector product (JVP): An efficient automatic differentiation operation computing the product of a Jacobian with a vector. "using a single Jacobian vector product (JVP) call using forward automatic differentiation."
- Monte Carlo Markov Chains (MCMC): Stochastic sampling algorithms that construct a Markov chain to sample from complex distributions. "Monte Carlo Markov Chains (MCMC)"
- Molecular Dynamics (MD): Simulation of molecular trajectories via numerical integration of Newtonian equations under a force field. "Molecular Dynamics (MD)~\citep{leimkuhler2015molecular}"
- partition function: The normalizing constant of a Boltzmann distribution obtained by integrating the unnormalized density. "with partition function "
- probability flow ODE: The ordinary differential equation whose vector field transports samples from the base to the data distribution. "the vector field of the probability flow ODE that transports samples from the noise distribution to the data distribution ."
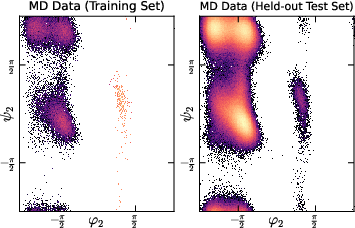
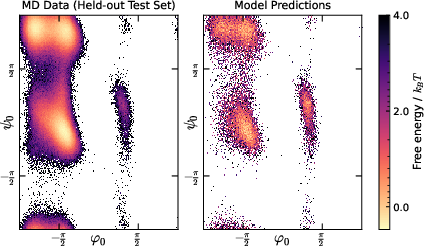
- Ramachandran plots: Visualizations of backbone dihedral angle distributions in proteins, revealing conformational modes. "with Ramachandran plots relegated to~\cref{sec:ramachandran_figures}"
- SE(3): The 3D roto-translation symmetry group, used for building rotation/translation-equivariant models. "a recent SE-equivariant architecture leveraging geometric vector perceptrons (GVPs)"
- self-normalized importance sampling (SNIS): An importance sampling method that normalizes weights to produce consistent estimators without knowing the partition function. "self-normalized importance sampling (SNIS) step to re-weight generated samples"
- Sequential Monte Carlo (SMC): A particle-based inference method that sequentially updates weighted samples to approximate target distributions. "SBG with SMC sampling (SBG SMC)"
- SO(3): The 3D rotation group, used to enforce rotational symmetry in molecular generative models. "soft SO(3) (rotation) equivariance"
- vector field: A function assigning a velocity to each point and time, defining the flow dynamics for sample transport. "the vector field of the probability flow ODE"
Collections
Sign up for free to add this paper to one or more collections.